Electron Microscopy (WIP)
Step-by-step visual guides for learning TEM. Focused on practice sitting in front of the microscope with visual elements.
WIP:
- Use real screenshots instead of pictures
- Better pictures: proportion, focus, FOV
- More visuals added to each session when needed
Disclaimer: Always follow https://barnum.su.domains/ for correctness. Only use this documentation if you are working with the authors and need quick visual references.
Available guides
| Guide | Instrument | Description | Readiness | Status |
|---|---|---|---|---|
| STEM (Spectra) | Spectra | STEM probe correction and imaging | 8/10 | Available |
| TEM (Spectra) | Spectra | Optional TEM alignment and image correction | 5/10 | Available |
| TEM (Titan) | Titan | TEM alignment and imaging | - | Coming soon |
| 4DSTEM | Spectra | 4DSTEM with Dectris detector | 2/10 | Available |
| EELS | Spectra | Electron Energy Loss Spectroscopy | 2/10 | Available |
| EDS | Spectra | Energy Dispersive X-ray Spectroscopy | 2/10 | Available |
| Tomography | Spectra | Electron tomography | - | Coming soon |
| Ptychography | Spectra | Ptychography imaging | - | Coming soon |
| MAPED | Spectra | Precession Electron Diffraction | - | Coming soon |
Instrument links:
Other resources:
Looking for volunteers!
We appreciate feedback, corrections, and contributions from the community!
- Found an error? Open an issue or submit a PR
- Want to add your institution’s SOP? Reach out to @bobleesj — we can help with writing and formatting as long as you have notes, Word docs, or rough drafts
- Have suggestions? See the GitHub repo for contribution guidelines
Acknowledgments
Authors thank Dr. Pinaki Mukherjee and Andrew Barnum for training @bobleesj and Guoliang Hu at Stanford SNSF.
Changelog
- Mar 1, 2026 - Restructure STEM and TEM guides, add pre-session checklist and end session procedures
- Feb 28, 2026 - Reorganize guide structure, rename directories, add safety warnings
- Jan 31, 2026 - Add STEM probe correction guide with screenshots from Andrew Barnum training
- Jan 31, 2026 - Add draft Titan TEM guide
- Dec 18, 2025 - Add EELS and EDS guides
- Dec 17, 2025 - Use mdBook to render static pages and host on GitHub
- Dec 14, 2025 - Begin Electron Microscopy training documentation, led by @bobleesj
Sample loading
Location
TITAN:
Sample loading is done outside of the room where Titan is hosted:

Spectra 300:
Sample loading can be done inside the Spectra room:

Sample holders are stored here at room temperature:
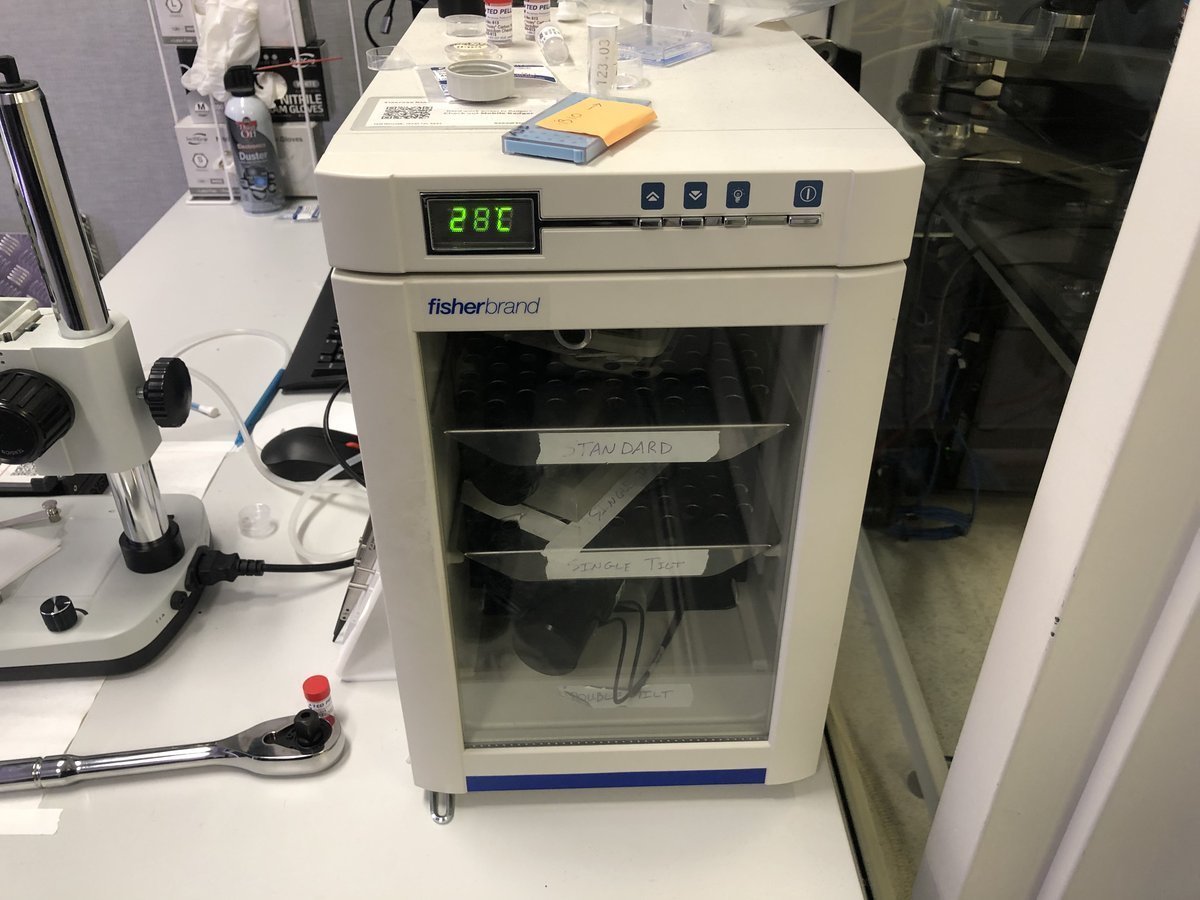

Single-tilt holder
Load sample:
FIXME: use technical terms for these objects… “pin” or “clip”?
-
Push the pin inside the tiny hole shown below:
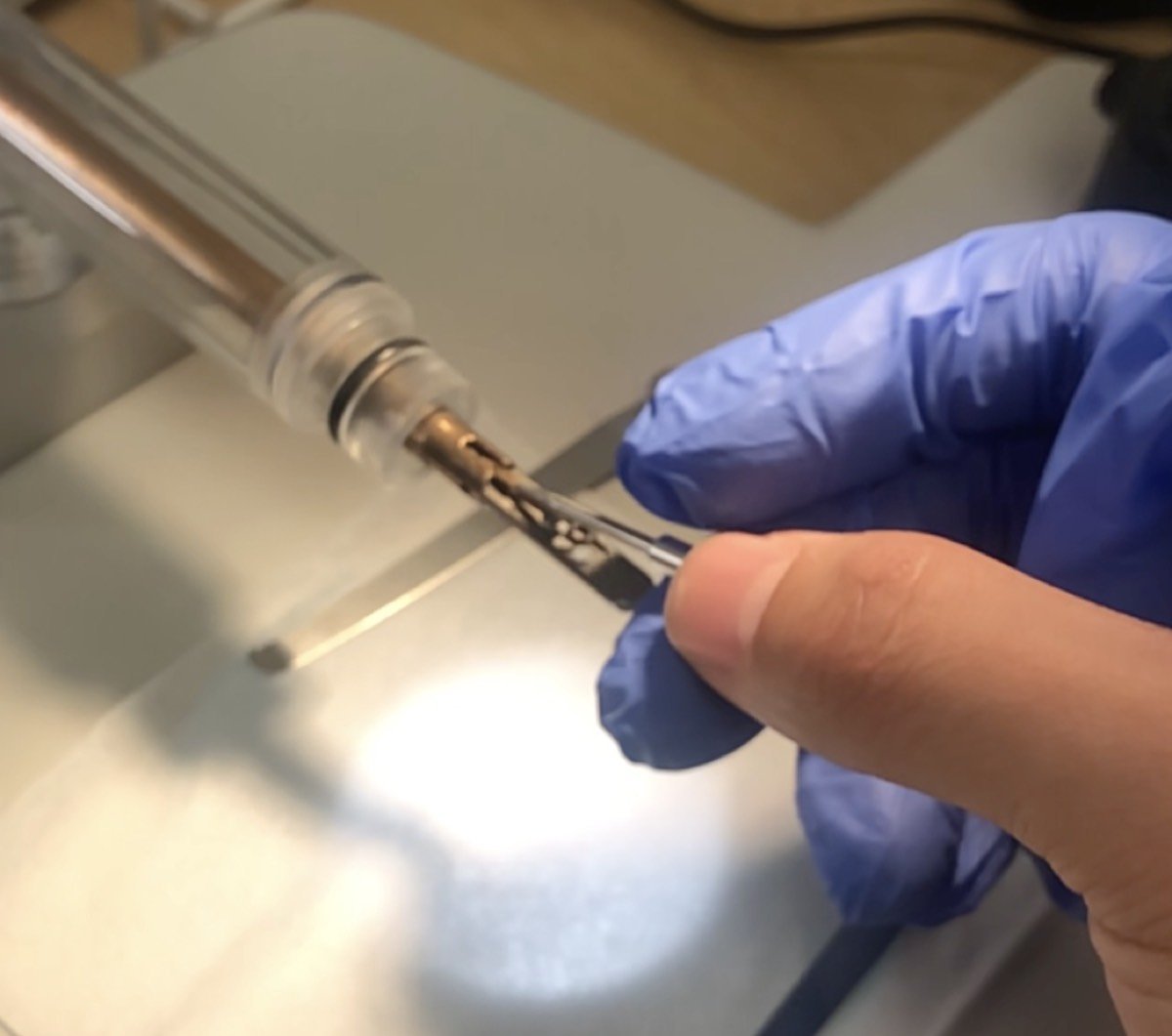

-
Lift the clip gently:
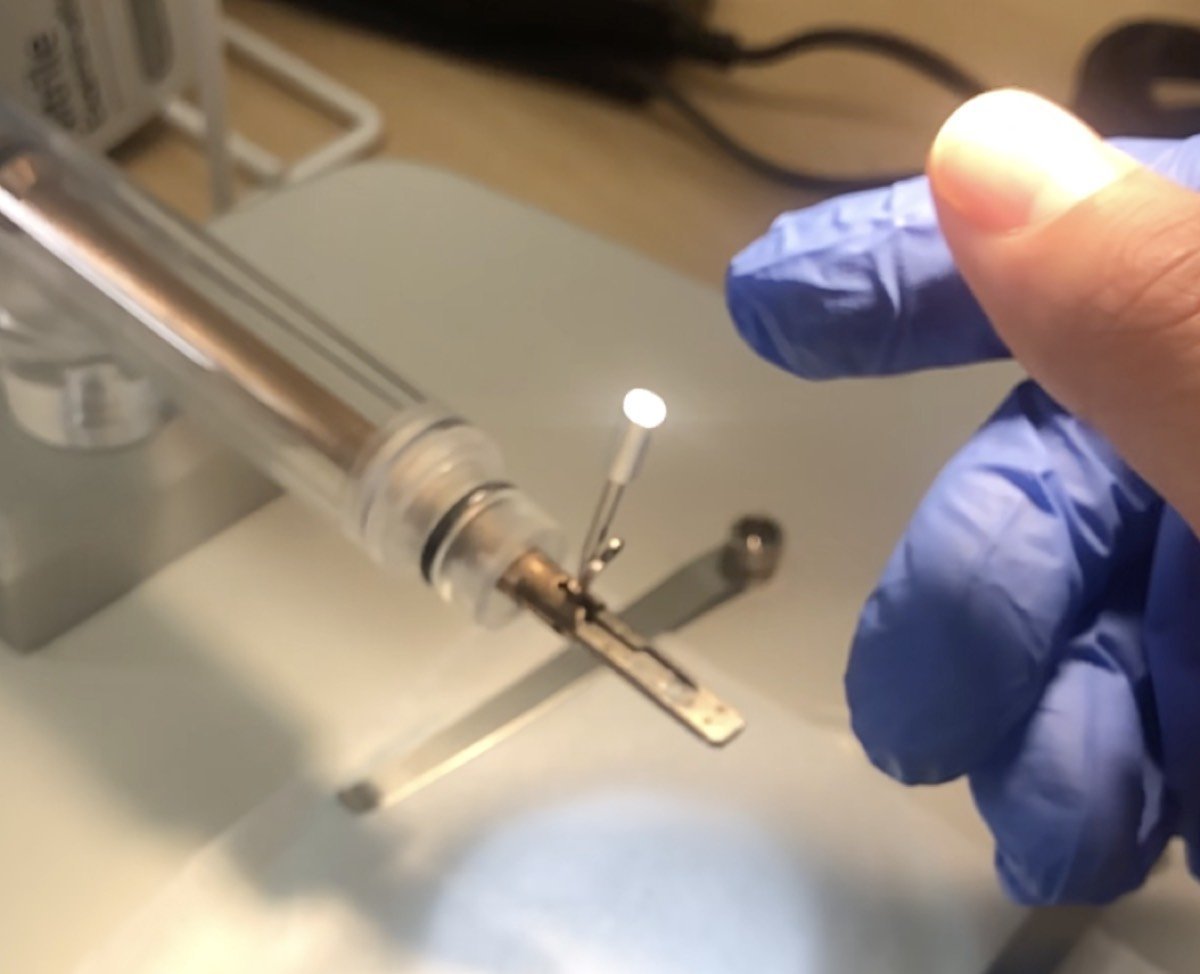

-
If using a copper grid, pinch the tip of the copper grid
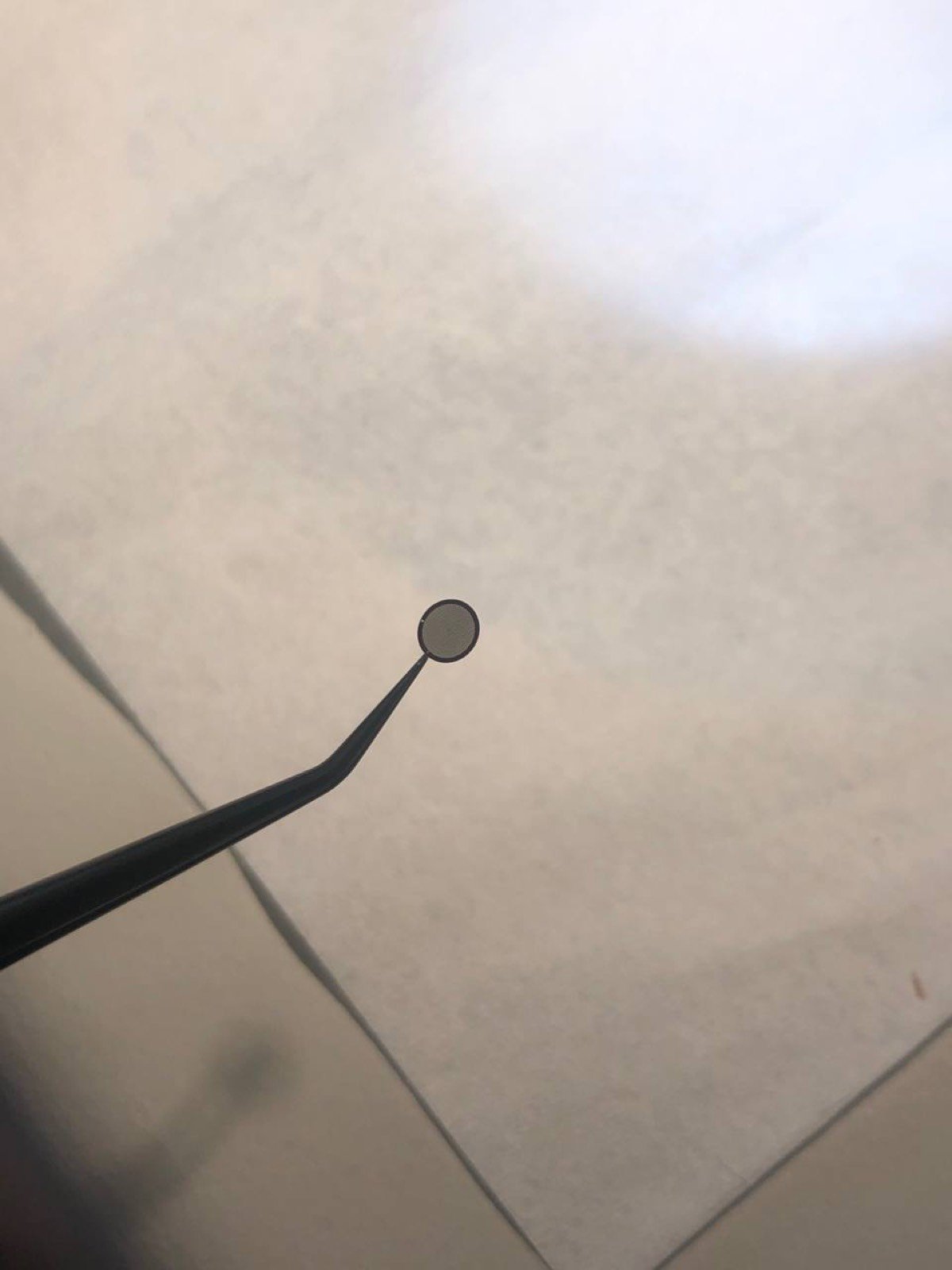

-
Place the sample, shiny side up for the standard sample
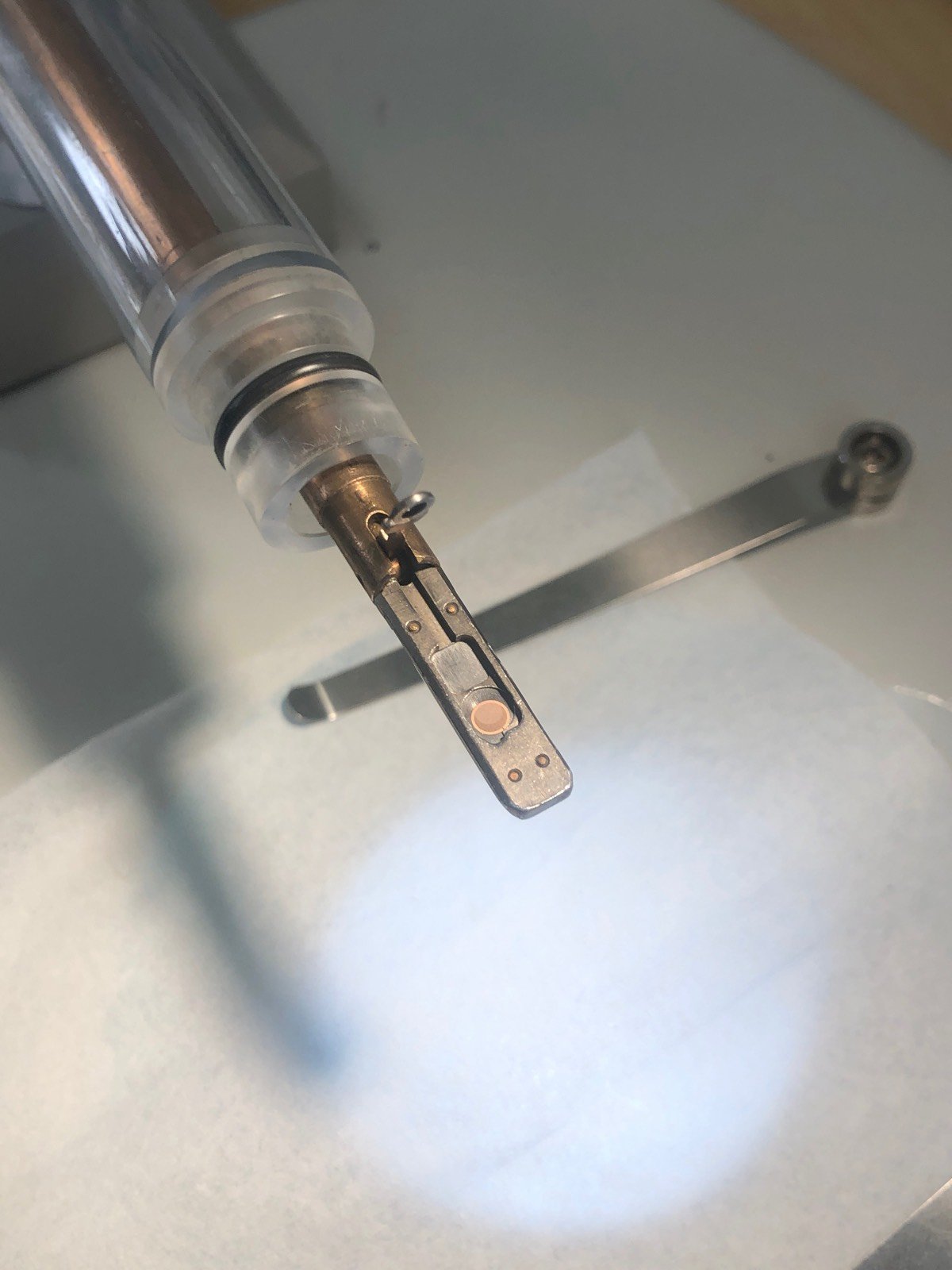

Double-tilt holder
Load sample:
-
Load the sample and washer (gold donut):
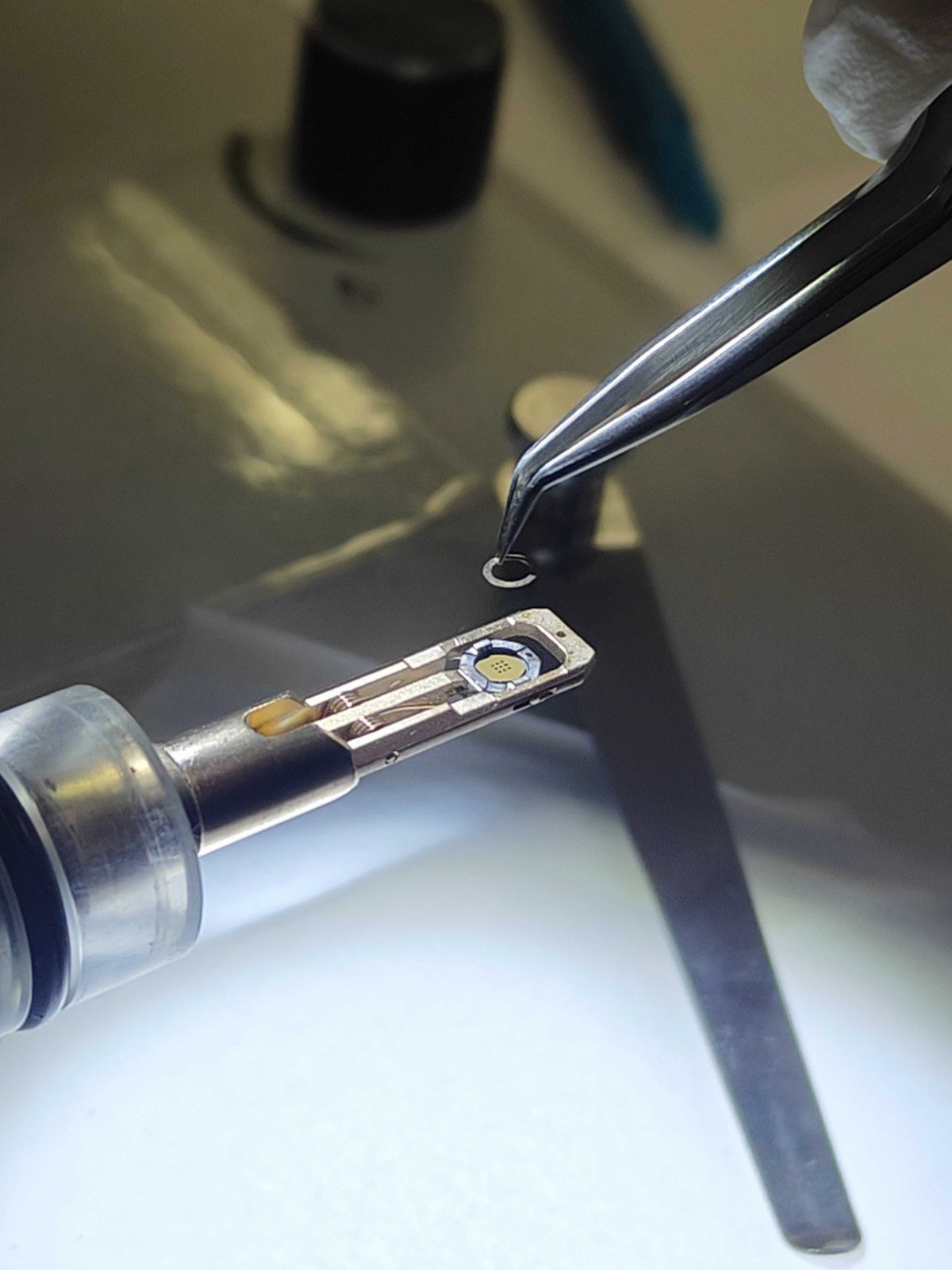

-
Add the cap and rotate the holder about the long axis to ensure the sample is secure:
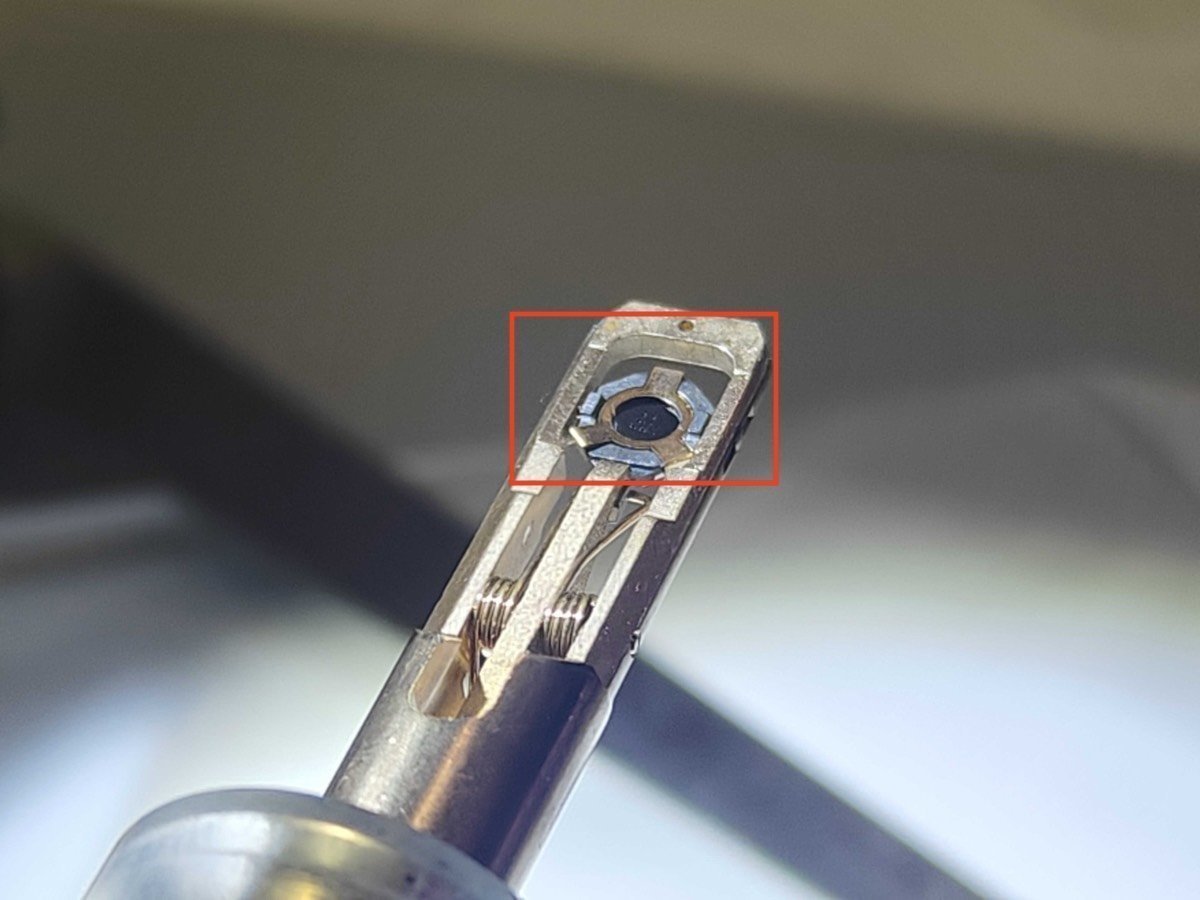

Unload sample:
-
Press down the very small hole gently as shown below:
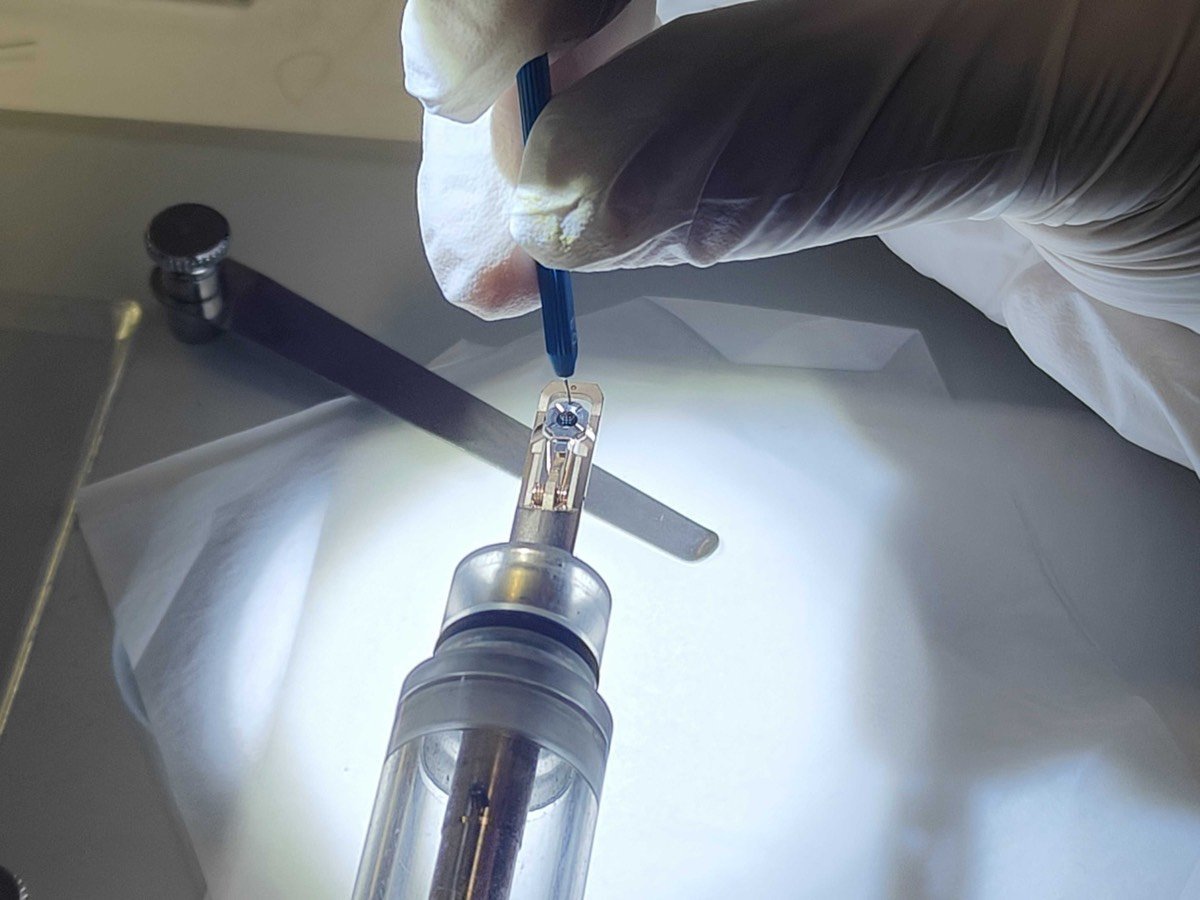

-
The three parts should all be disassembled and placed on the bottom:
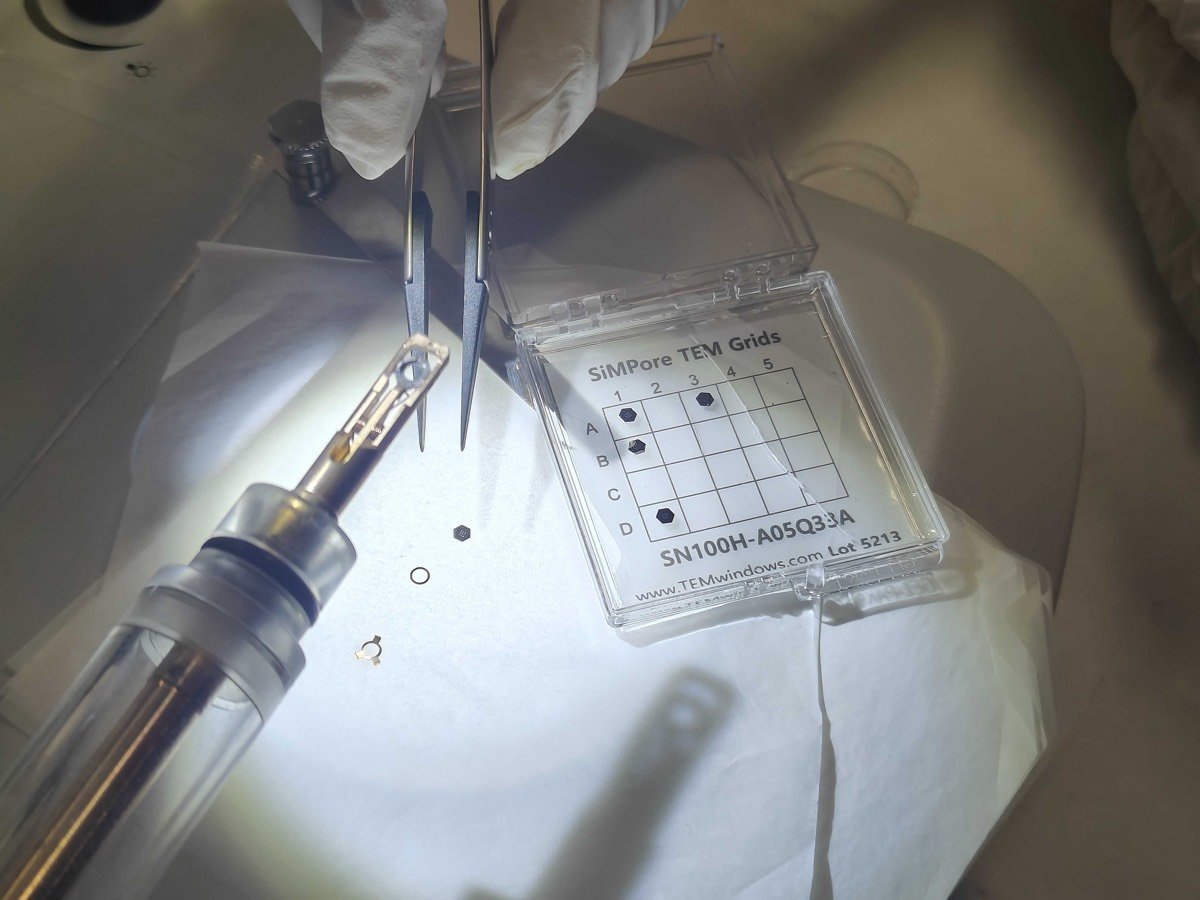
Tomography holder
Unload sample
-
The sample grid is held between the two “arms” of the metal strips.
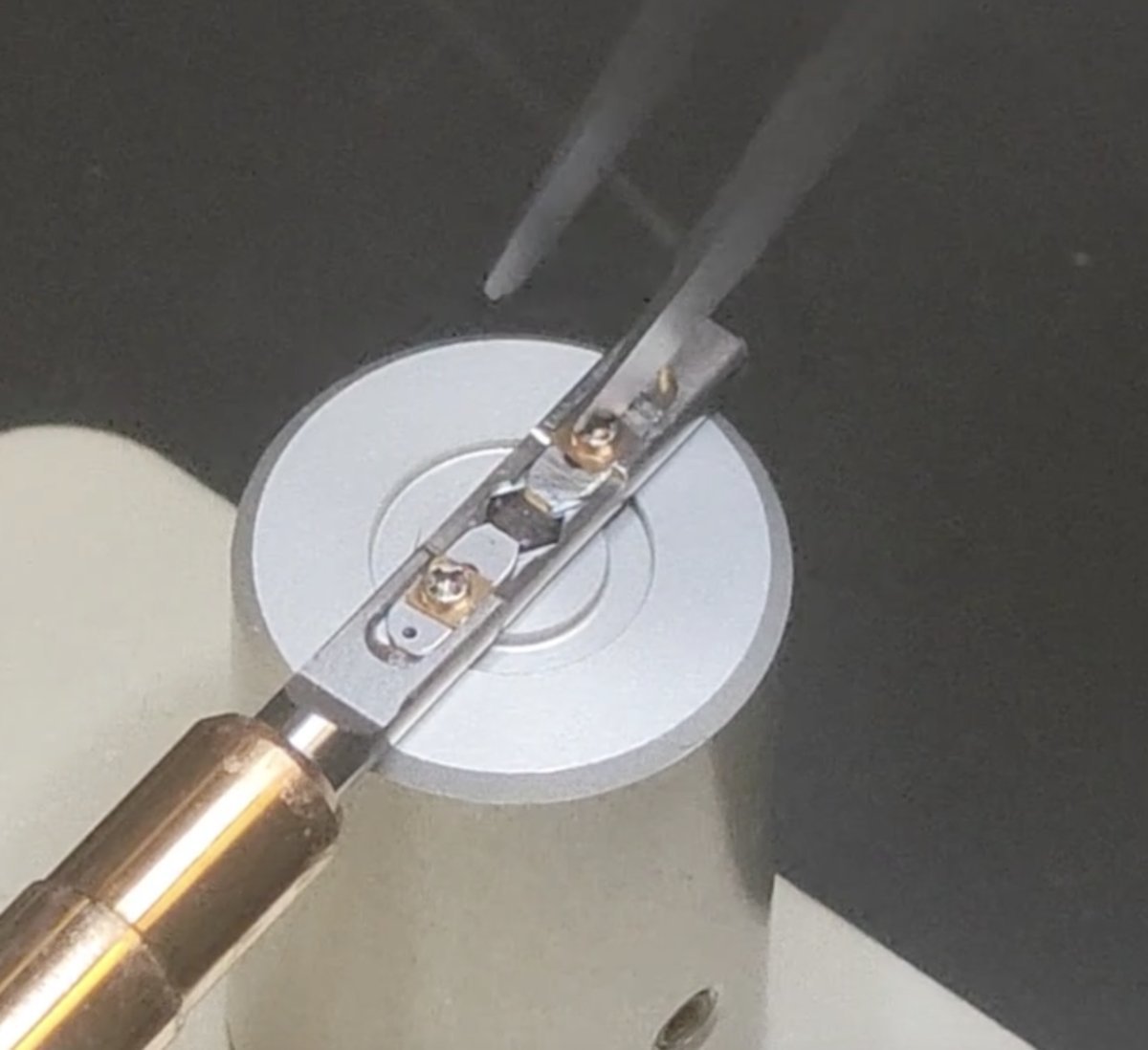

-
Rotate the screw counterclockwise about 90 degrees. Repeat for the other “arm”.
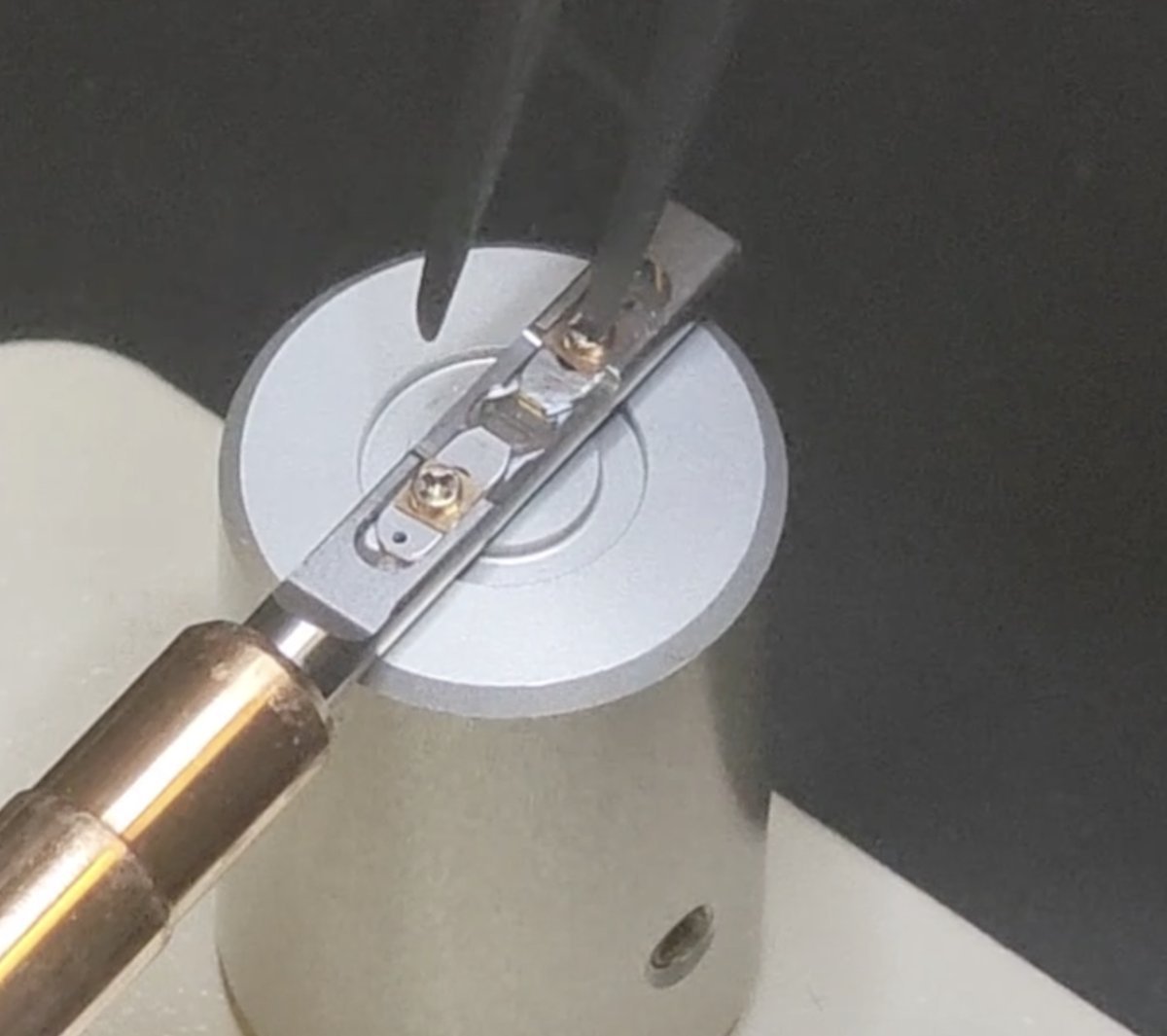

-
Move the two strips away from each other to free the sample
From:
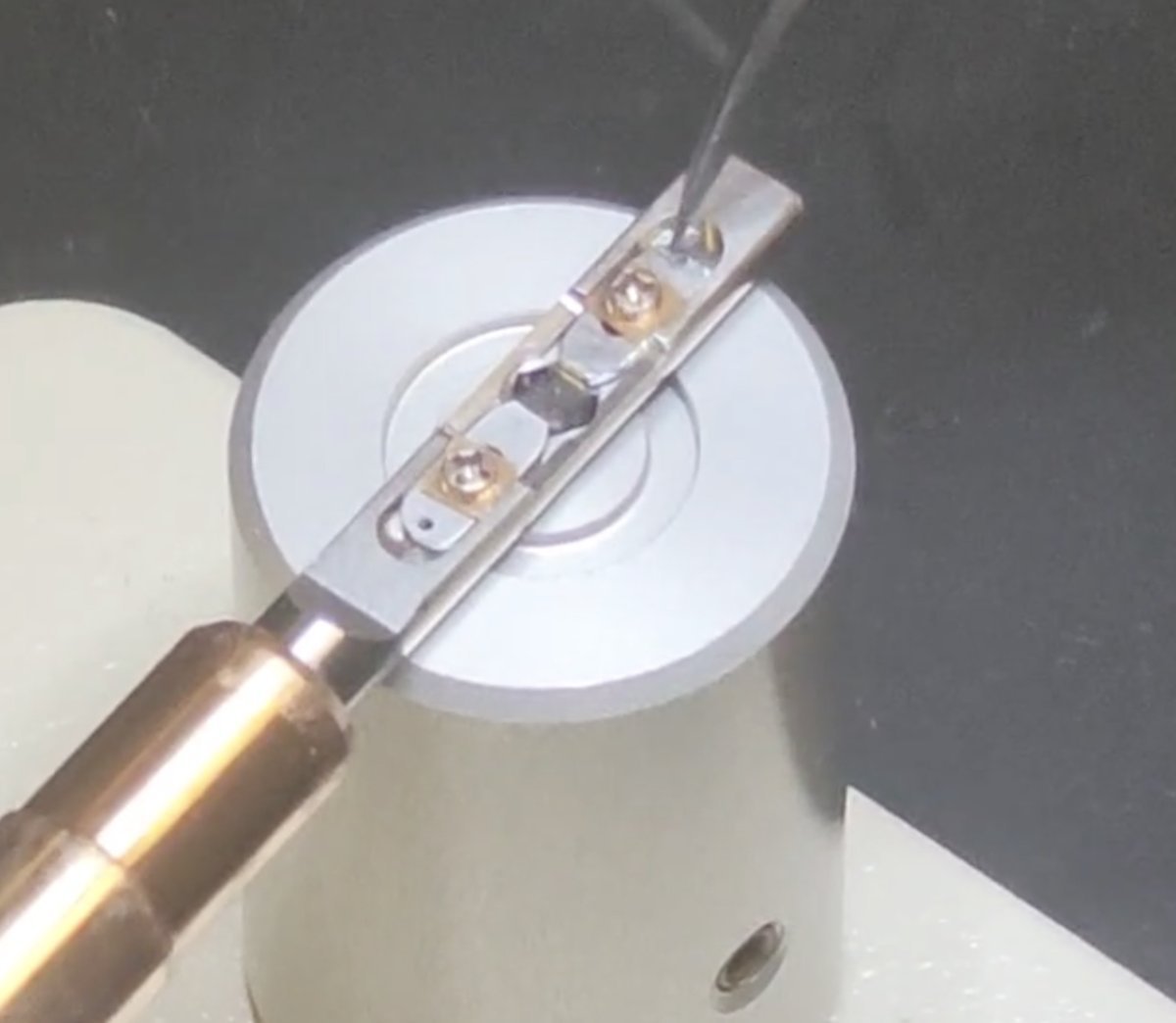

To:
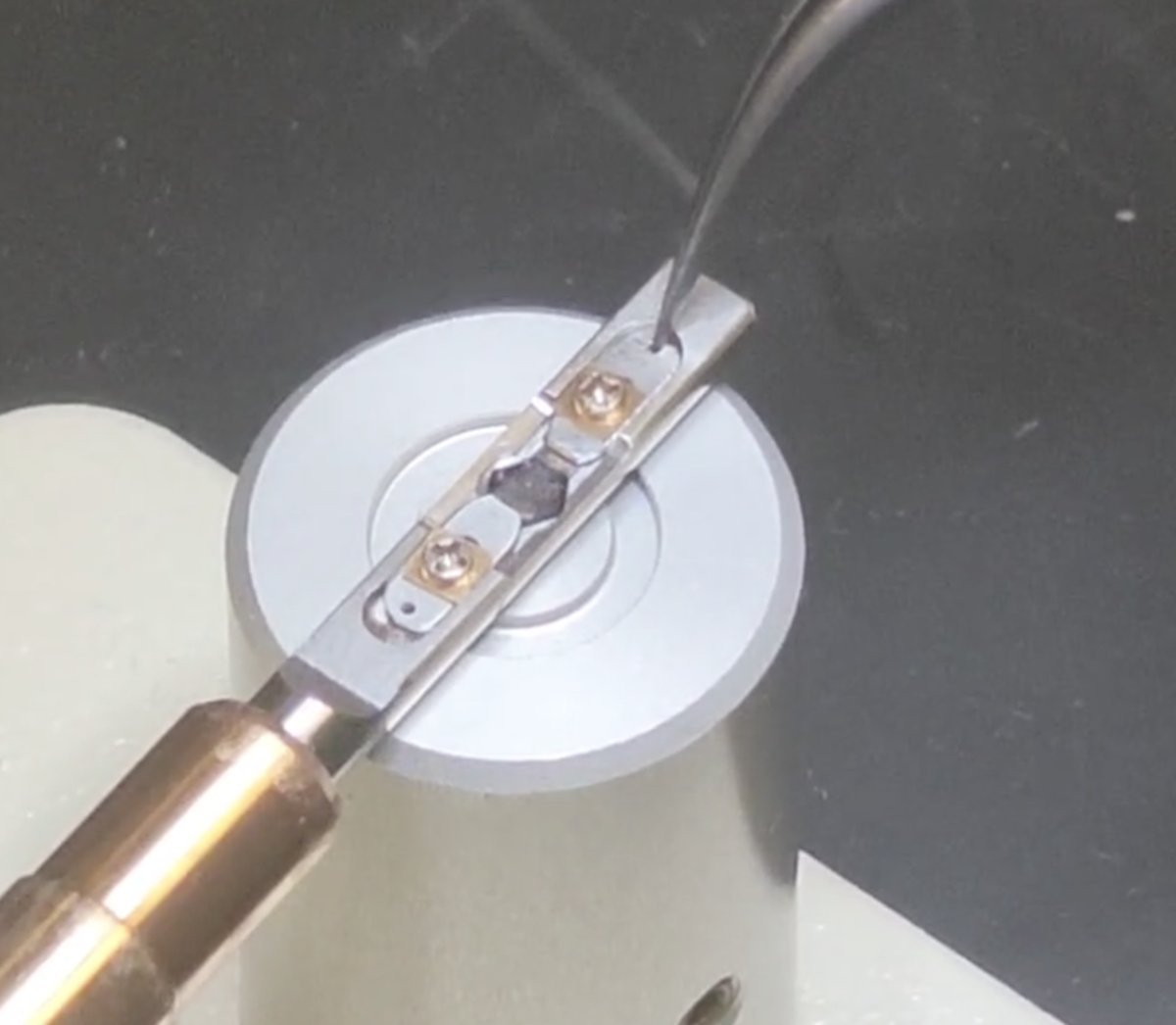

-
Remove the sample
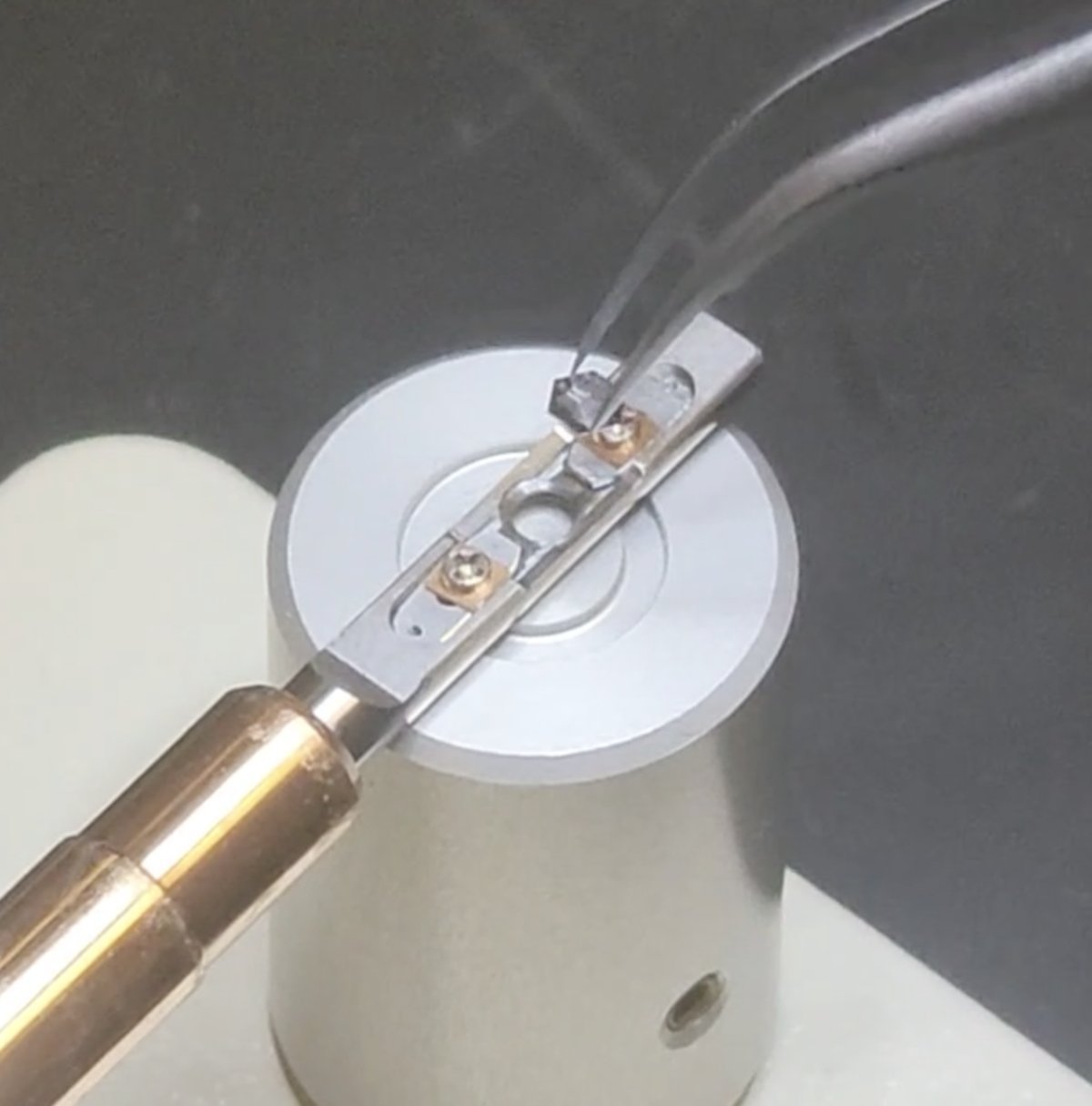

Load sample
Follow the process in reverse: load a new sample, pull the two strips closer together, and rotate the screws clockwise.
Changelog
- Dec 20, 2025 - Rnhance holder insersion process with visuals
- Dec 18, 2025 - Reorganize into Start/End session sections for use across all tutorials by @bobleesj
- Dec 17, 2025 - Add sample loading region by @bobleesj
- Dec 15, 2025 - Add tomography holder section by @bobleesj, with images taken by Guoliang Hu
- Dec 13, 2025 - Add single-tilt, double-tilt holder section by @bobleesj, with images taken by Guoliang Hu
TEM (Titan)
Coming soon.
This guide covers TEM alignment and imaging on the FEI Titan at Stanford SNSF.





Changelog
- Feb 28, 2026 - Add Titan instrument photos
Spectra 300 STEM Alignment Guide (DRAFT)

This guide covers STEM alignment on the Spectra 300 at Stanford SNSF (Stanford Nano Shared Facilities). Screenshots and instructions are provided by Andrew Barnum. Written instructions and images are organized by Sangjoon Bob Lee.
Check before starting your session
First, visually confirm the following from the previous user to ensure no damage has occurred.
- Standard gold nanoparticle sample on a single-tilt holder is loaded.
- Logbook is checked for any notes from the previous user.
- Start your session on NEMO.
- Screen is inserted.
- Beam is blanked.
- Column valves are closed.
- Turbo pump is off.
- Stage tilt is at 0° (alpha and beta) and the stage has been reset.
- Arina detector is retracted.
- Arina detector is turned off.
- All holders are capped and placed in the holder box.
- No errors are found across all software programs including TEMUI.
Report immediately in the logbook if anything has occurred or contact supervisors.
After you have checked the states,
- Follow any special instructions and warnings posted on NEMO.
- Emergency contacts are available.
Acronyms:
mulXY- Multifunction X/Y knobs on hand panelTEMUI- TEM User Interface (software)
Workstation layout:
| Monitor | Software | Purpose |
|---|---|---|
| Bottom left | TEMUI | Microscope control, vacuum, alignments |
| Bottom right | Velox | Live imaging, acquisition |
| Top left | Probe Corrector S-CORR | Aberration measurement & correction |
| Top right | Velox image gallery | Captured images from Velox |
Overview
This guide covers three main phases:
| Phase | Procedures | Time |
|---|---|---|
| Part 1: Setup & Alignment | Vacuum check, eucentric height, STEM mode configuration, direct alignments, monochromator tune, HAADF setup | 5-10 min |
| Part 2: Probe Correction | Correct aberrations (C1A1, Tableau) to achieve sub-angstrom probe | 10-15 min |
| Part 3: Imaging | Image acquisition, Sherpa fine-tuning (optional) | varies |
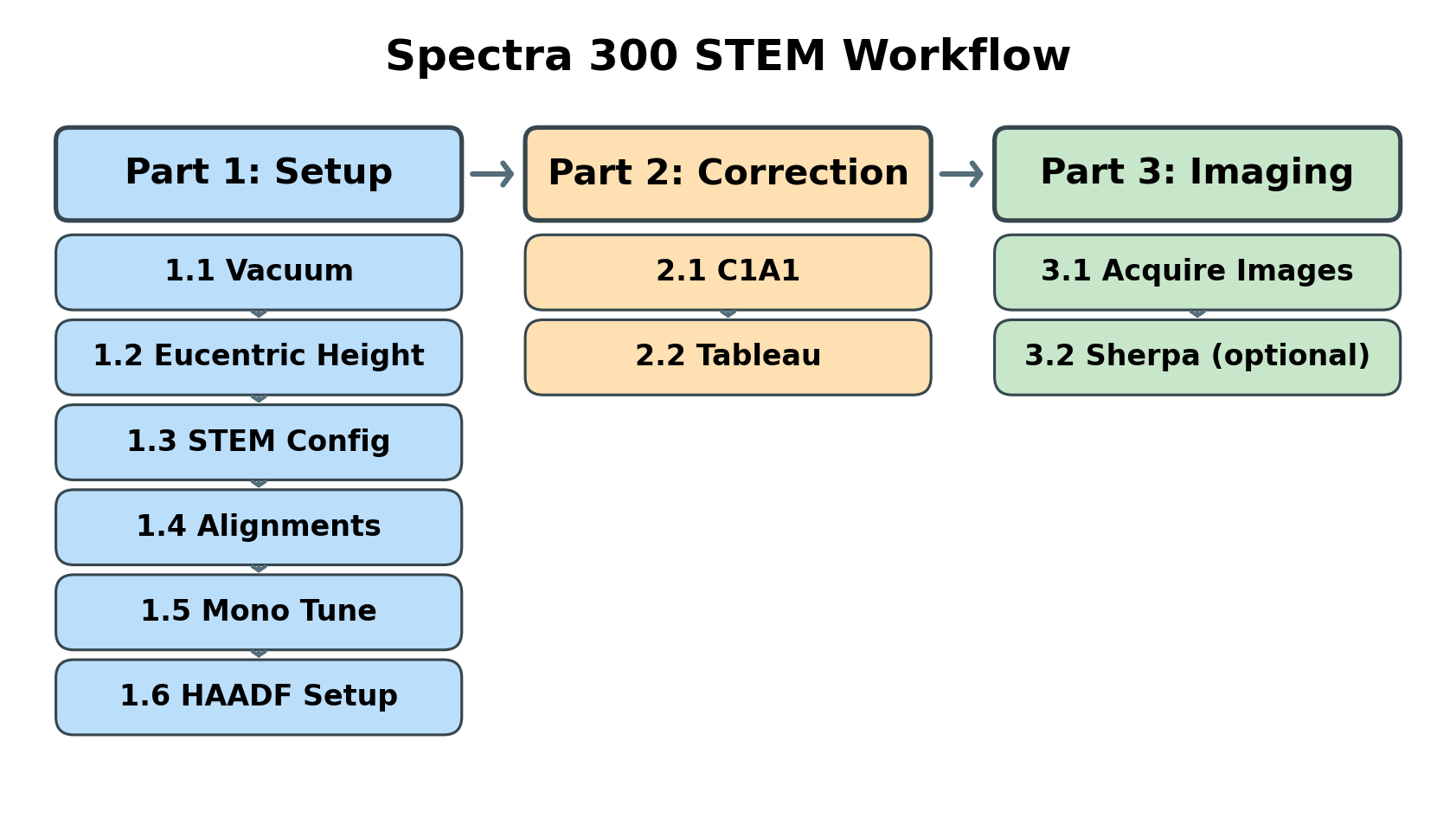

Part 1: Setup & Alignment
1.1 Vacuum check
Before imaging, verify that the vacuum system is ready and the column valves can be safely opened. Poor vacuum conditions can damage the electron source and contaminate the sample.
-
Check vacuum status
-
In
TEMUI, openSetuptab. -
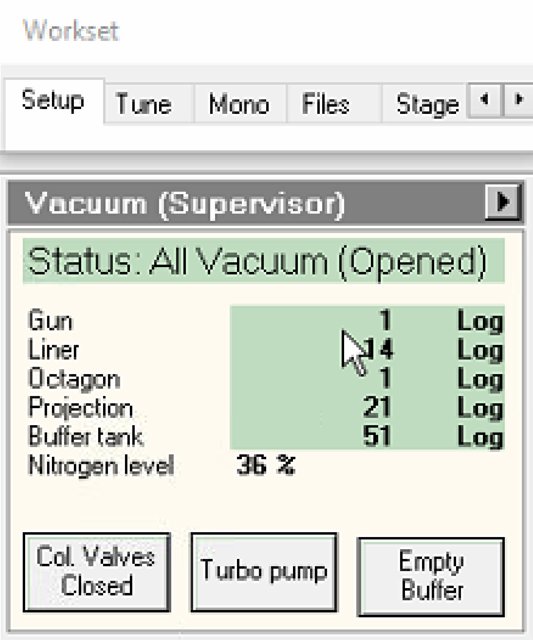
Locate vacuum status panel. Status shows “All Vacuum (Closed)” and
Col. Valves Closedbutton is yellow.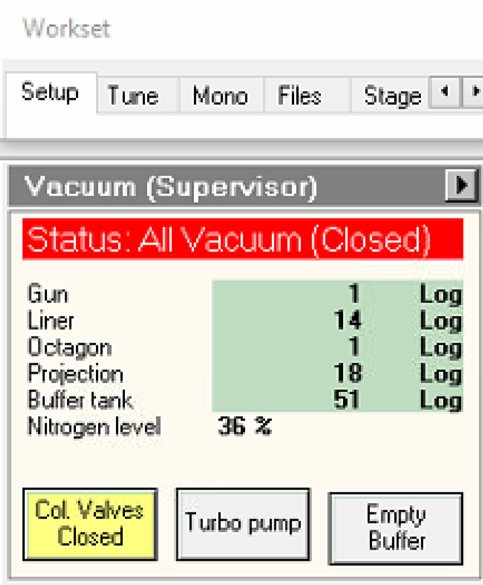
-
-
Verify vacuum pressure
-
Check vacuum pressure values on the log scale (lower = better):
Gauge Typical Log Value Notes Gun 1 Critical for source lifetime Liner 14 Column vacuum Octagon 1 Sample area Projection 19-21 Below sample Buffer tank 41-51 Empty if above 51 -
If buffer tank pressure is above 51, click
Empty Buffer.- Pumps cycle audibly.
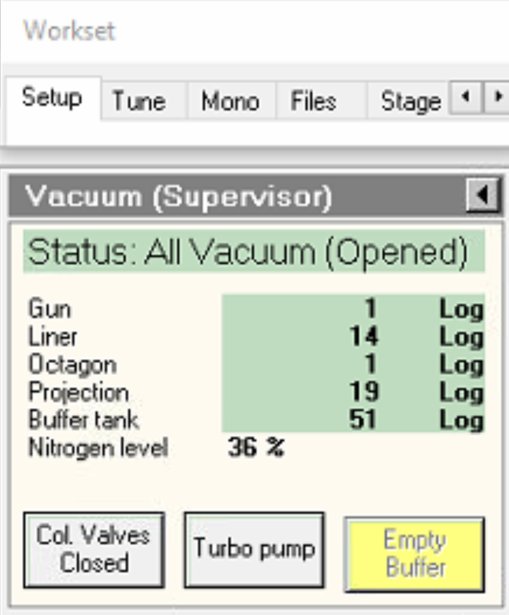
-
Wait for value to decrease before proceeding.
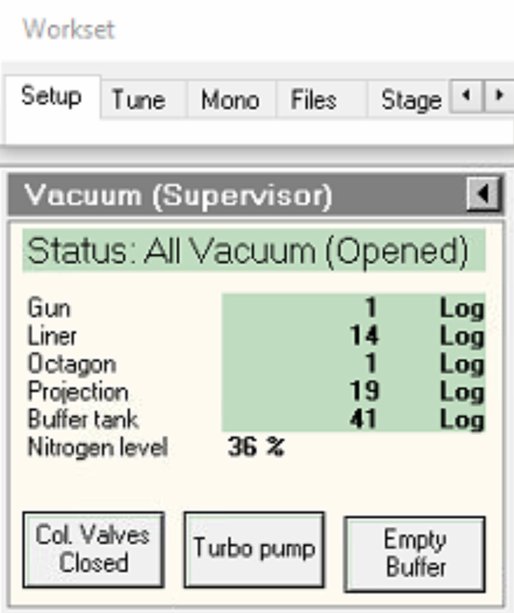
-
-
Open column valves
-
Click
Col Valves Closedbutton.- Status changes to “All Vacuum (Opened)”.
NOTE: The system only allows opening if vacuum levels are acceptable. Once opened, the electron beam path is clear from gun to sample.
-
1.1 Check convergence angle
For this guide, we use a convergence angle of 30.0 mrad.
- In
TEMUI, navigate to theTunetab, then selectAperture. - Set Condenser 1, Condenser 2, and Condenser 3 to
2000,70, and1000.
NOTE: These aperture values determine the convergence angle. For example, setting Condenser 2 to
50instead of70gives a convergence angle of 21.5 mrad. A smaller aperture restricts the beam to a narrower range of incident angles, blocking higher-angle electrons.
1.2 Find eucentric height
Complete eucentric height alignment after loading each sample and before imaging. Do not skip this step. At eucentric height, the sample remains stationary when tilted. This is essential for accurate imaging and tomography. The ronchigram “blow-up” method provides a quick way to find this position.
-
View ronchigram
-
Verify the
Diffractionbutton is pressed on the hand panel with the red light turned on.NOTE: The ronchigram is the diffraction pattern formed when the convergent probe is stationary. When defocused, it contains shadow images of sample features, making structure visible during z-height adjustment.
-
In
TEMUI, view the ronchigram in the main display.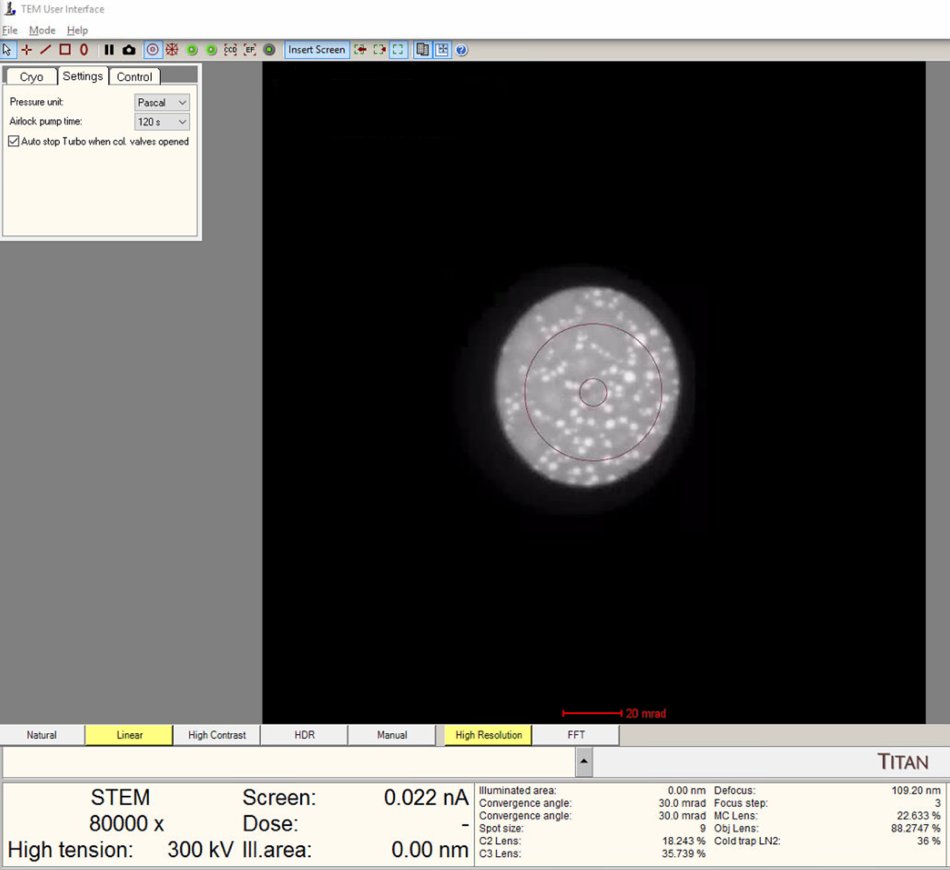
-
Position probe on a sample region that scatters electrons (not over a hole or vacuum).
-
-
Adjust z-axis to find blow-up point
-
Lower magnification to 5,000x. A wider field of view makes ronchigram changes easier to observe.
-
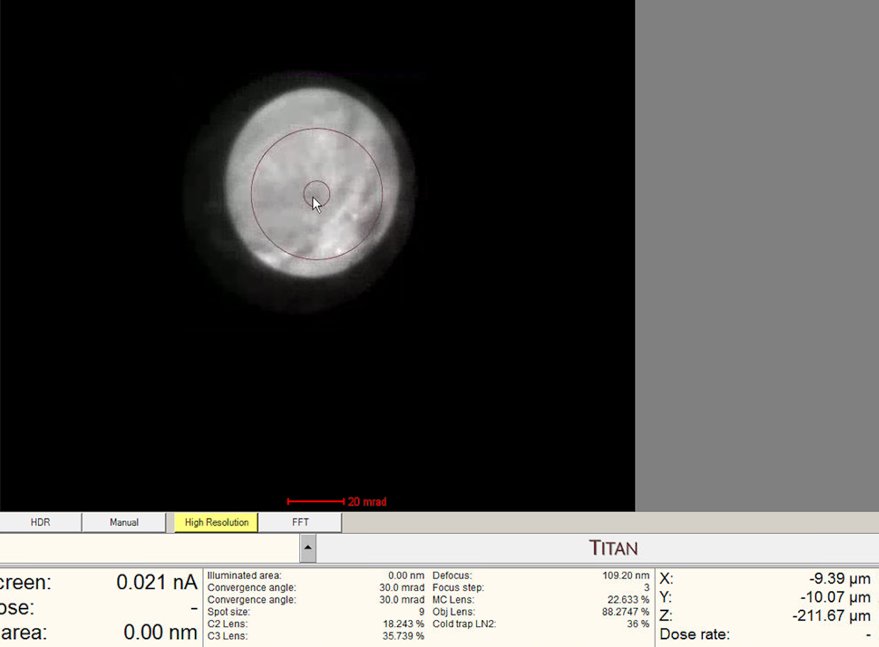
Find a region where there is a sharp contrast at a boundary, as shown in the following image.
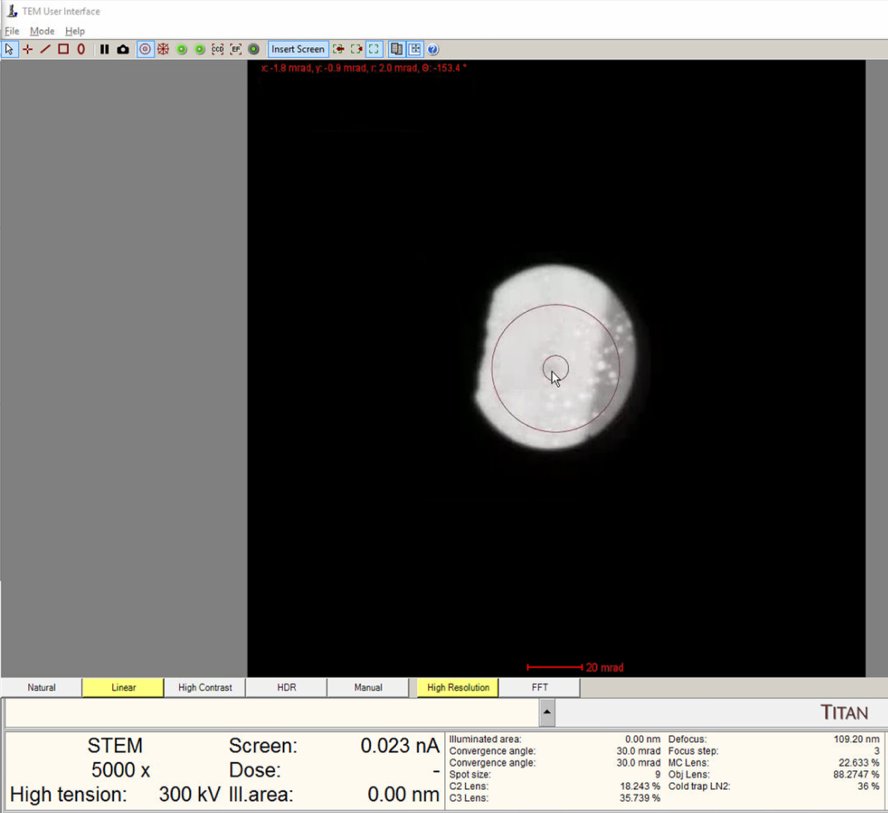
-
Use z-axis buttons on hand panel to move stage up or down.
- Buttons are pressure sensitive: press harder for faster movement.
- Start with gentle presses for fine control.
- Notice that as you adjust the z-axis, the ROI also shifts. Use the joystick to remain on the sharp contrast boundary region.
-
Watch the ronchigram while adjusting z-height. The pattern “zooms” in or out as the sample moves through focus.
-
Continue adjusting. The ronchigram expands when approaching eucentric height, also referred to as the “blow-up” point.
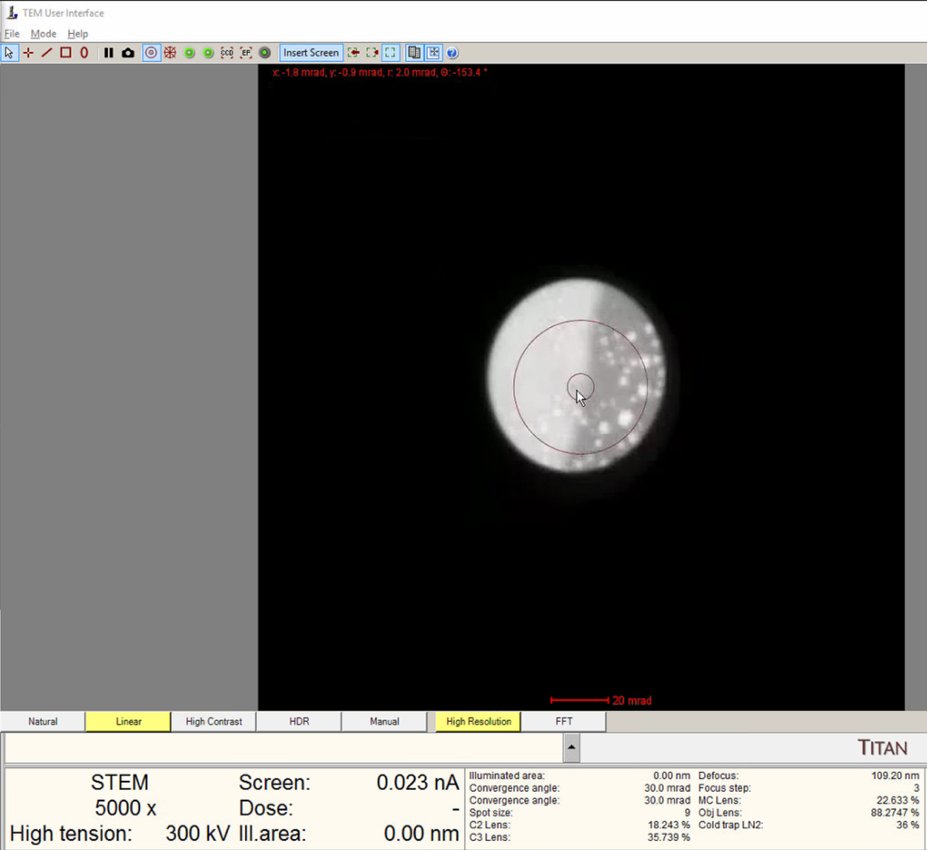
-
Find the “blow-up” point where the ronchigram appears infinitely magnified: shadow image features expand until they fill the entire display.
- If the ronchigram starts shrinking again, reverse direction.
-
1.3 STEM mode configuration
Before performing alignments, configure the STEM imaging parameters and verify detector settings.
-
Enable descan
-
In
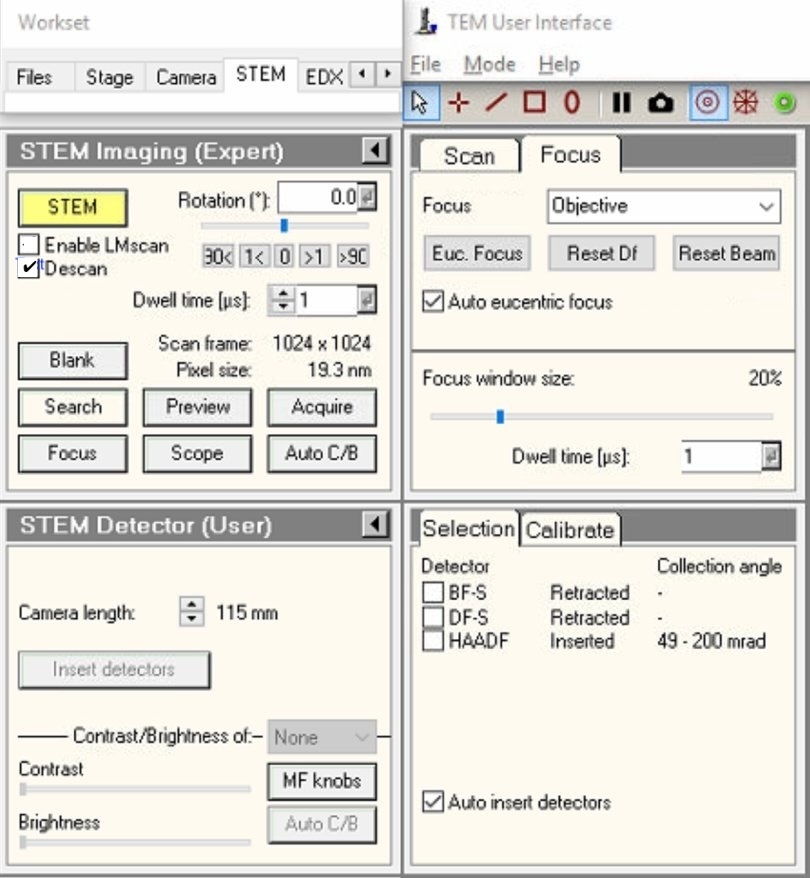
TEMUI, locate theSTEM Imaging (Expert)panel. -
Enable
Descancheckbox.NOTE: Descan compensates for beam movement during scanning, keeping the diffraction pattern stationary on the detector.
-
-
Verify HAADF is retracted
-
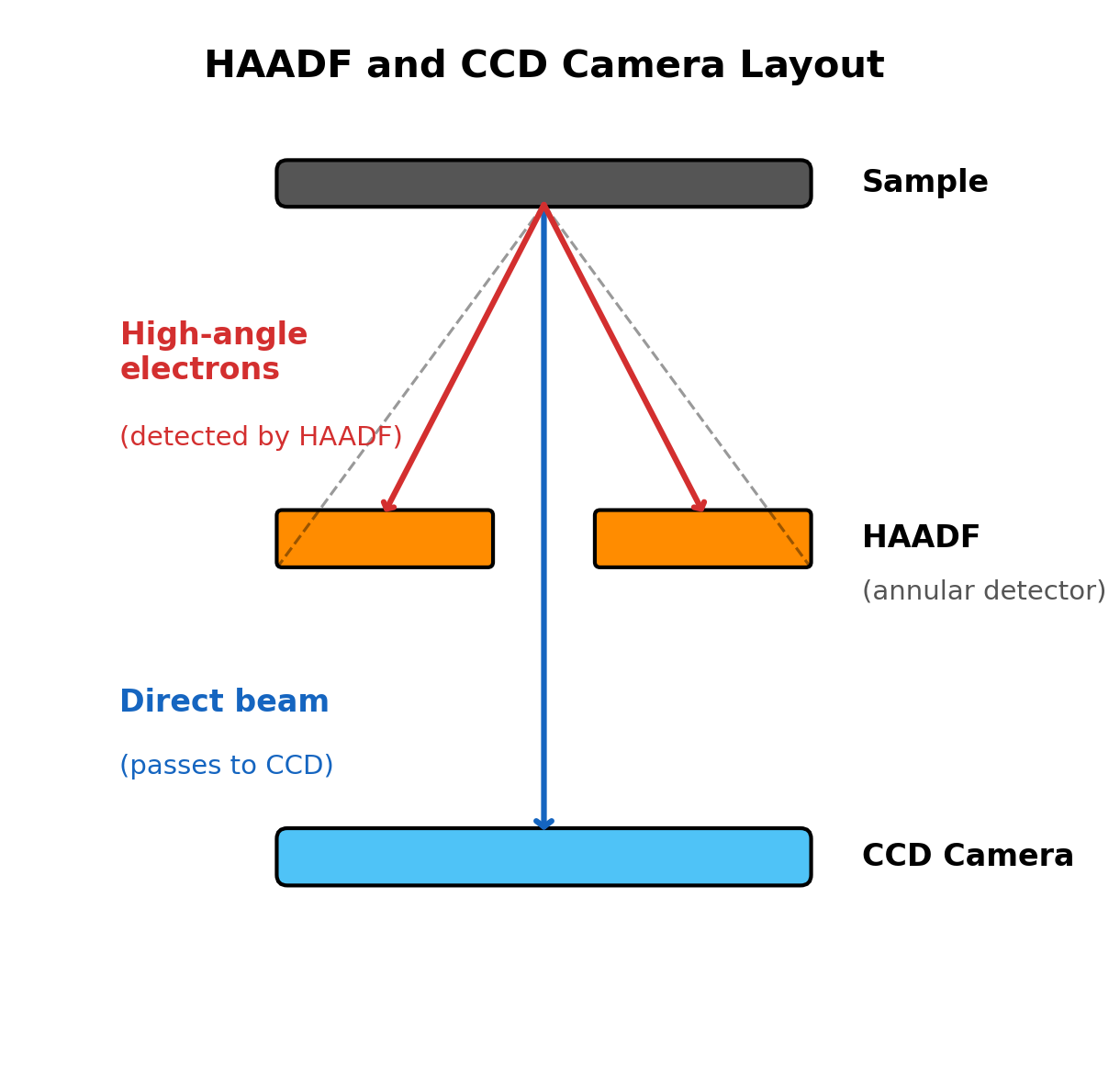
Locate the
Selectionpanel. -
Verify detector states:
- BF-S (Bright Field): Retracted
- DF-S (Dark Field): Retracted
- HAADF: Retracted
-
Toggle
HAADFcheckbox on then off to confirm retracted state.NOTE: HAADF must be retracted during ronchigram alignment. The HAADF is a ring-shaped detector with a central hole. If inserted, high-angle electrons hit the ring instead of the camera below, blocking part of the ronchigram.
-
-
Set detector layout in Velox
-
Open
Veloxacquisition software. -
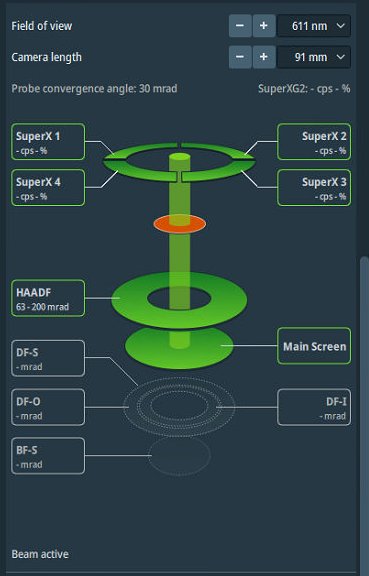
Open detector layout display.
-
Set camera length to 91 mm.
NOTE: Camera length determines detector collection angles. The layout display shows the angular ranges for each detector.
-
-
Configure beam settings
-
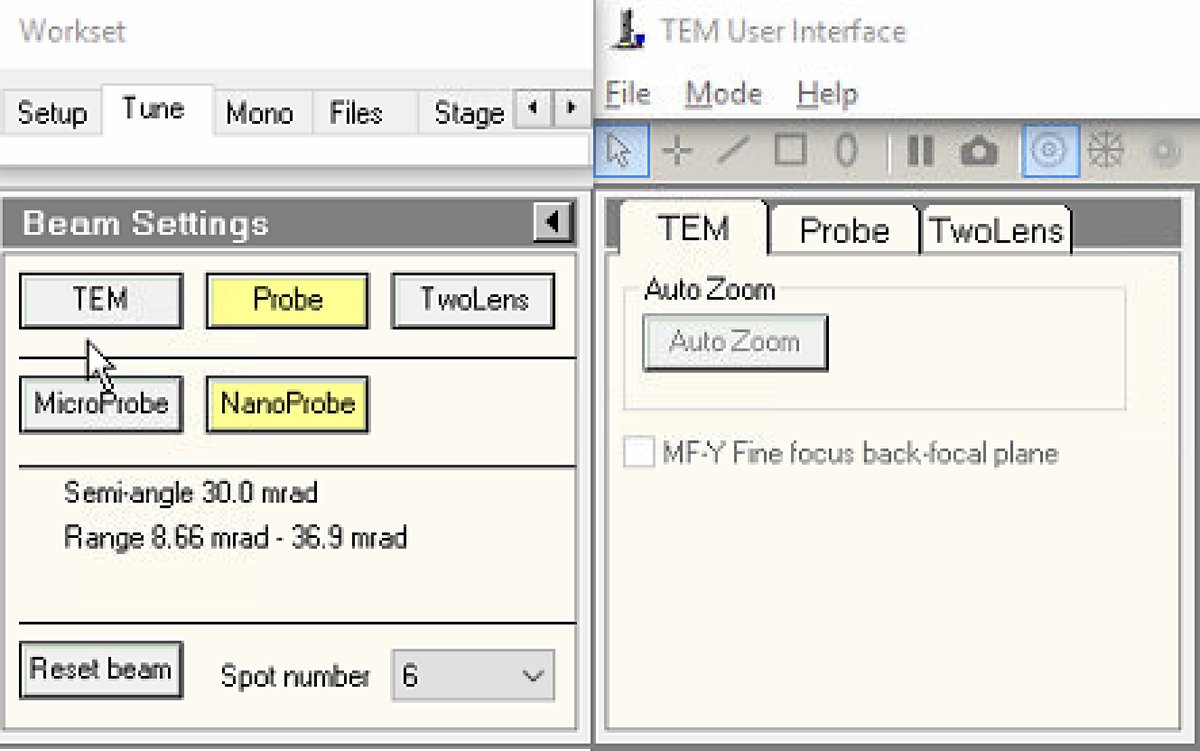
In
TEMUI, go to theTunetab and locate theBeam Settingspanel. -
Select
Probemode.- Button is highlighted yellow when active.
-
Select
NanoProbemode. -
Set spot number to 6.
NOTE: NanoProbe provides a smaller, more coherent probe than MicroProbe. Lower spot numbers produce smaller probes with lower current; higher numbers produce larger probes with more current.
TODO: Verify spot number convention for Spectra 300. On some Thermo Fisher systems, it is the opposite (spot 1 = most current).
-
1.4 Direct alignments
The basic alignments center the electron beam and align it through the optical column. Proper alignment is essential for optimal resolution and probe symmetry.
-
Set magnification and open Direct Alignments
-
Set magnification to 200-300kX using the magnification knob.
-
In
TEMUI, navigate toTunetab, thenDirect Alignments. This panel provides access to all fundamental beam alignment procedures. -
Select
Diffraction Shift and Focus alignmentto begin.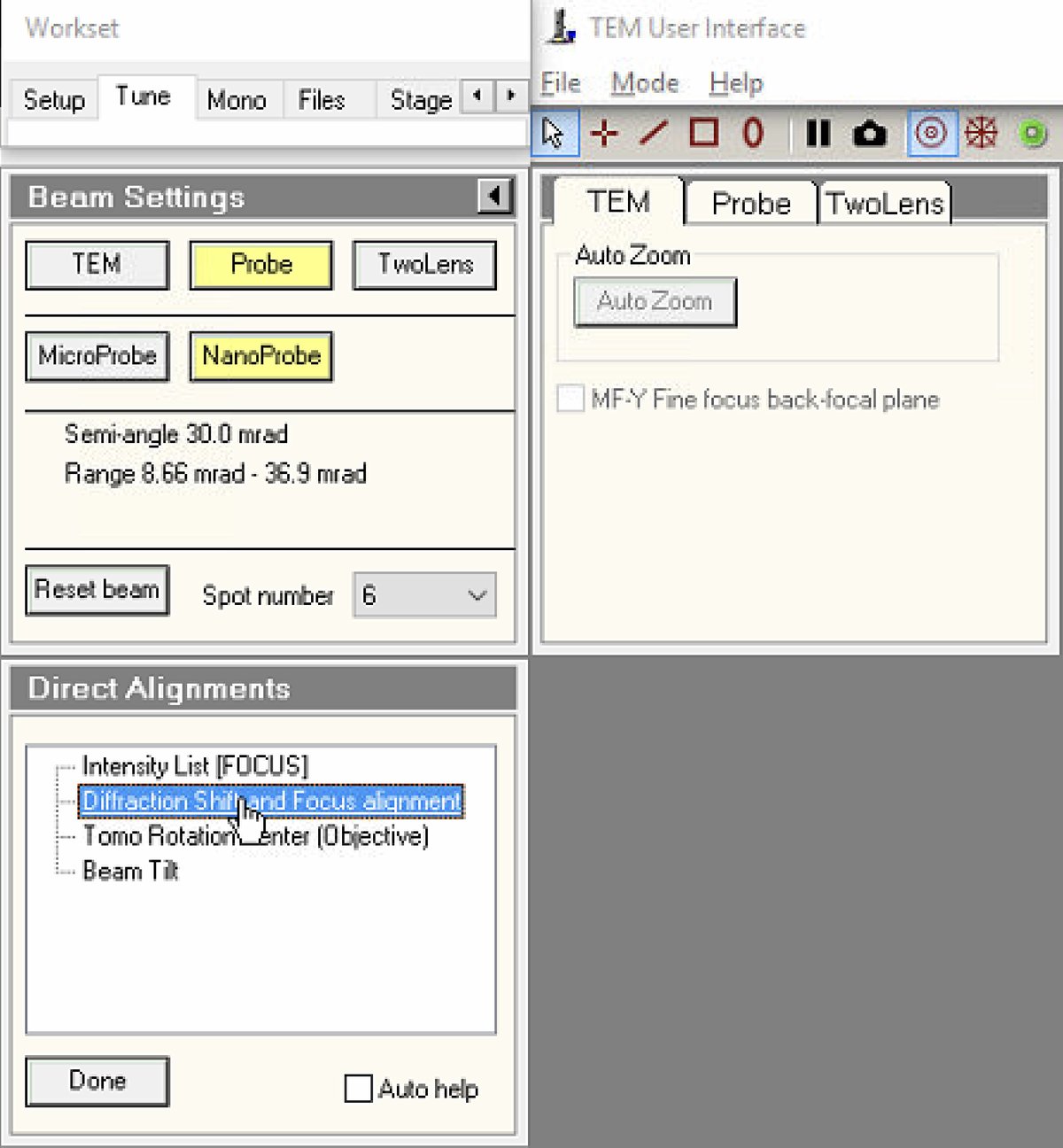
-
-
Center ronchigram
-
Observe the ronchigram position on the display. If the ronchigram is shifted from center, use the
mulXYknobs to bring it back.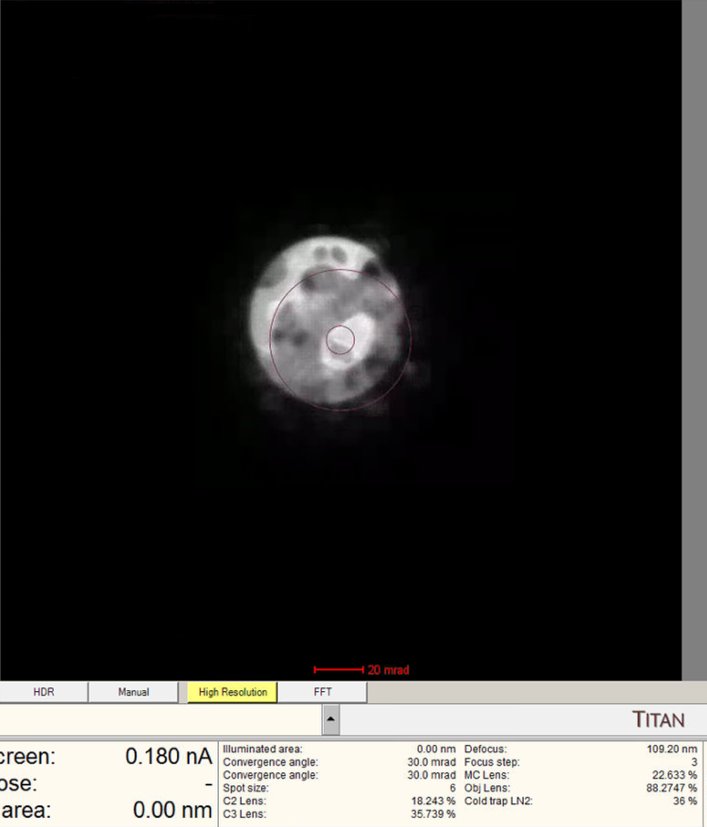
-
The
mulXYknobs now control diffraction shift. Adjust until the ronchigram is centered.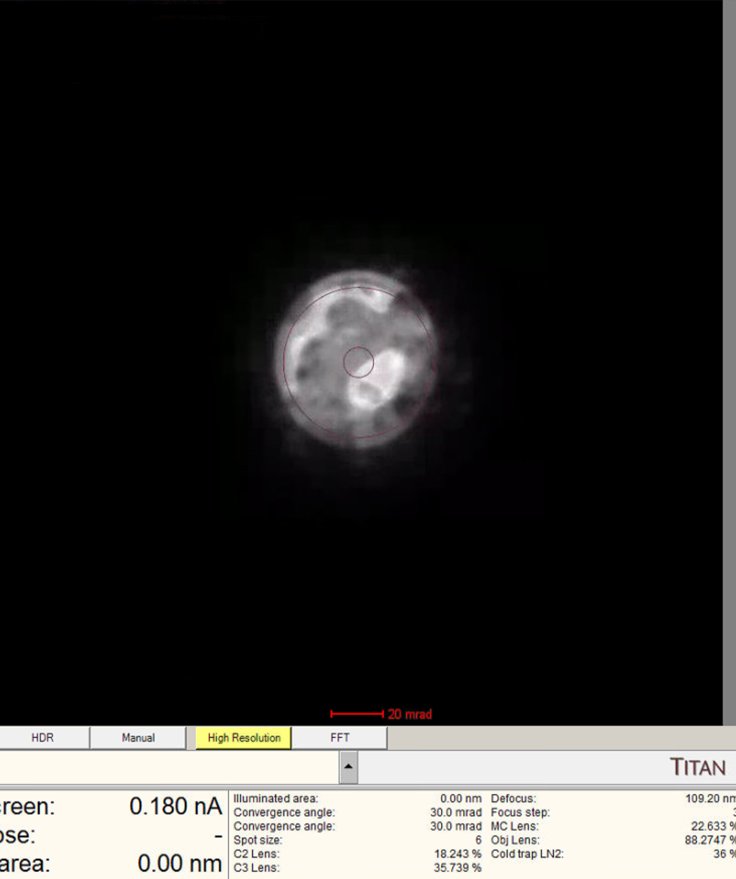
-
-
Reset STEM AutoTuning
-
In the quick dropdown menu, select
STEM AutoTuning. This panel stores automatic alignment adjustments from previous sessions. -
Click
Resetunder Settings to clear stored values. This establishes a known baseline. Previous user adjustments persist and interfere with fresh alignments if not reset.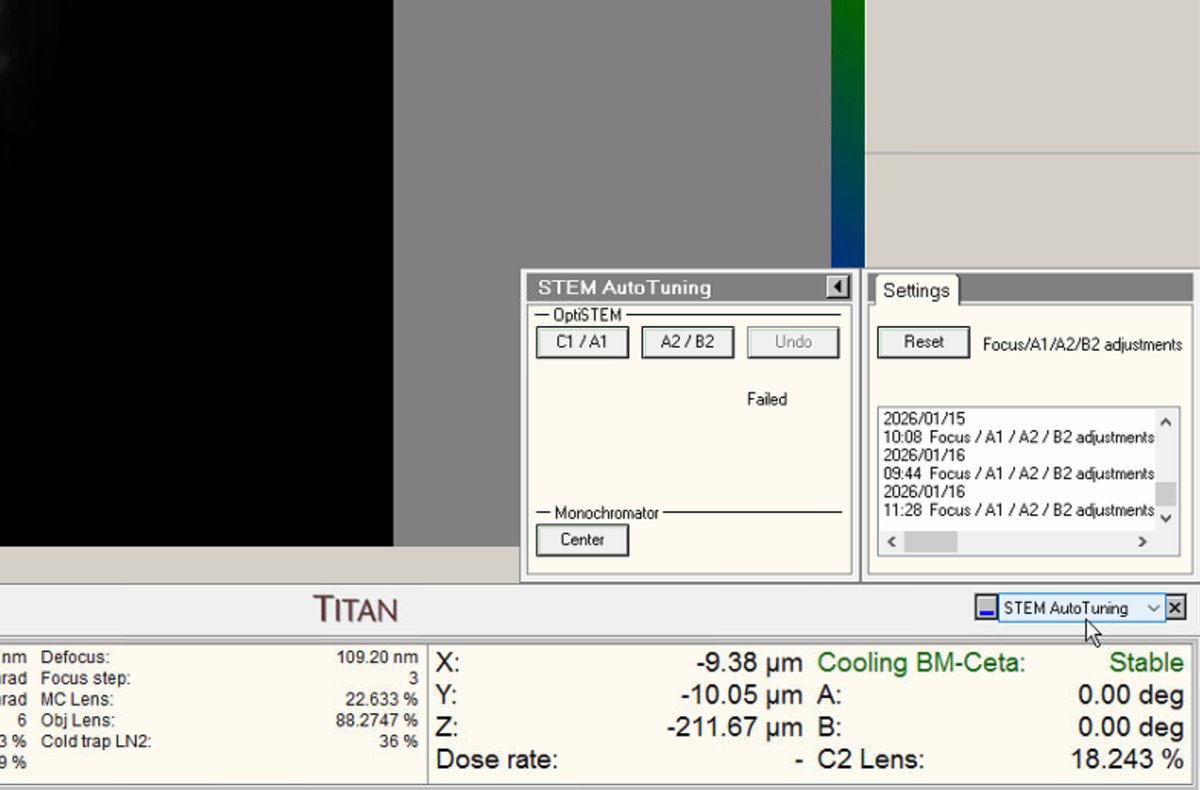
-
-
Switch to probe image mode
-
Press the
Diffractionbutton on the hand panel to enter probe image mode (STEM scanning). The red light should turn off once pressed.No beam found? In the following step you will click
Beam Shiftand adjust themulXYknobs. Watch the screen current: it changes from 0.000 nA to 0.001 nA, etc. This means you are shifting the beam position near the screen. You will see dim light coming from the edges. Press theFineandCoarsebuttons to adjust the sensitivity of themulXYknobs.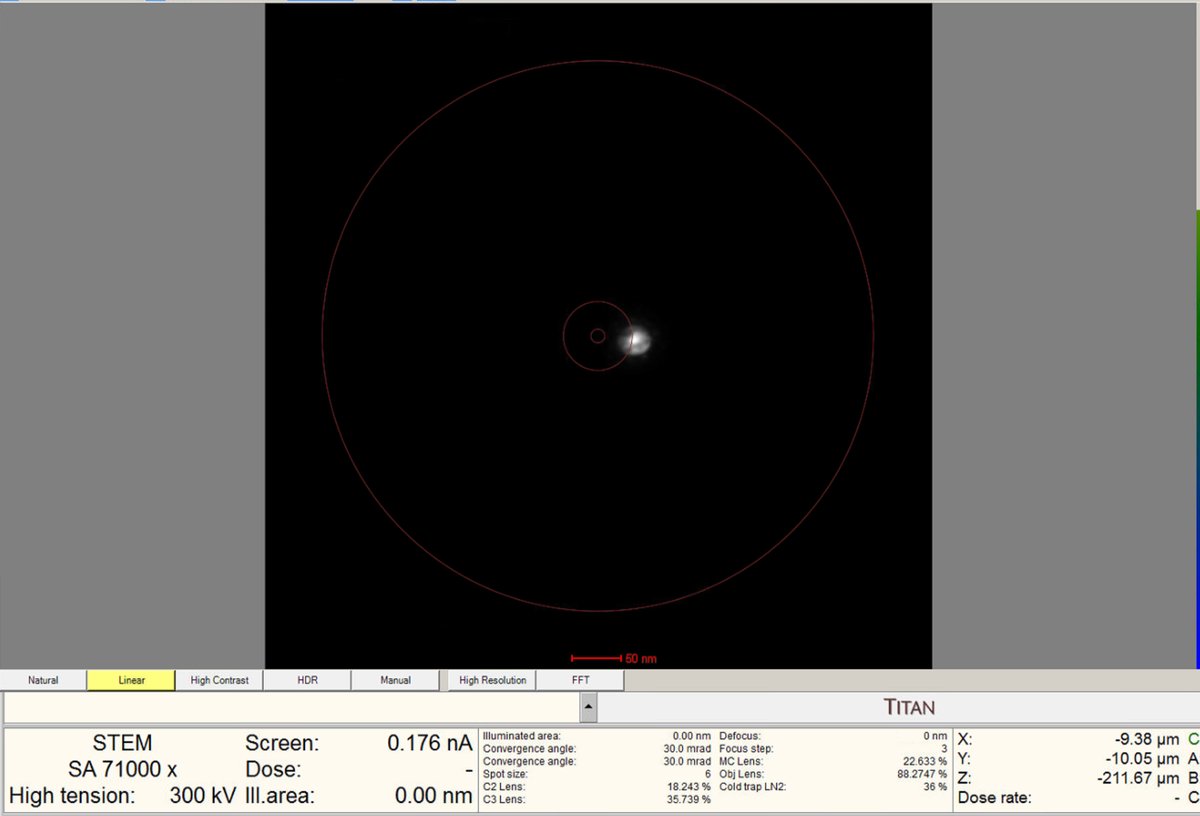
Diffraction mode vs. Probe image mode
Mode Probe Display Diffraction mode Stationary Ronchigram - diffraction pattern from convergent probe Probe image mode Scanning STEM image - probe scans to build up image pixel by pixel The
Diffractionbutton on the hand panel toggles between these two modes.
-
-
Align beam shift
-
Click on
Beam shiftin the Direct Alignments panel. ThemulXYknobs now control alignment beam shift. -
Use the
mulXYknobs to center the beam on the screen. The beam responds smoothly to knob movements. If the beam moves too quickly, pressFineon the hand panel to reduce sensitivity. -
Important: If the beam is lost after clicking beam shift, reduce magnification until the beam is visible, center using the
mulXYknobs. -
Click
Doneonce the beam is properly centered.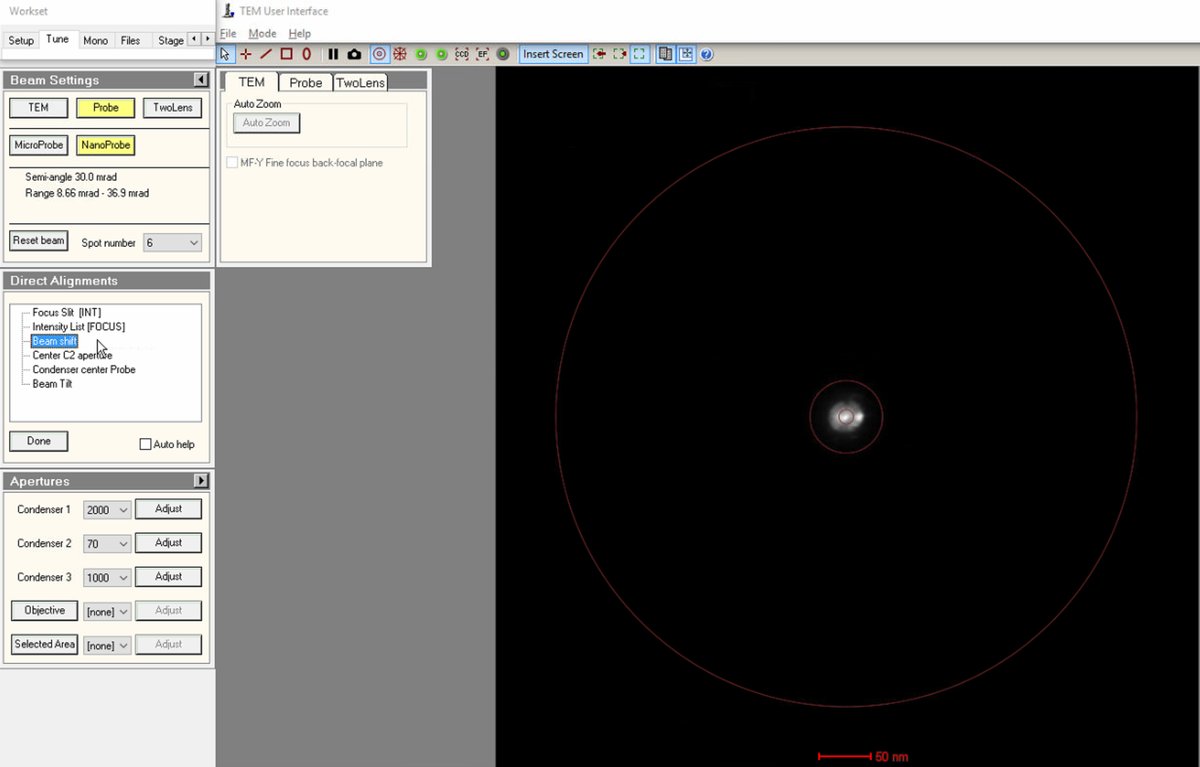
-
-
Center C2 aperture
-
Select
Center C2 aperturefrom the alignment options. The system oscillates the C2 lens, causing the beam to expand and contract rhythmically. -
Watch the beam movement carefully. The beam pulses in and out. The goal is to make this expansion/contraction perfectly concentric (no lateral movement).
-
Use the
mulXYknobs to adjust the aperture position. -
Click
Donewhen the movement is concentric.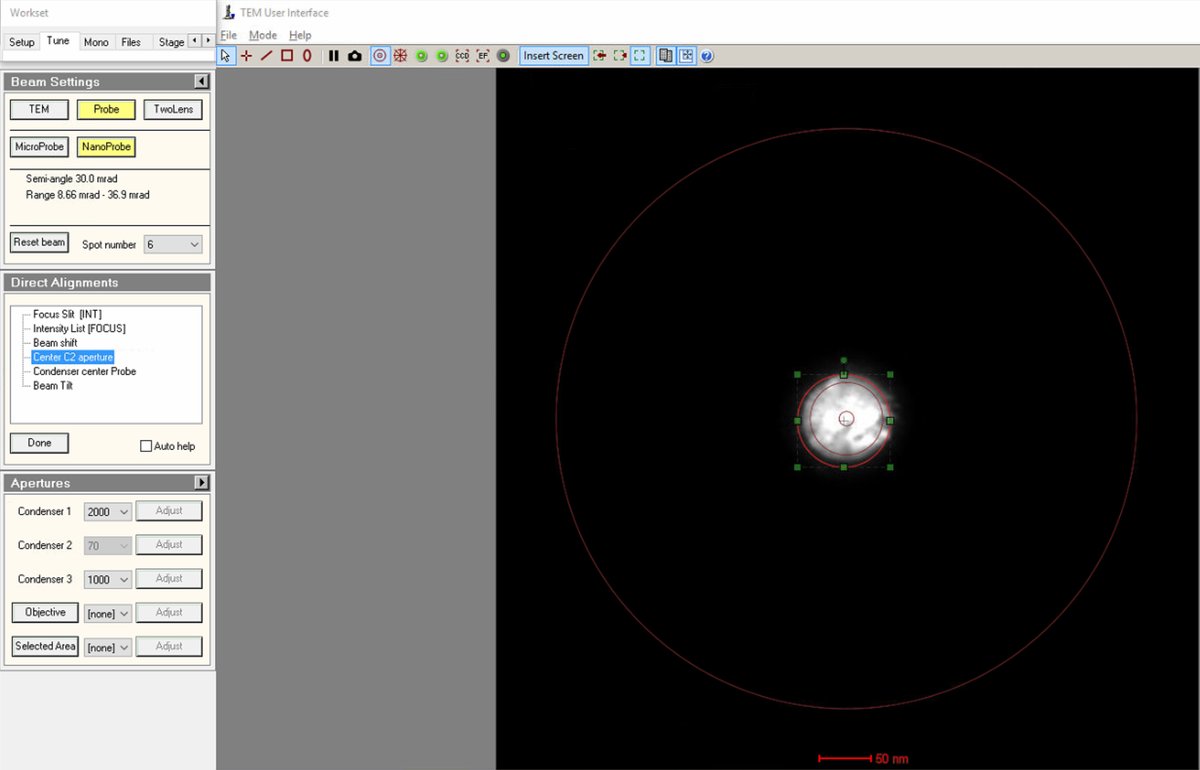
-
-
Align beam tilt
-
Select
Beam Tiltfrom the alignment options. This alignment minimizes the lateral shift of the beam when tilting. -
Use the
mulXYknobs to minimize lateral movement of the beam. When properly aligned, the beam changes angle without shifting position. -
Reduce the lateral x and y movements as much as possible using the
mulXYknobs. -
Click
Doneonce the lateral movement is minimized.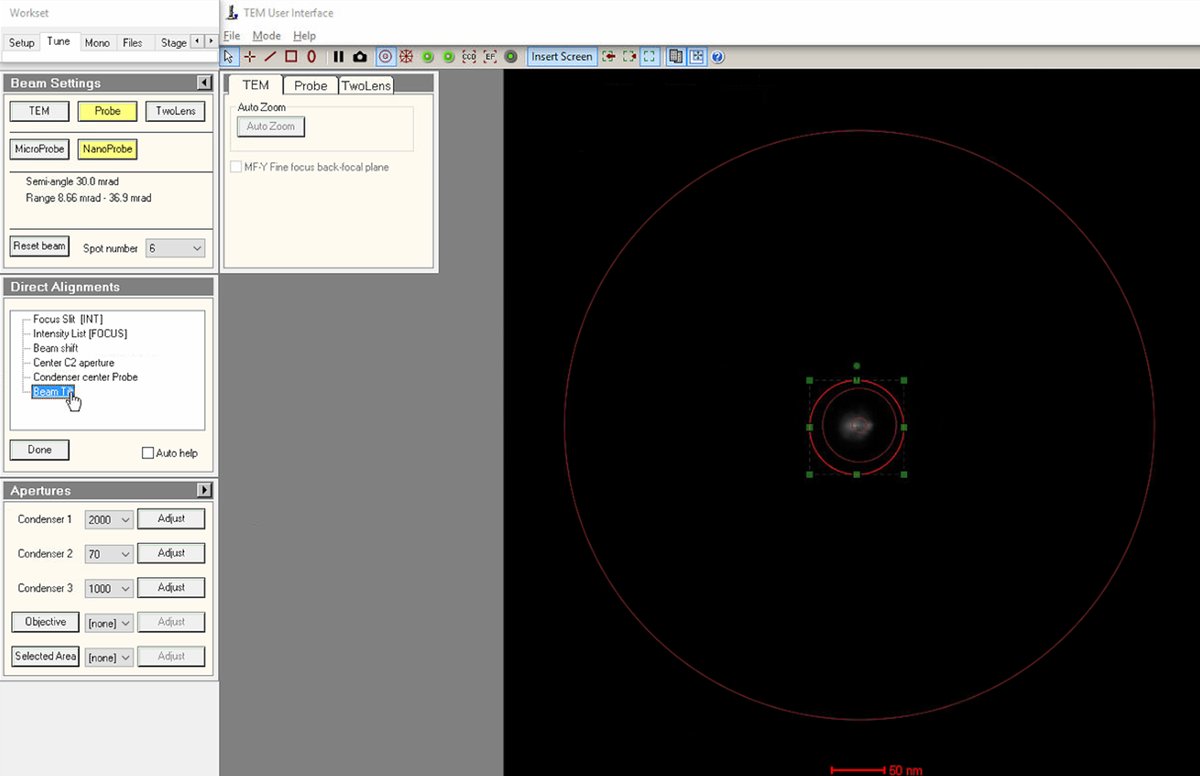
-
-
Verify final diffraction shift
-
Press the
Diffractionbutton on the hand panel to switch back to diffraction mode (view the ronchigram). The red light should now be on. -
Return to
Diffraction Shift and Focus alignmentfor a final centering check. -
Use
mulXYto center the ronchigram precisely on the display. Centering confirms the beam is on the optical axis. -
Click
Doneto complete the direct alignments.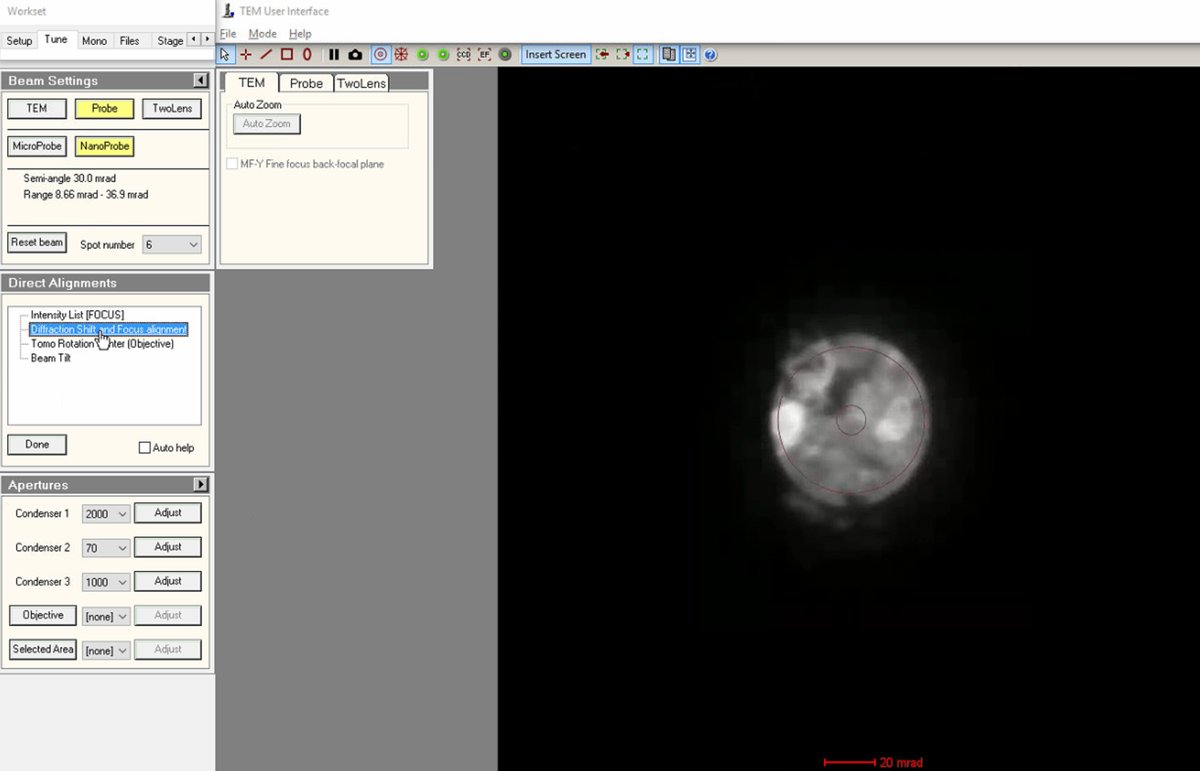
-
Note: Mode changes (diffraction ↔ probe image, TEM ↔ STEM) disable descan. Re-enable descan after each mode switch. If the image looks distorted, verify
Descanis enabled inSTEM Imaging (Expert).
1.5 Monochromator tune
Before proceeding to probe correction, check that the monochromator is properly aligned and not partially blocking the beam. The monochromator selects a narrow energy spread from the electron source, improving resolution but reducing beam current.
-
Open Monochromator Tune
-
In
TEMUI, go to theMonotab and locate theMonochromator Tune (Expert)panel. This panel provides controls for adjusting the monochromator position and focus. -
Click on both
ShiftandFocusbuttons to enable adjustment mode. At this point, the intensity knob controls monochromator focus and themulXYknobs control monochromator shift.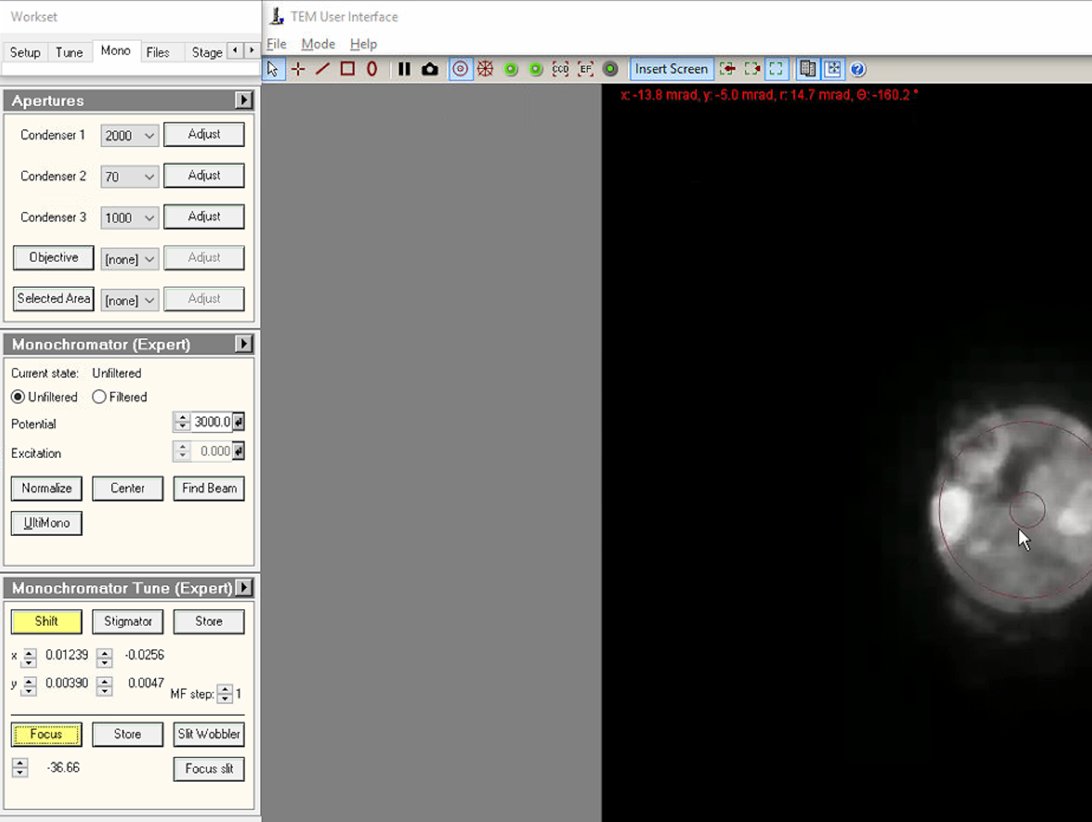
-
-
Adjust focus
-
Adjust the intensity knob to bring the Focus value close to 0. As the monofocus approaches zero, the screen current increases because more electrons pass through the monochromator slit.
-
While adjusting Focus toward 0, also adjust the
mulXYknobs to ensure the beam isn’t blocked. The screen current inTEMUIshould be above 15 nA when focus is near zero.No beam visible? Click
Linearin the detector settings to switch from log to linear display mode. If the beam is still missing, return to the initial focus value and slowly bring it back toward 0 while adjustingmulXY, as in the Beam Shift alignment. -
Watch the current readout while adjusting.
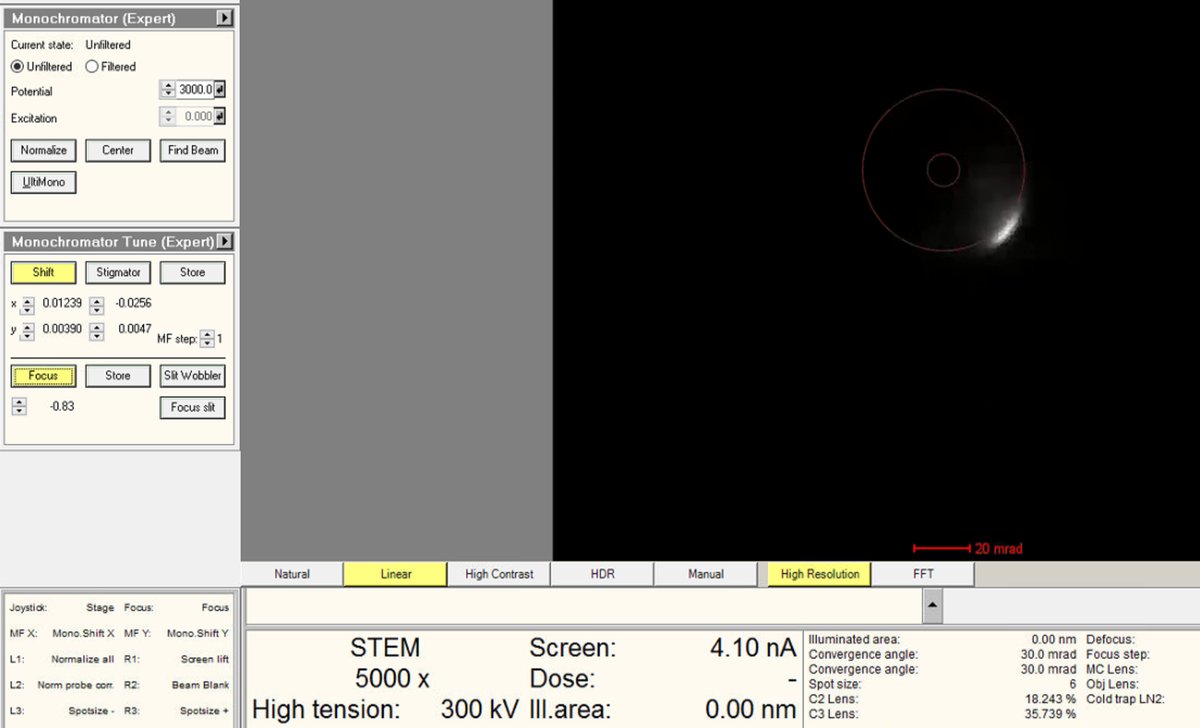
-
-
Center and adjust current
-
Adjust the
mulXYknobs to center the beam through the monochromator.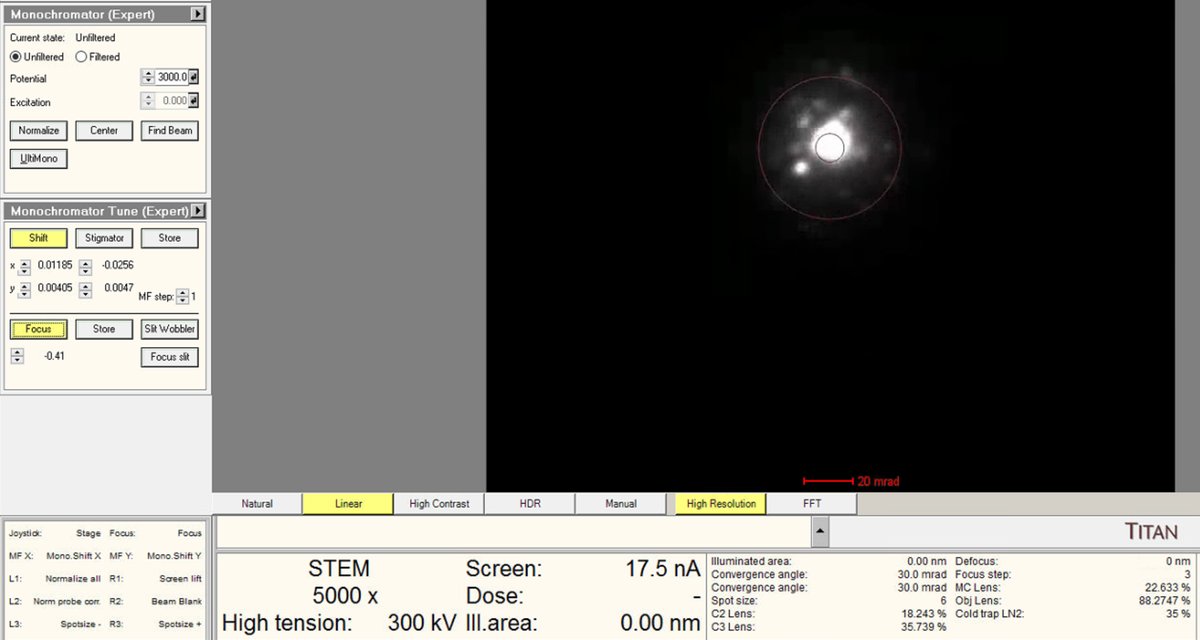
-
Use the intensity knob to achieve the target beam current (~0.150 nA for high-resolution STEM).
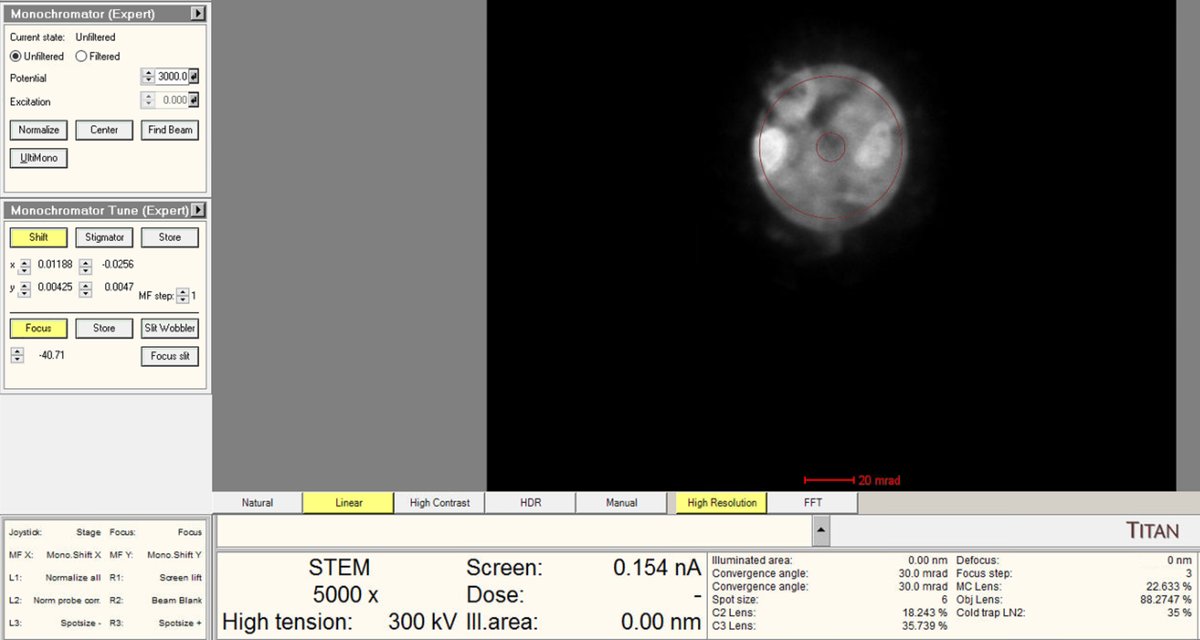
-
-
Deselect Shift and Focus
- Click
Shiftagain to deselect both buttons (they toggle together). - This returns the intensity knob and
mulXYknobs to their normal functions. Verify the current readout shows the target value before proceeding.
- Click
-
Re-verify eucentric height
- Use the z-axis controls to return to the “blow-up” point (eucentric height). Monochromator adjustments affect focus; re-verify eucentric height.
1.6 HAADF imaging setup
Before running aberration correction, set up HAADF (High-Angle Annular Dark Field) imaging to view the sample and find a suitable region. HAADF provides Z-contrast imaging where heavier atoms appear brighter.
-
Switch to HAADF
-
In the
Veloxacquisition software, clickSTEMto enter STEM mode, -
Click
HAADF. This automatically inserts the HAADF detector. -
Verify in
TEMUIthat the HAADF detector shows “Inserted” status with the correct collection angle (63-200 mrad).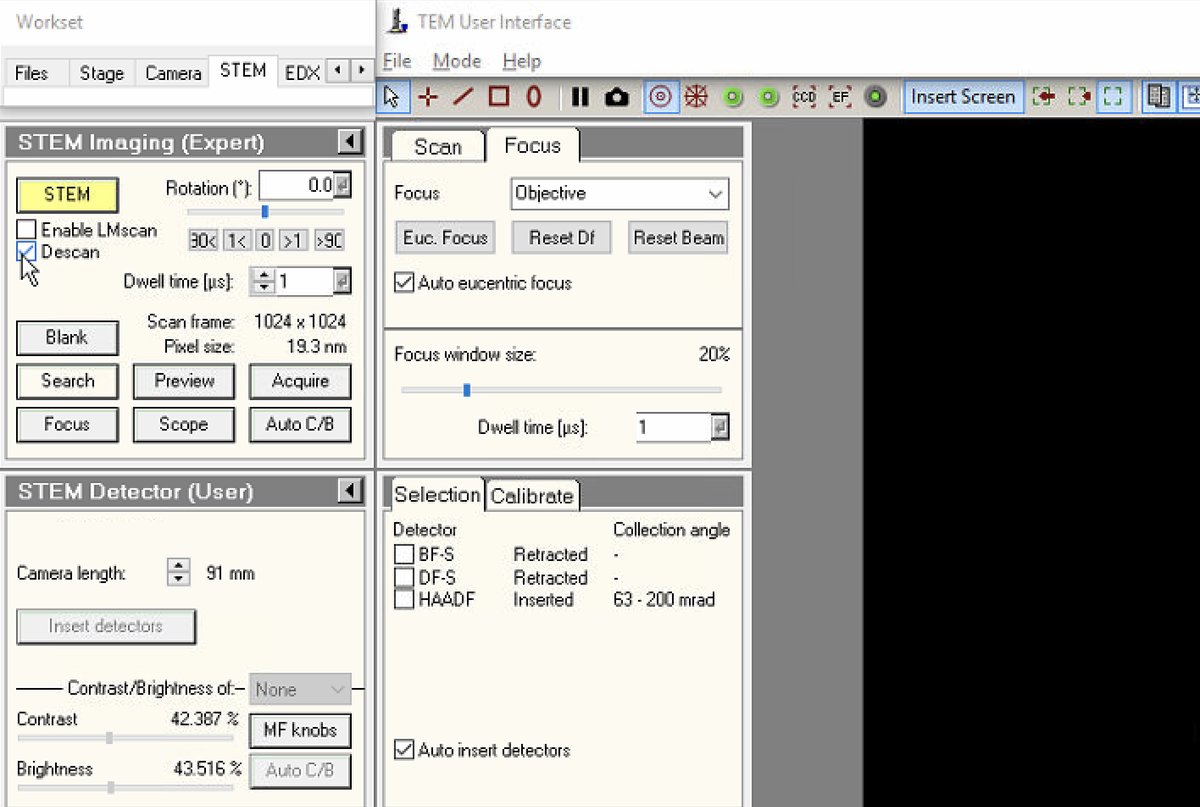
-
-
Verify Descan is enabled
- In
TEMUI, go toSTEM Imaging (Expert)and verifyDescanis enabled. Mode changes disable descan; re-enable after switching modes.
- In
-
Start live scanning
-
Click the play button in
Veloxto start live scanning. -
The image is saturated (all white) initially. Detector signal adjustment follows in the next step.
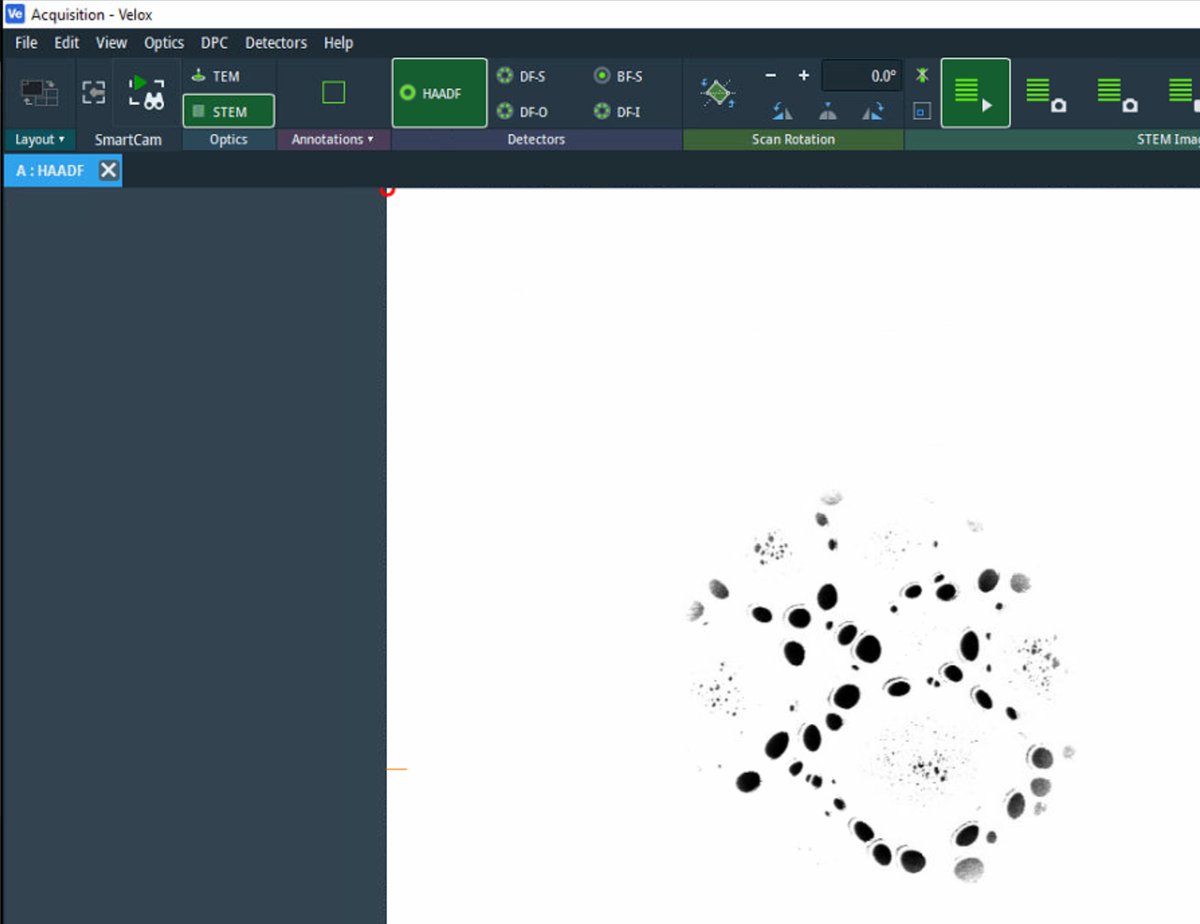
-
-
Adjust detector signal
-
In
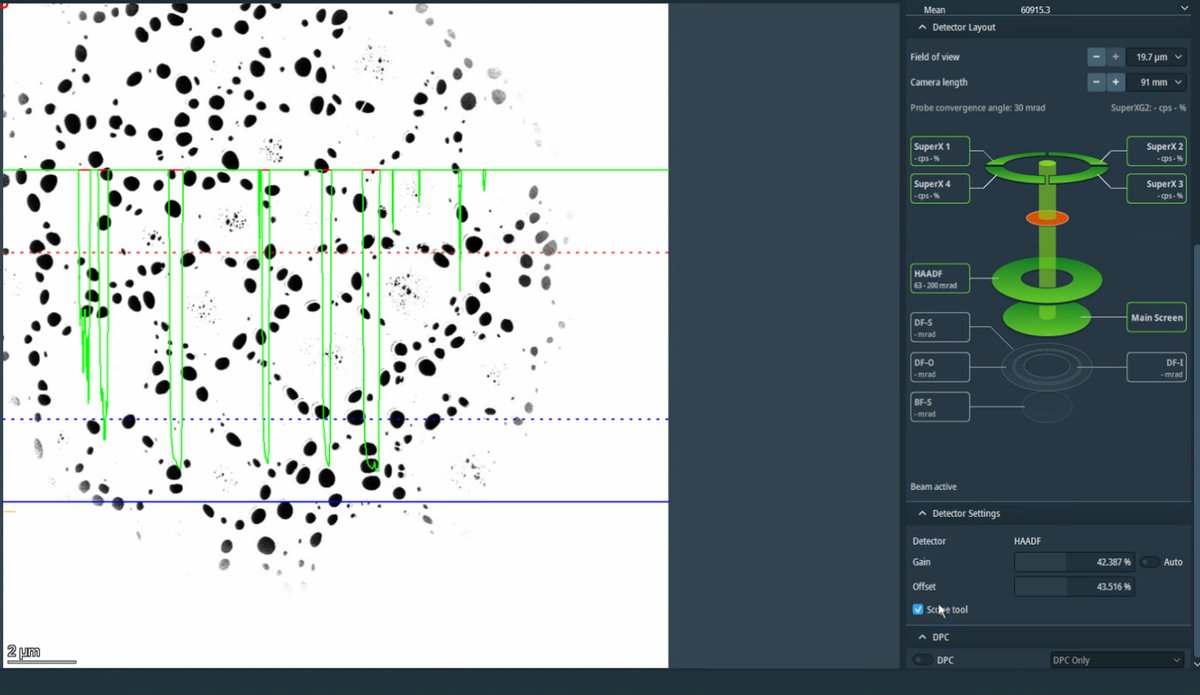
Velox, clickScope toolto enable signal adjustment. -
Adjust
GainandOffsetso the signal does not go above the dotted red lines.Signal math: Display = (Gain × Signal) + Offset
- Gain (= Contrast): Multiplier that stretches the signal. 100% = no change, 200% = double the contrast
- Offset (= Bias/Brightness): Shifts the baseline as a percentage of the display range
-
If the signal is clipping at zero (bottom of display), increase
Offsetto shift the signal up.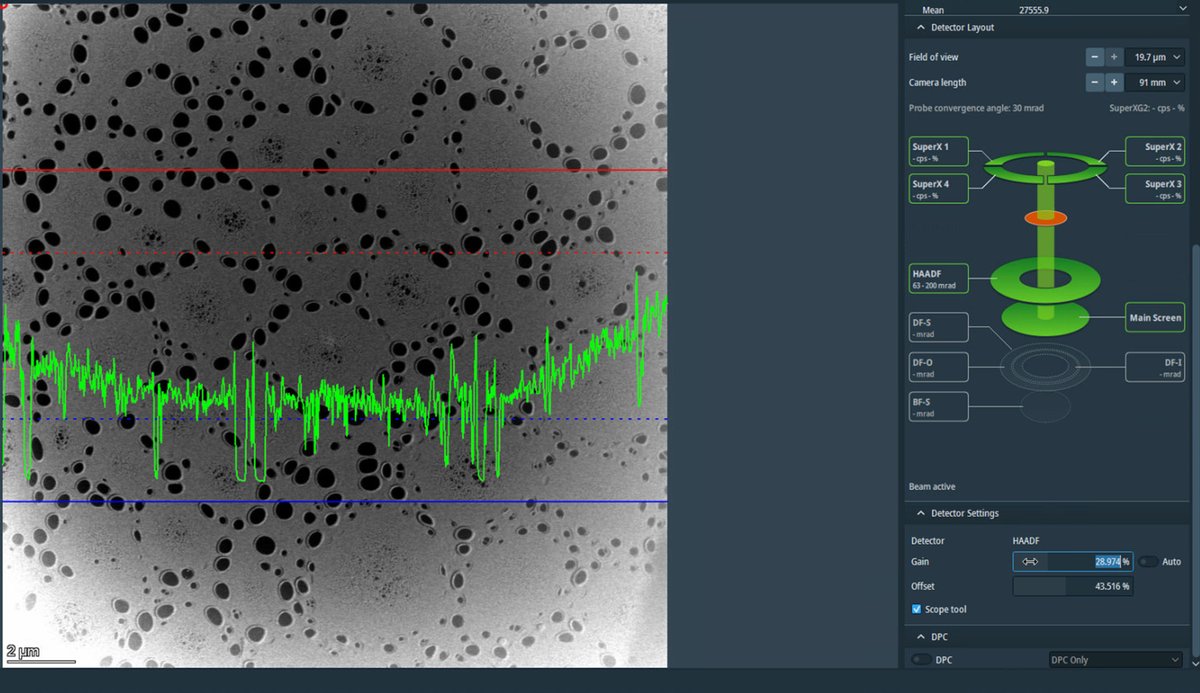
-
Adjust the magnification to ~20,000x using the magnification knob. Once adjusted, uncheck
Scope toolto turn it off.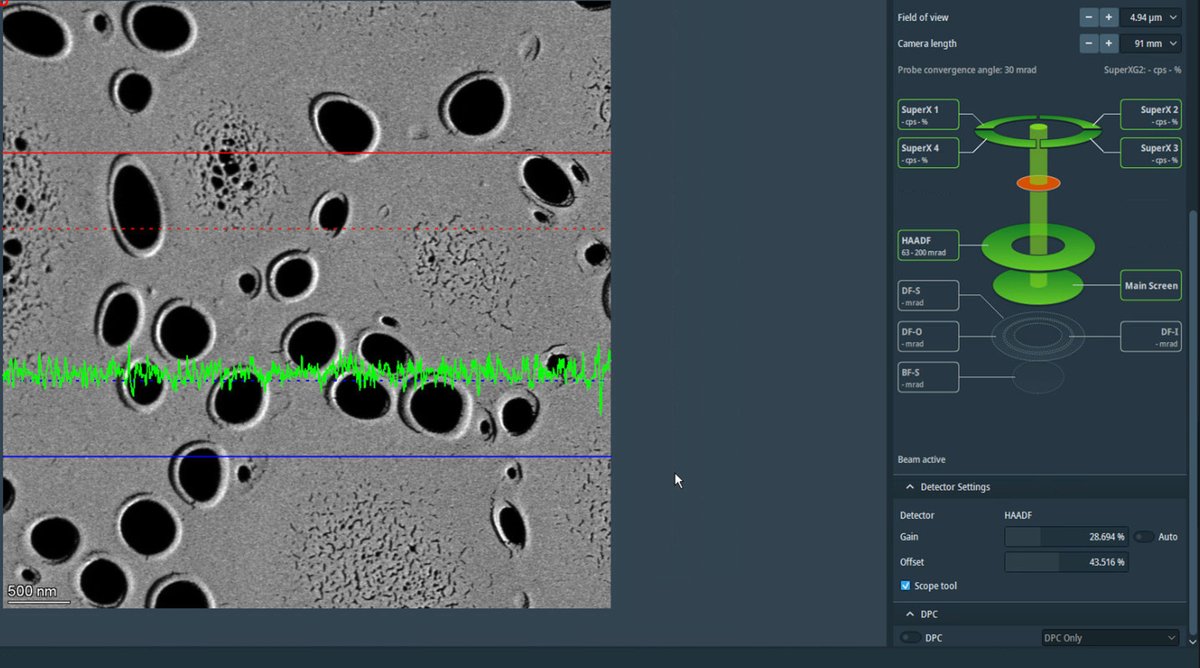
-
-
Find sample boundary
-
Reduce magnification to ~10,000x and navigate with the joystick to find a suitable region.
-
Locate a boundary region with particles at the edge of a support film, with vacuum visible.
-
This type of region provides excellent contrast for aberration correction.
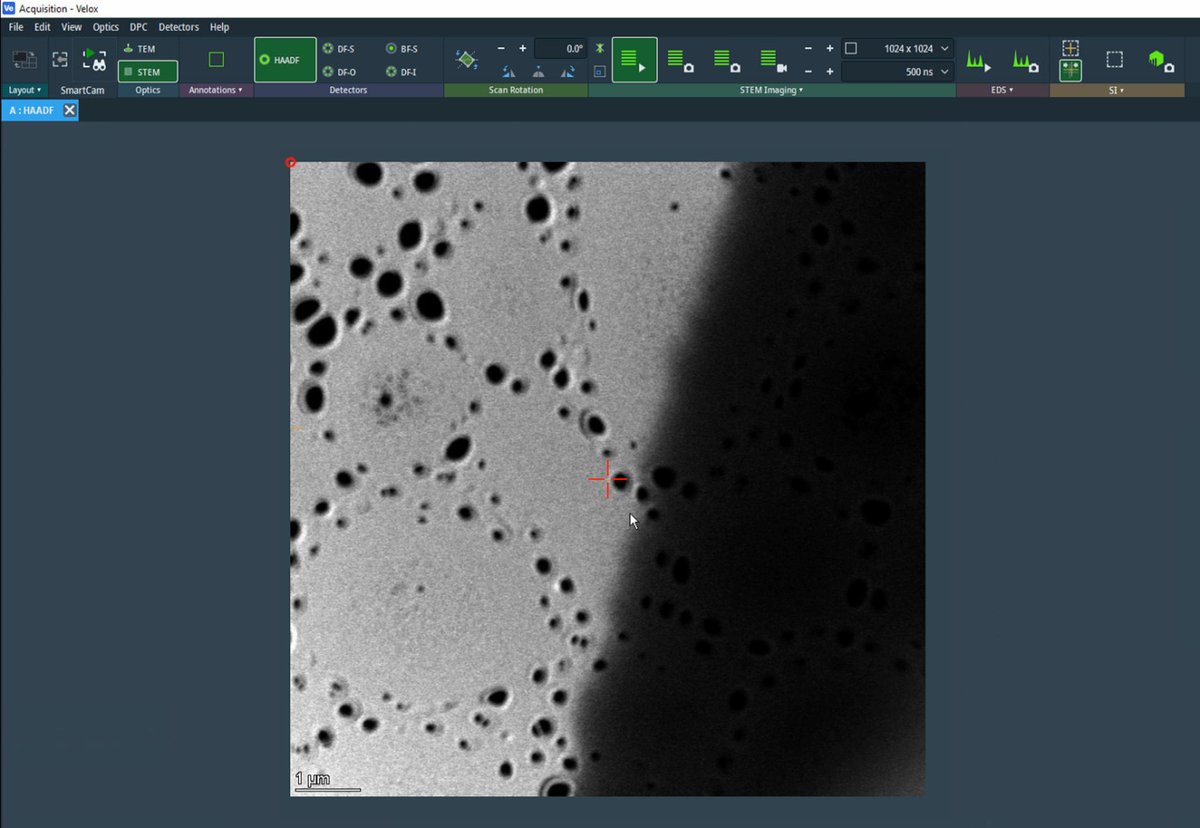
-
-
Adjust focus
-
Once a suitable boundary is found, increase magnification to ~160,000x. Alternate between magnification and z-axis adjustments until focus is sharp.
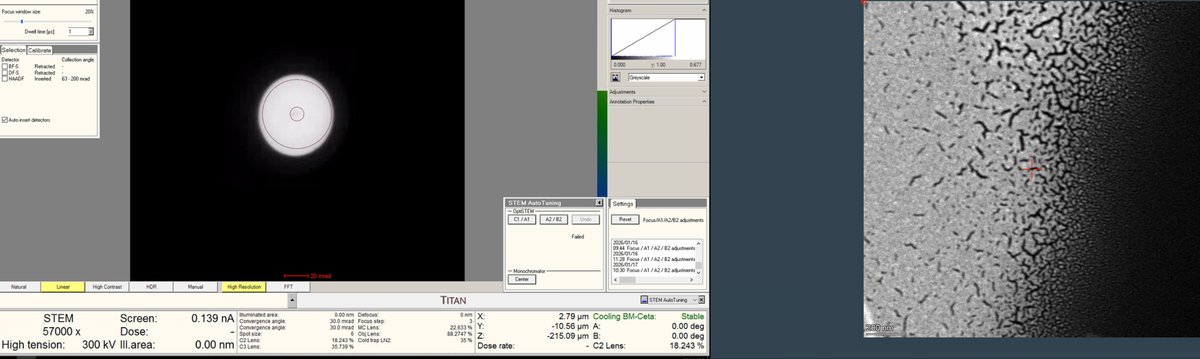
-
Adjust the z-axis while watching the HAADF image. Features become sharper as focus is approached.
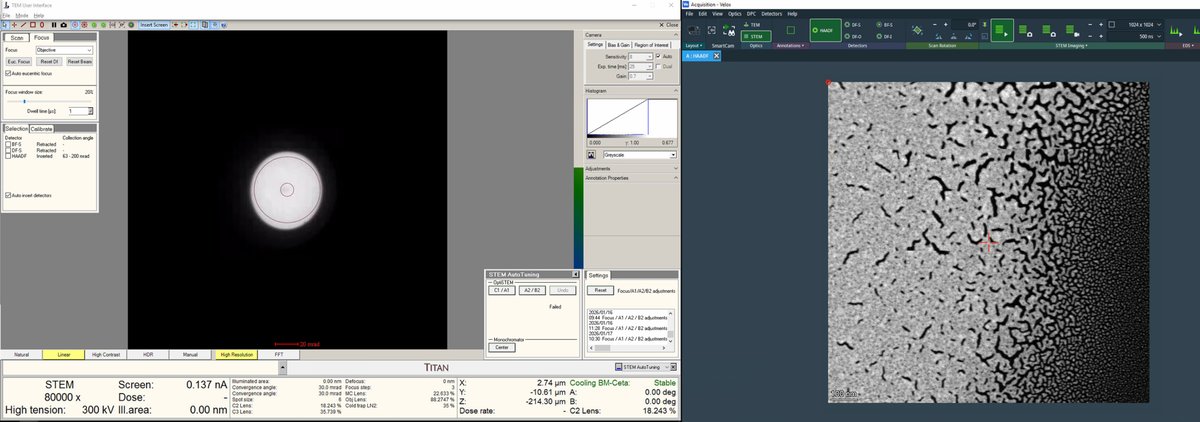
-
Continue adjusting magnification and z-axis.
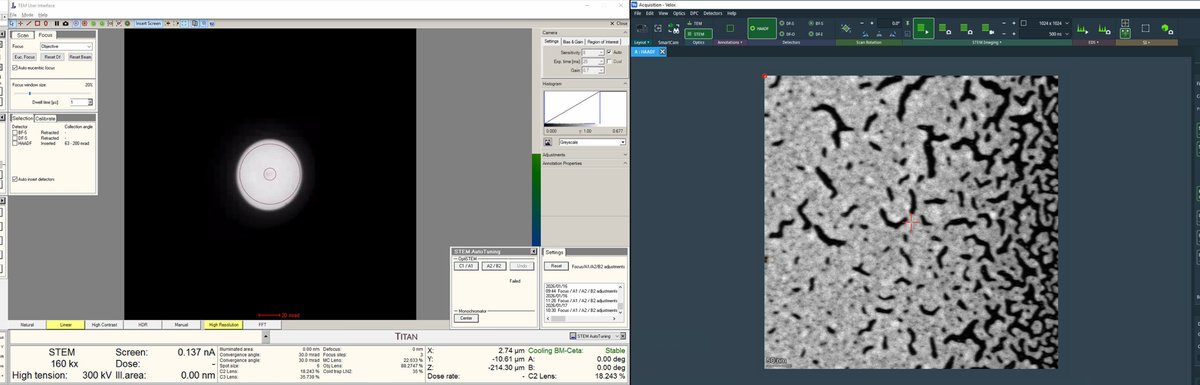
-
Finalize the position with sharp features and distributed particle sizes.
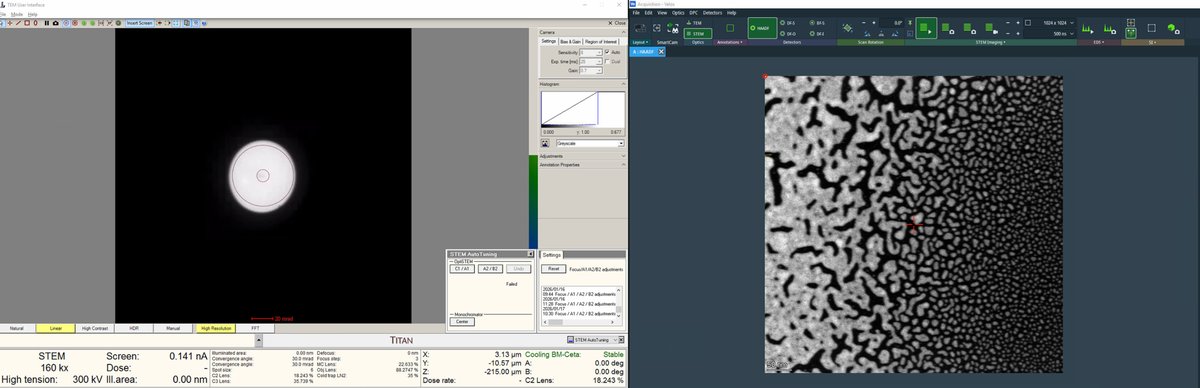
Distributed particles are important for aberration measurement. Aberrations vary with position relative to the optical axis (e.g., coma increases further from center). The correction algorithm requires ronchigram data from multiple positions to accurately fit the aberration coefficients.
-
Part 2: Probe Correction
Before starting probe correction, retract the HAADF detector. C1A1 and Tableau both analyze the ronchigram, and the HAADF ring blocks high-angle electrons from reaching the camera below. In TEMUI, verify the HAADF detector shows “Retracted” status.
TODO: Confirm with staff whether HAADF must be retracted for C1A1/Tableau on Spectra 300 S-CORR.
Aberrations distort the electron probe and degrade image resolution. The goal of probe correction is to achieve a flat, aberration-free ronchigram. The figure below shows how individual aberrations affect the ronchigram appearance:
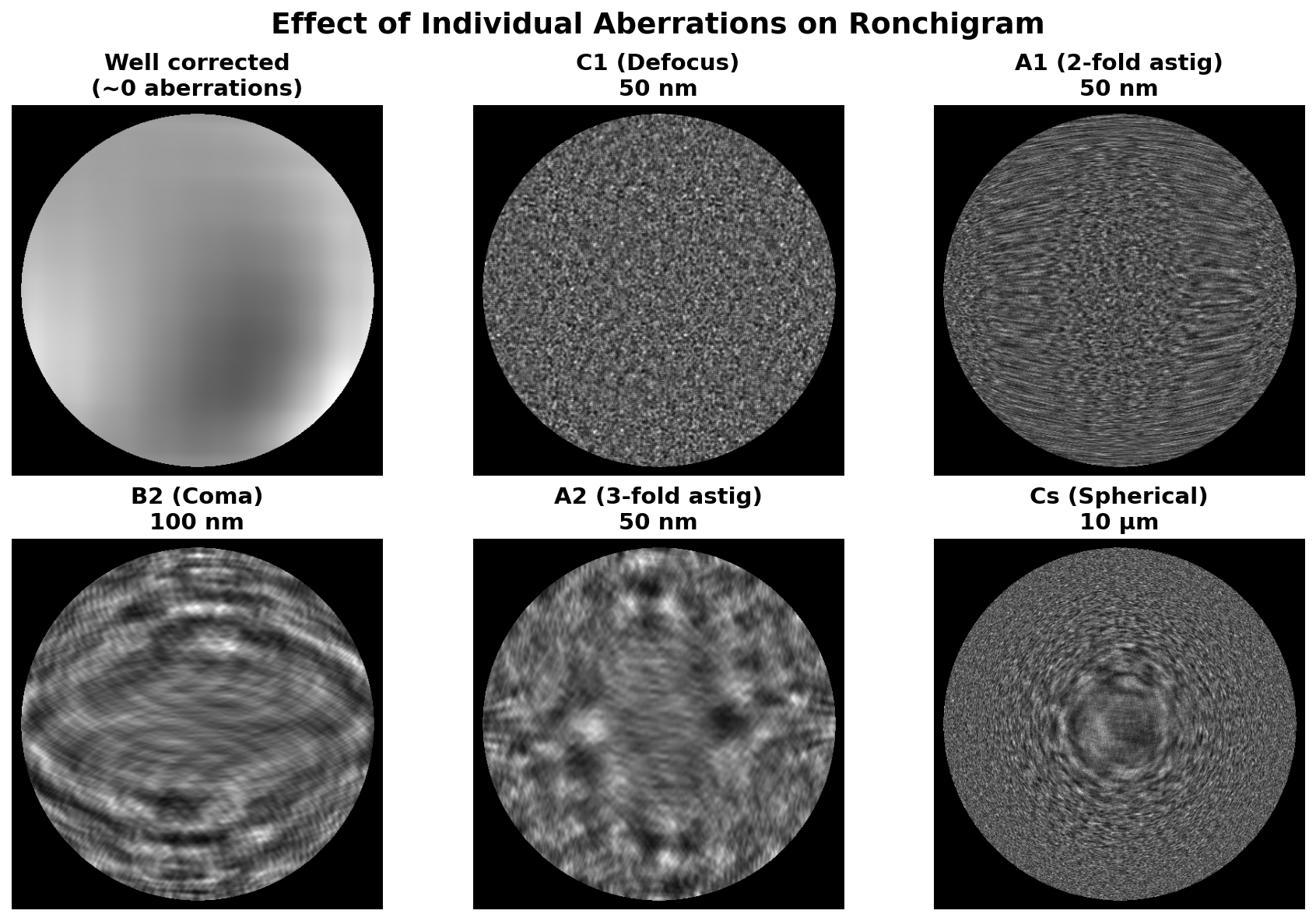
Interactive demo: Explore how aberrations affect the ronchigram at bobleesj.github.io/electron-microscopy-website/ronchigram
Probe correction uses two tools in the Probe Corrector S-CORR software:
| Tool | What it corrects | How it works |
|---|---|---|
| C1A1 | First-order: defocus (C1) and 2-fold astigmatism (A1) | Continuous ronchigram measurement; click buttons to apply |
| Tableau | Higher-order: A2, B2, C3, S3, A3 | Single measurement sequence with beam tilts; then apply |
The correction workflow:
The following workflow is covered in this section. Follow the steps below, then use this diagram as a quick reference:
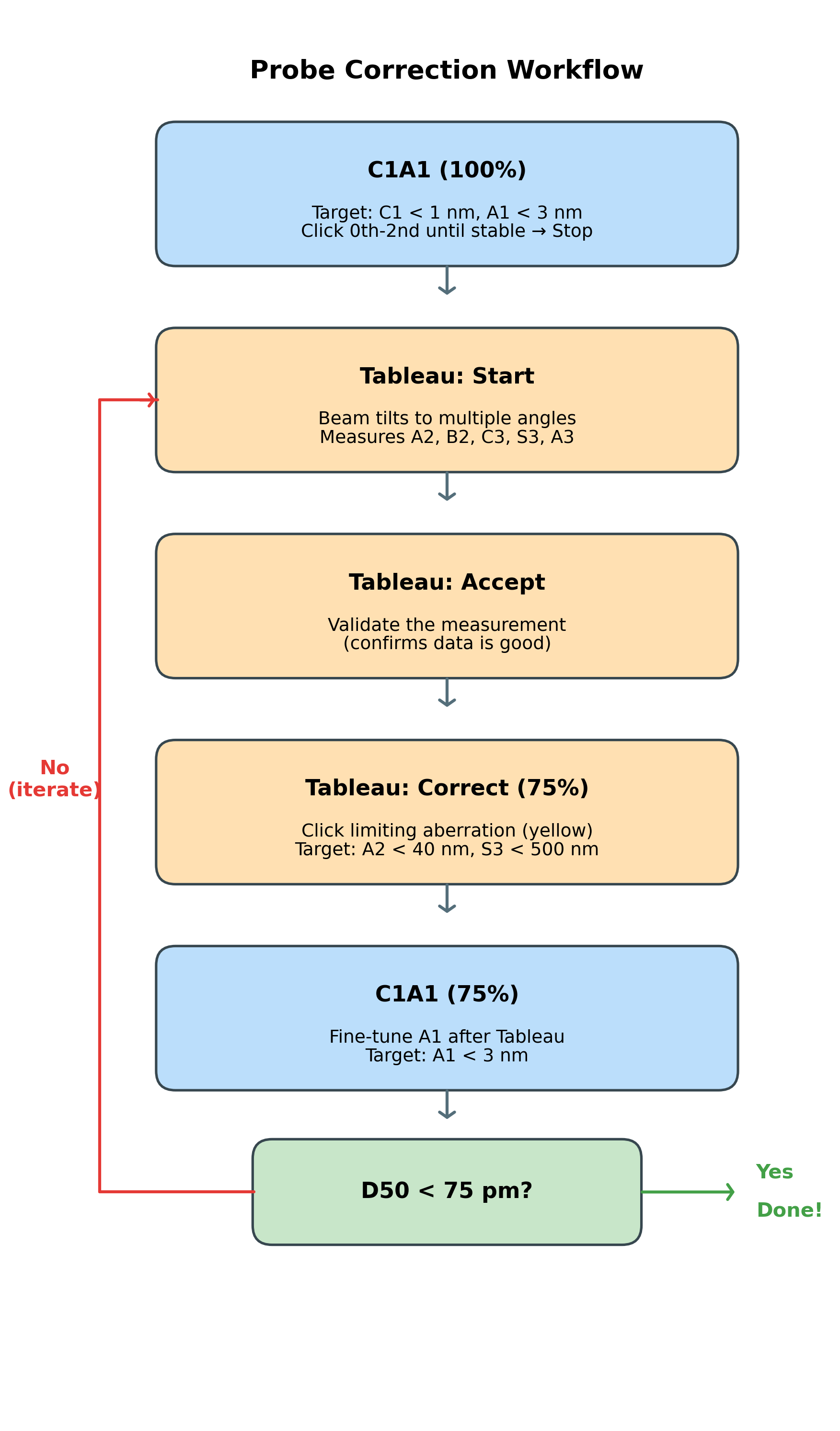
2.1 C1A1 correction
C1A1 corrects first-order aberrations: defocus (C1) and 2-fold astigmatism (A1). These are the dominant aberrations that must be corrected before higher-order Tableau measurement. The C1A1 procedure analyzes the ronchigram to measure and correct these aberrations iteratively.
-
Open Probe Corrector
-
On the top left monitor, open the
Probe Corrector S-CORRsoftware (main interface for aberration measurement and correction). -
Check the mode indicator in the top right. Verify it shows
STEM@300KV: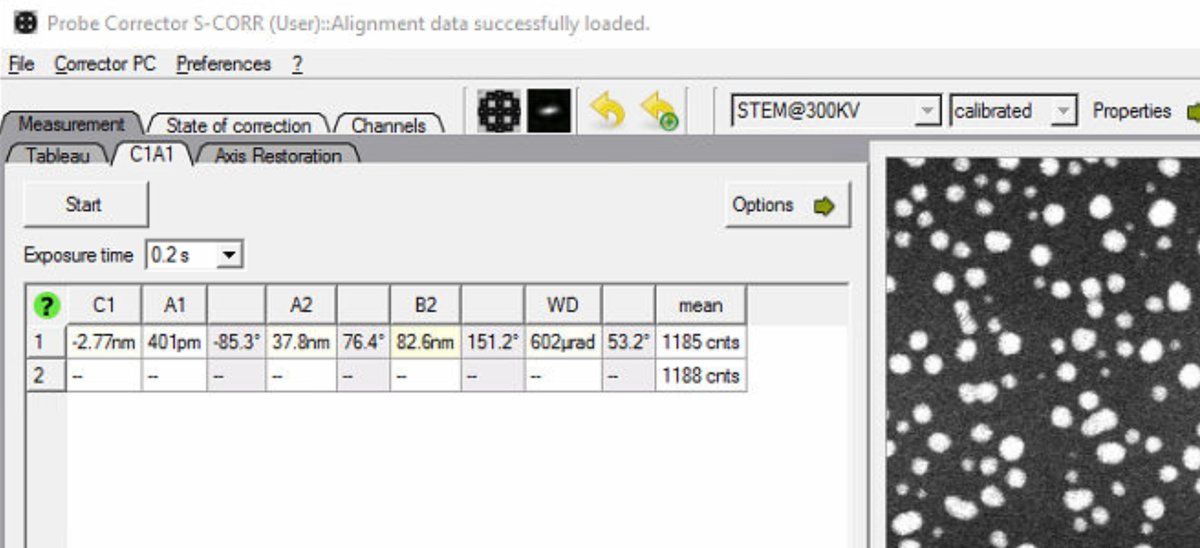
-
If
MC_STEM@300KVappears instead, the system is in monochromated STEM mode. Follow the steps below to reset to standard STEM mode. IfSTEM@300KVis displayed, skip ahead to “Configure C1A1 options.”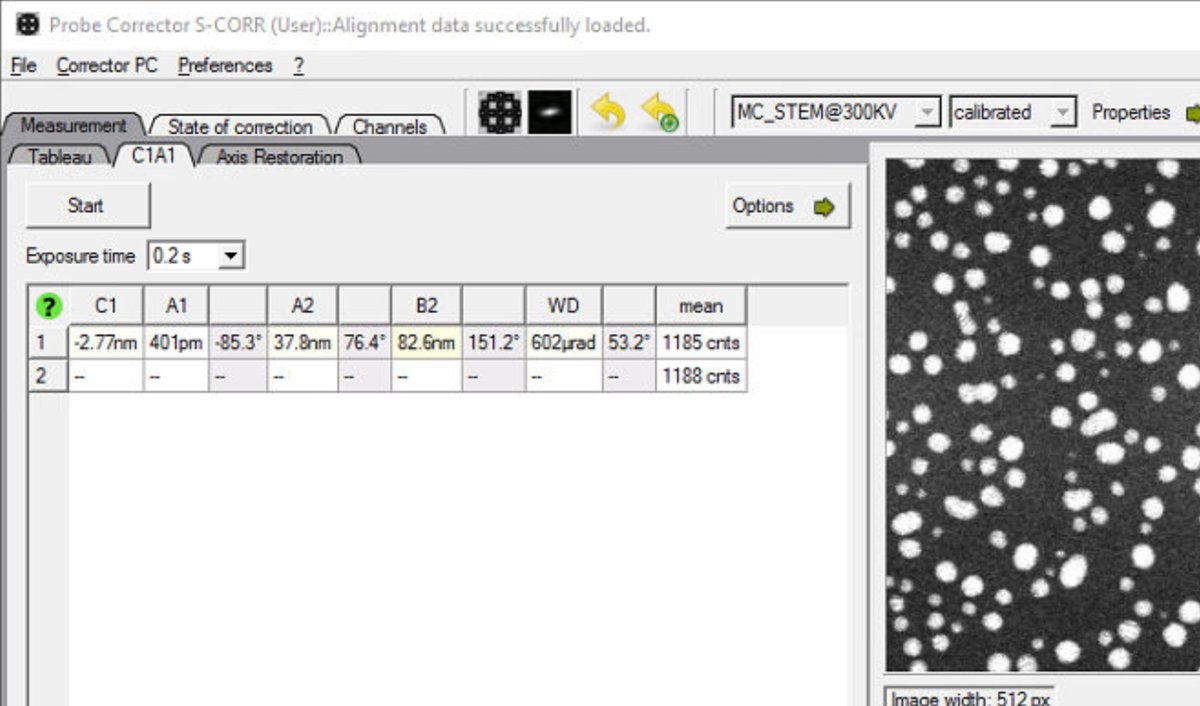
-
To reset, in TEMUI go to
Mono, then openMonochromator (Expert)and clickFilter:
-
Then click
Unfilterto reset to standard STEM mode:
-
-
Configure C1A1 options
-
In the Probe Corrector software, click
Optionsto expand the configuration panel: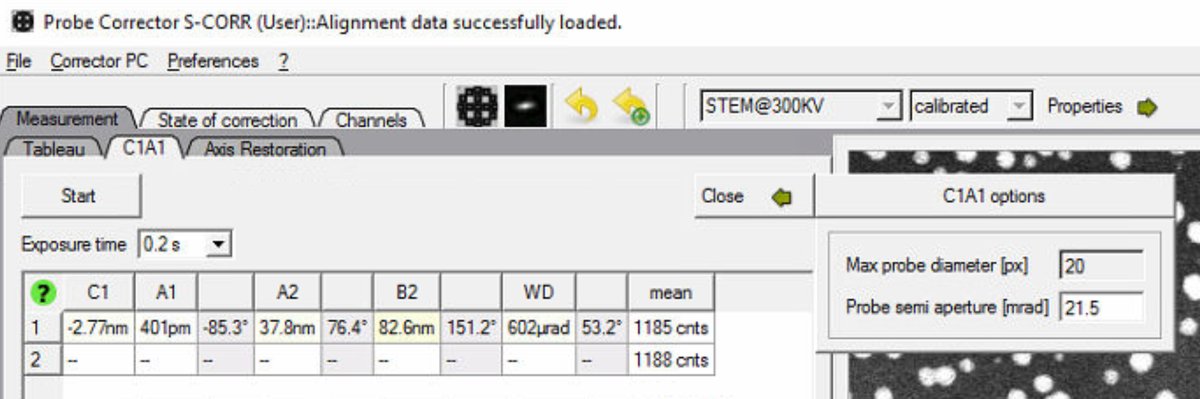
-
Set
Probe semi apertureto 30 mrad: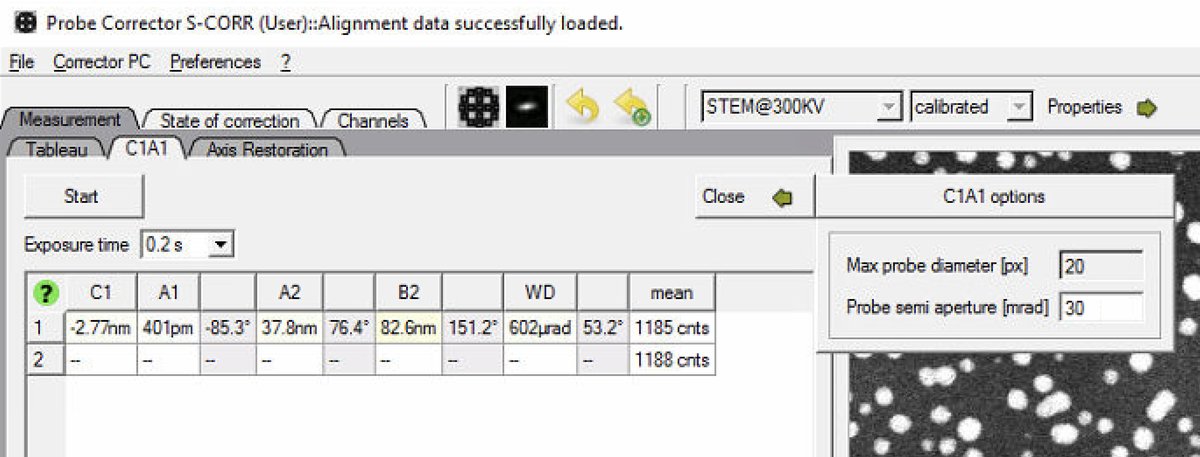
-
-
Switch to diffraction mode
-
Stop live scanning in
Veloxby clicking the play button, then ensureDiffractionmode is on on the hand panel. C1A1 analyzes the ronchigram, so diffraction mode (stationary probe) is required, not probe image mode (scanning):
-
Click the
Beam Blankbutton to unblank the beam. Stopping the scan automatically blanks the beam. The Probe Corrector software requires an unblanked beam to read the ronchigram:
-
-
Run C1A1
-
Go to the
C1A1tab in theProbe Correctorsoftware. Before clicking Start, verify the ronchigram is visible on the left monitor: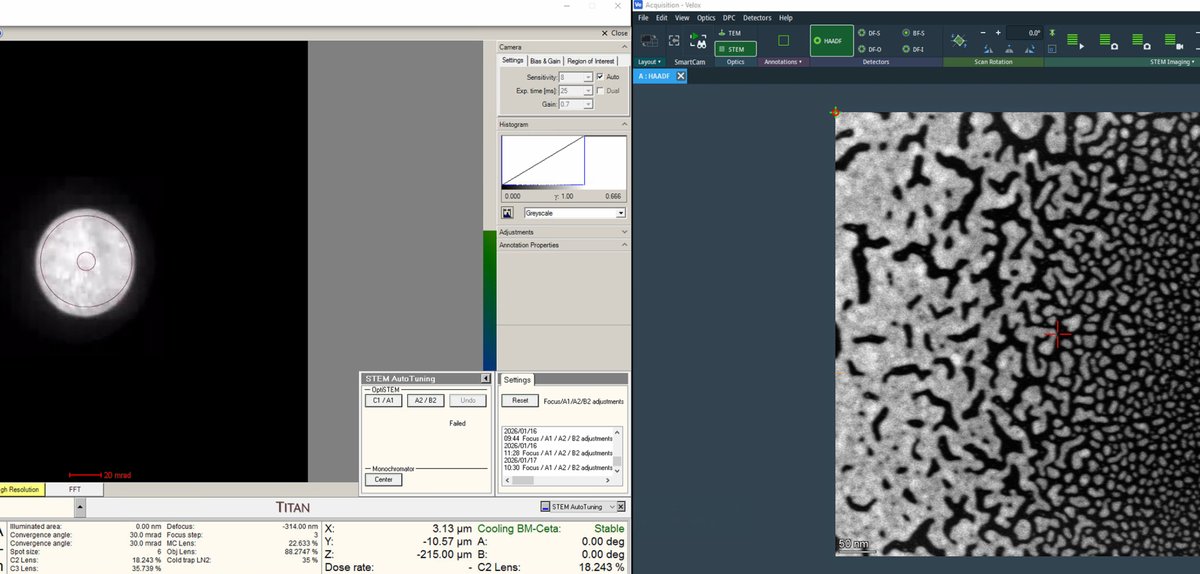
-
Click
Startto begin aberration measurement. The software continuously analyzes the ronchigram and displays measured aberration values (C1, A1, A2, B2, WD) in the table. -
Set Auto correct to 100% for the first iteration.
-
Click
0th-2ndto apply corrections for all first and second order aberrations.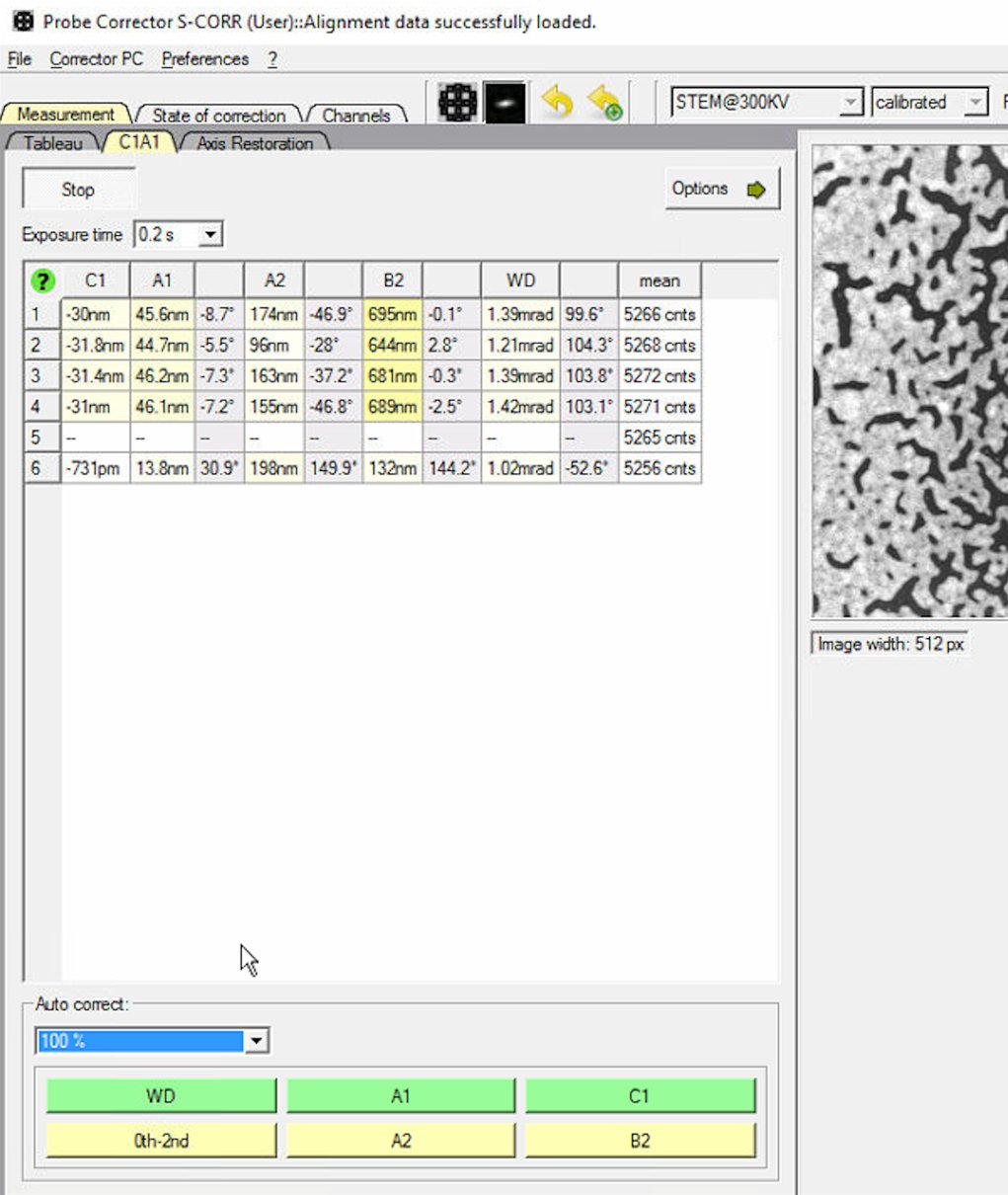
-
-
Iterate C1A1
-
Click
0th-2ndrepeatedly to apply corrections. Each row represents one measurement cycle. Watch the aberration values decrease with each iteration.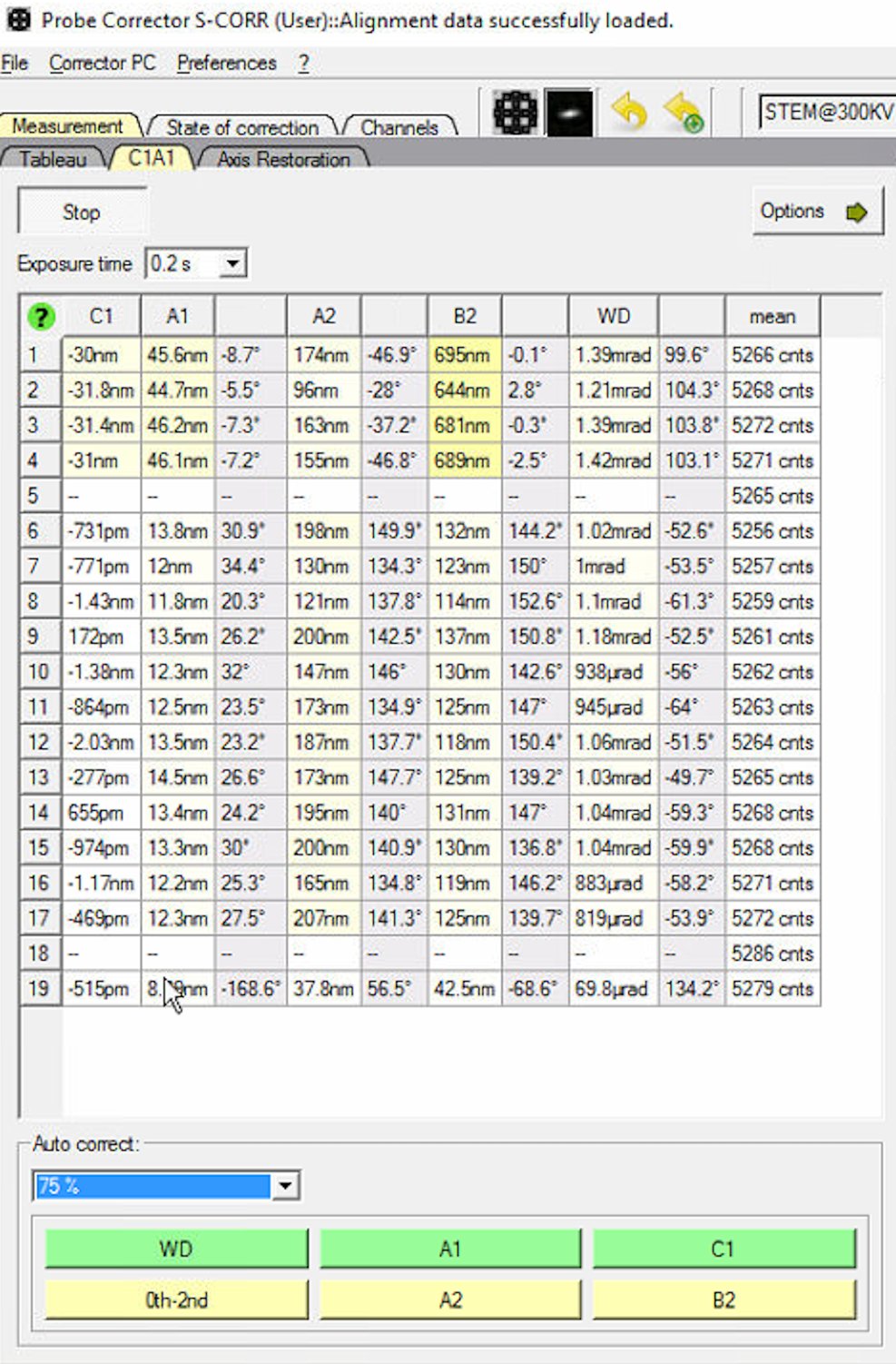
-
Reduce the Auto correct percentage to 75% after several iterations (typically 3 to 5) to prevent overcorrection.
If A1 is still high but other values are good, click
A1specifically to correct only astigmatism.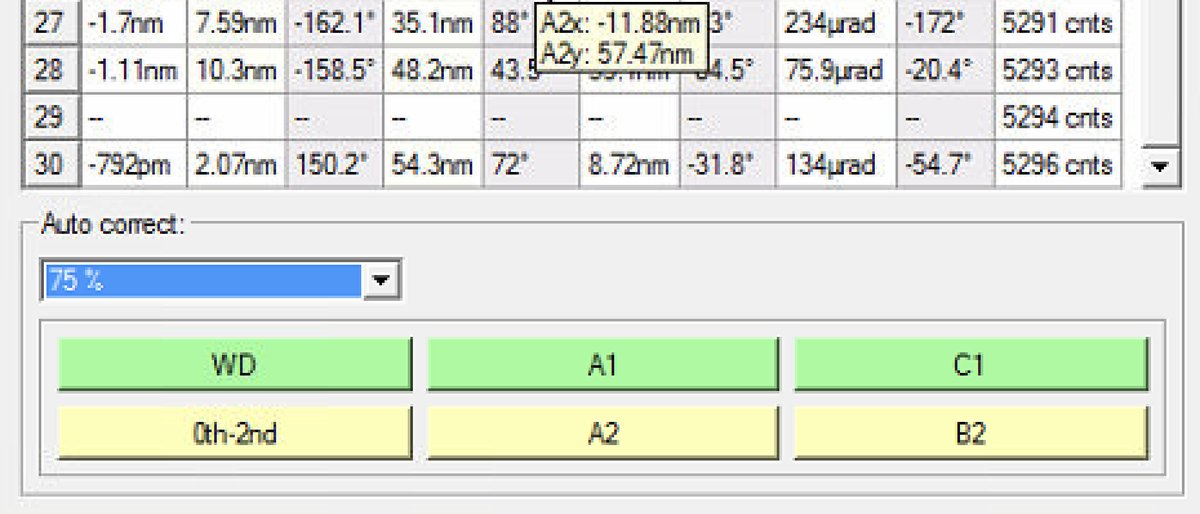
-
When to stop: C1A1 values are stable when they no longer decrease significantly between iterations. Target: C1 (defocus) < 1 nm and A1 (astigmatism) < 3 nm. Click
Stopwhen values are stable.
-
2.2 Tableau measurement
Tableau measures higher-order aberrations (A2, B2, C3, S3, A3) by acquiring ronchigram patterns at multiple beam tilt angles. The software analyzes how the ronchigram changes with tilt to extract the full aberration function. Tableau is more comprehensive than C1A1 and necessary for highest resolution.
-
Open Tableau tab
-
Switch to the
Tableautab in theProbe Correctorsoftware for full aberration measurement and correction. -
Select
Standardfor Tableau type. This acquires a sufficient number of tilt positions for accurate measurement without taking excessive time. -
Set the Outer tableau tilt to 40 mrad. Larger tilts probe higher-order aberrations but require more time.
-
Verify the Probe semi aperture is set to 30 mrad to match the beam settings.
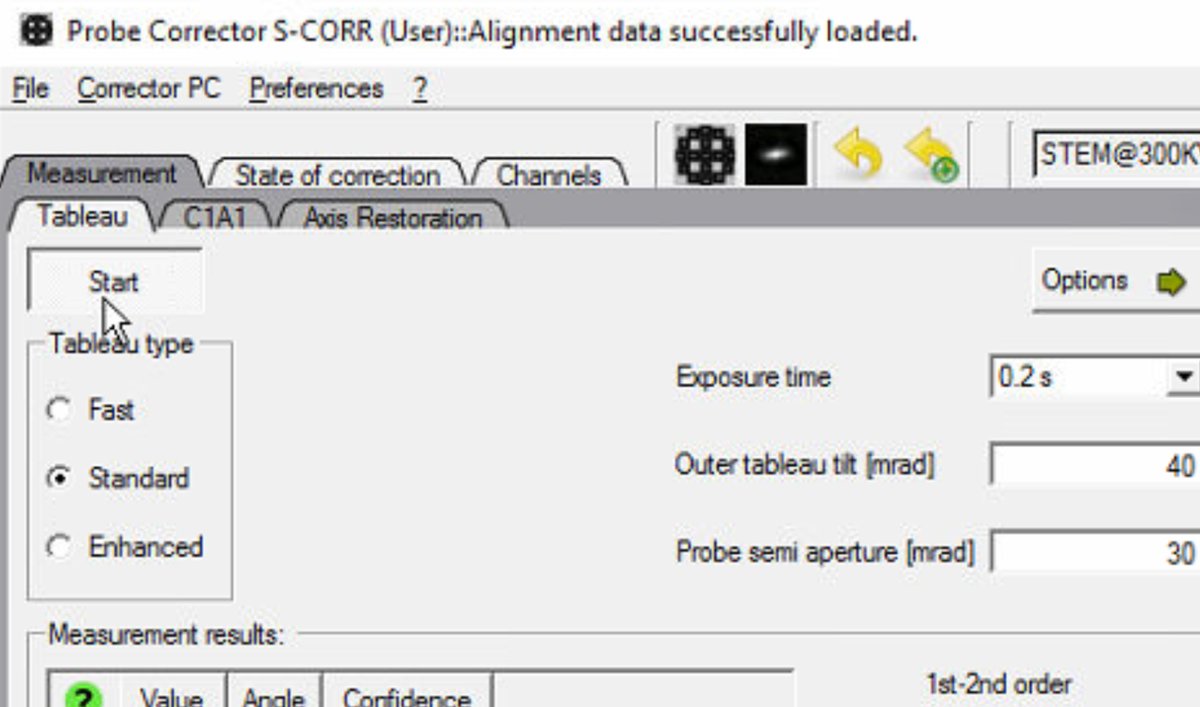
-
Click
Optionsand select theA5toggle. It measures up to 5th order aberrations.
-
-
Run Tableau measurement
-
Click
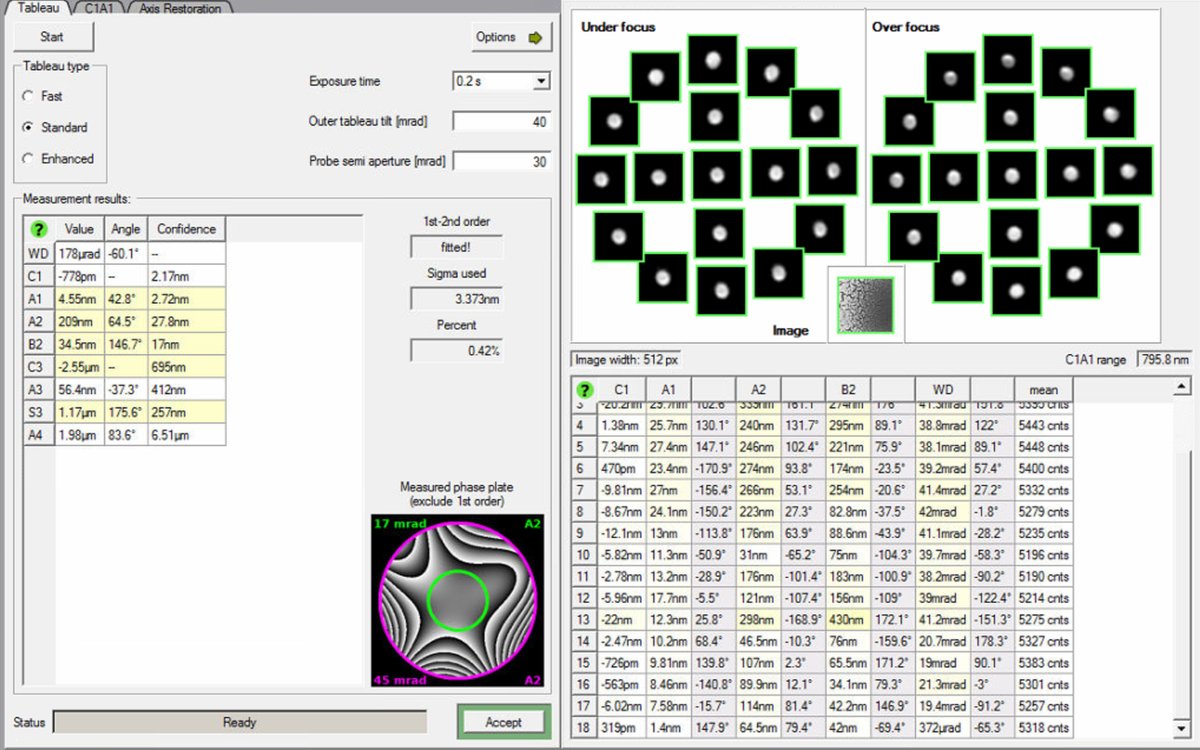
Startto begin the Tableau measurement. The software automatically tilts the beam to multiple angles and acquires ronchigram images at each position. -
Wait for measurement completion. The ronchigram shifts across the screen as the software captures patterns at different tilts and focus levels (under-focus and over-focus at each tilt). This movement is expected.
-
Do not modify the stage position during measurement. If the beam is unstable, stop and ask staff.
-
-
Accept measurement
- Click
Acceptafter measurement completes. This validates the data for corrections.
- Click
-
Review measurement results
-
Click the
State of correctiontab. This shows all measured aberration coefficients in three columns:- Estimation: Just measured values
- Latest accepted measurements: Previously applied corrections (yellow = outside limits)
- Estimation in image coordinate system: Values transformed to image coordinates
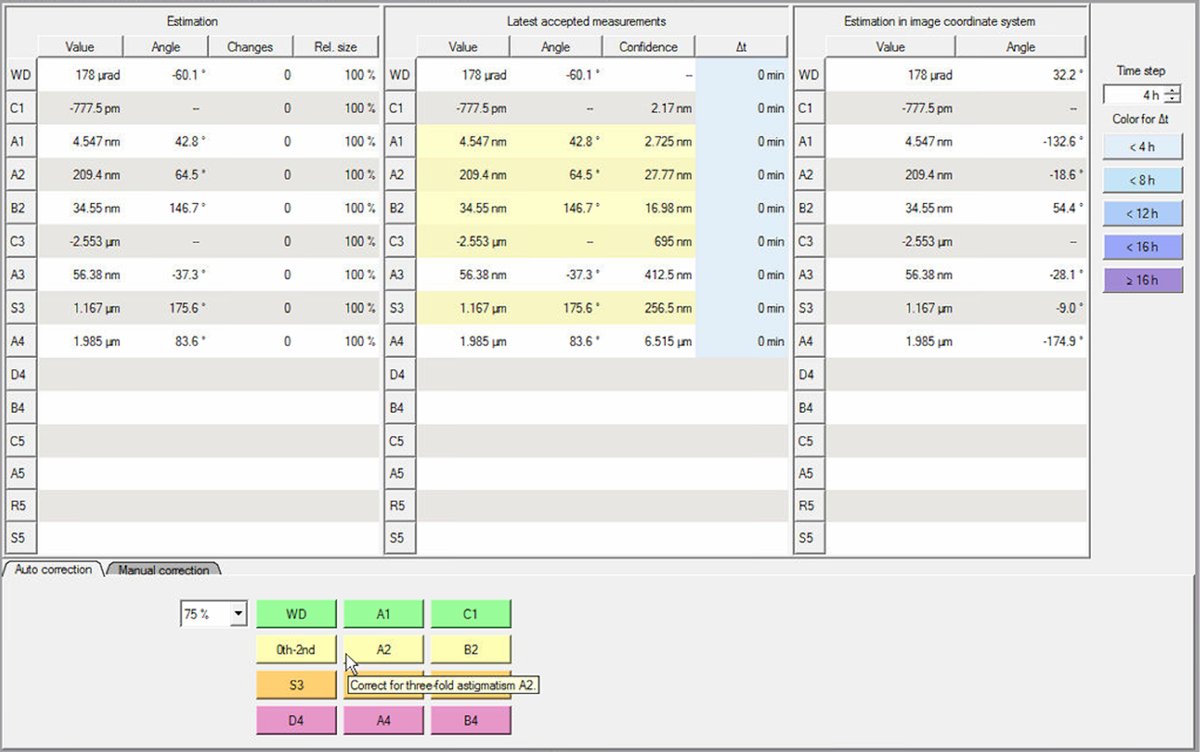
-
Check the phase plate visualization on the right. A well corrected probe has a flat, symmetric phase plate. Strong asymmetric patterns indicate uncorrected aberrations:
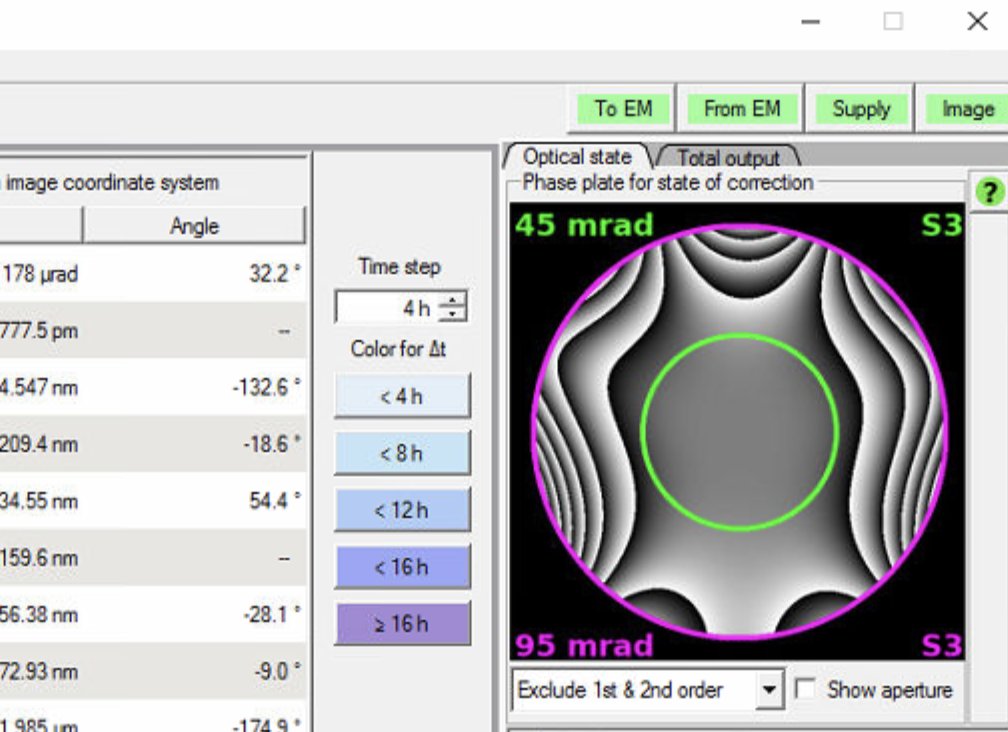
-
-
Apply corrections
-
Set Auto correct to 75% to prevent overcorrection. Yellow highlighted values in the “Latest accepted measurements” column indicate aberrations outside acceptable limits. Correct these first. In this example, S3 (1.167 μm) and C3 (-2.553 μm) are highlighted yellow:
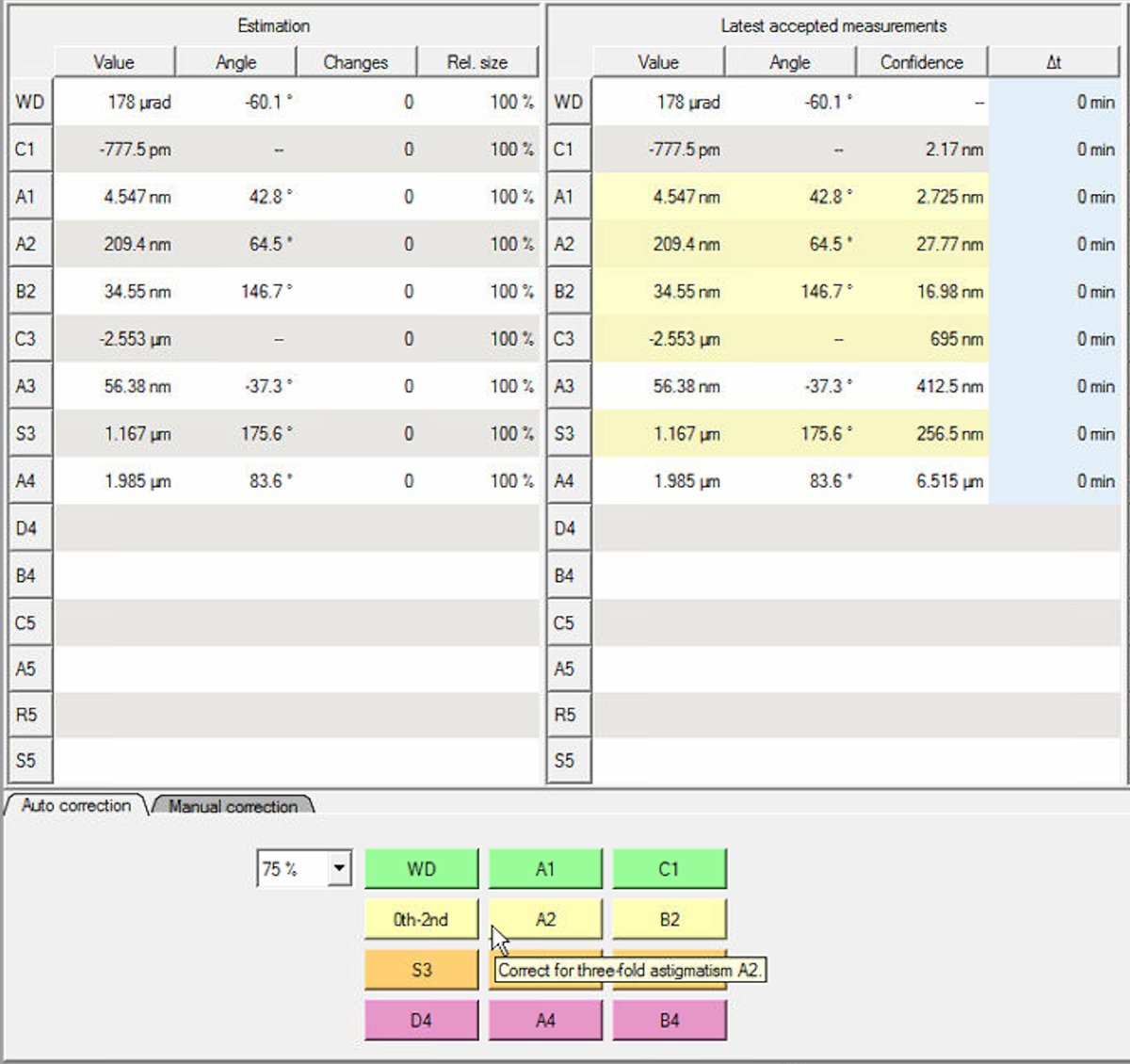
Note: either clicking
B4orD4can have a significant impact onC1andA1values.-
Click the aberration buttons at the bottom to apply corrections. The phase plate visualization shows the limiting aberration. Correct this one first. Click the button repeatedly until the value improves sufficiently, then move to the next limiting aberration.
-
The “Changes” column tracks how many corrections have been applied. After correcting S3 and C3, the values improve significantly:
- S3: 1.167 μm → 72.93 nm
- C3: -2.553 μm → -159.6 nm
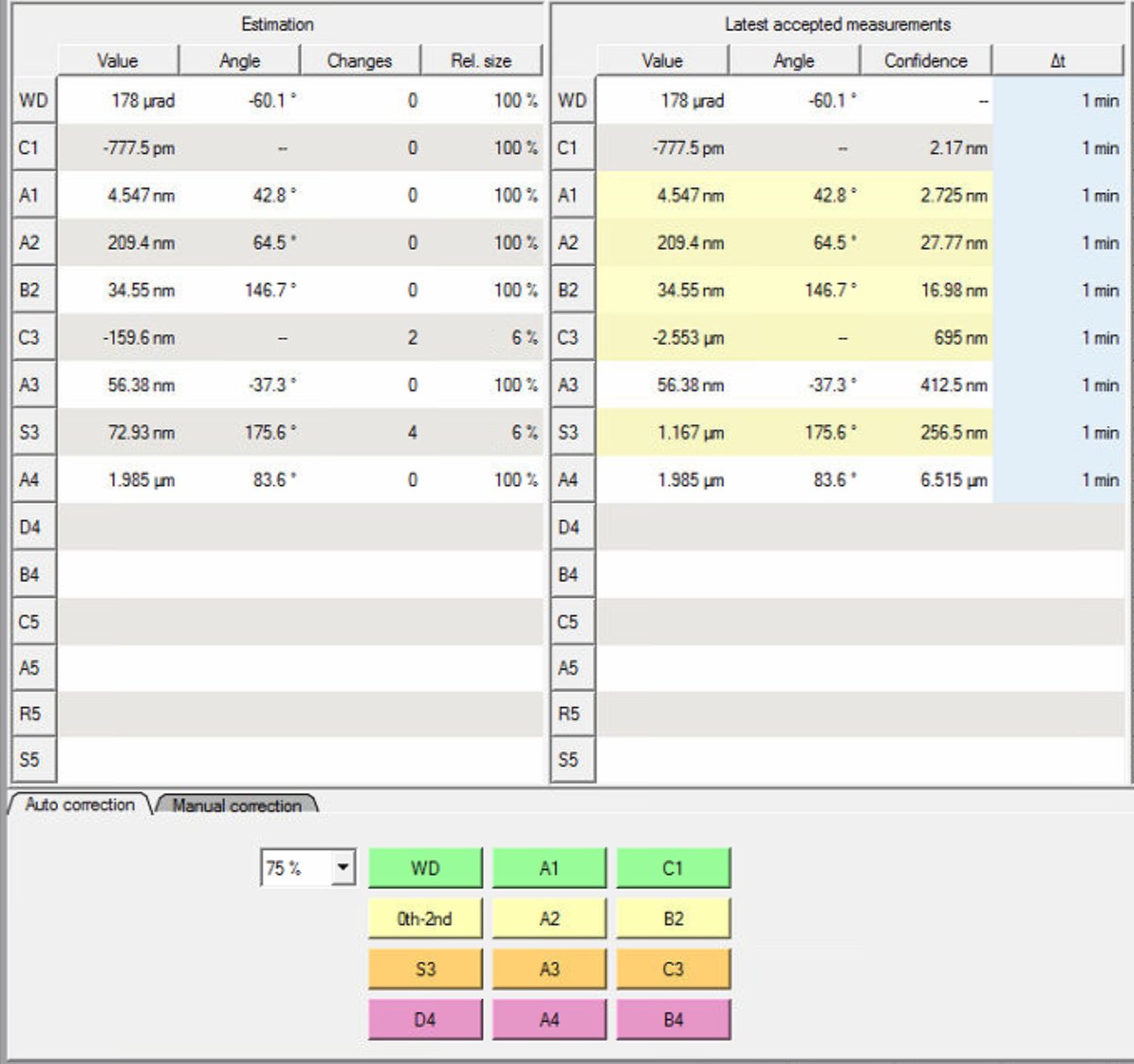
-
-
Run full measurement
-
After applying corrections, run another complete Tableau measurement to verify the improvements.
-
Check the aberration surface and phase plate displays. A well-corrected probe shows:
- Flat aberration surface with green in the center (minimal phase variation across the aperture)
- Symmetric phase plates without strong directional features
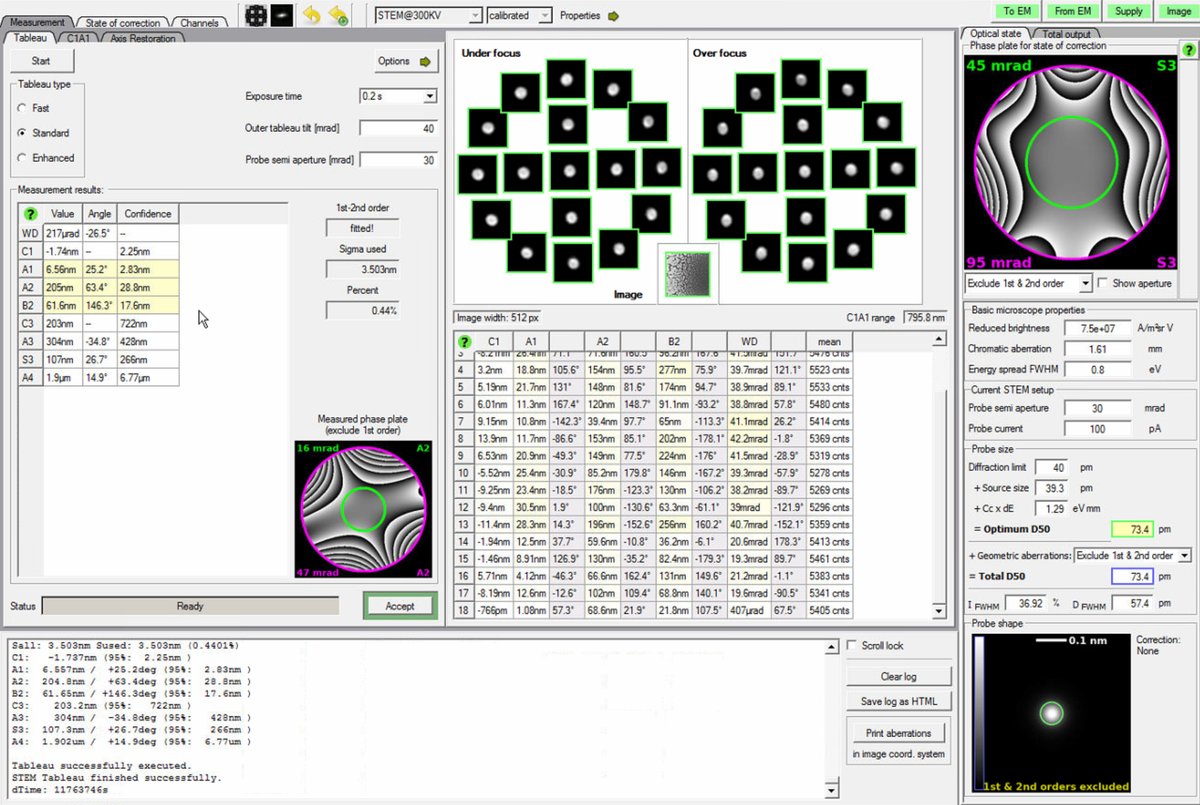
Target values (30 mrad semi-aperture):
Parameter Target C1 < 1 nm A1 < 3 nm A2 < 40 nm B2 < 25 nm C3 < 1.5 μm A3 < 1 μm S3 < 500 nm TODO: CONFIRM WITH ANDREW for IDEAL TOTAL D50
-
-
Verify with C1A1
- Return to the
C1A1tab in the Probe Corrector software. Tableau correction can sometimes introduce small first-order errors. - Click
Startto begin C1A1 measurement again. - Click
A1to correct any residual astigmatism introduced by Tableau. - Click
0th-2ndif defocus also needs adjustment. - Iterate between Tableau and C1A1 if necessary until all values are within specification.
- Return to the
-
Check resolution
-
The
State of correctionpanel displays resolution estimates on the right side: Total D50 and Optimum D50. D50 represents the probe diameter containing 50% of the beam intensity (smaller = better resolution).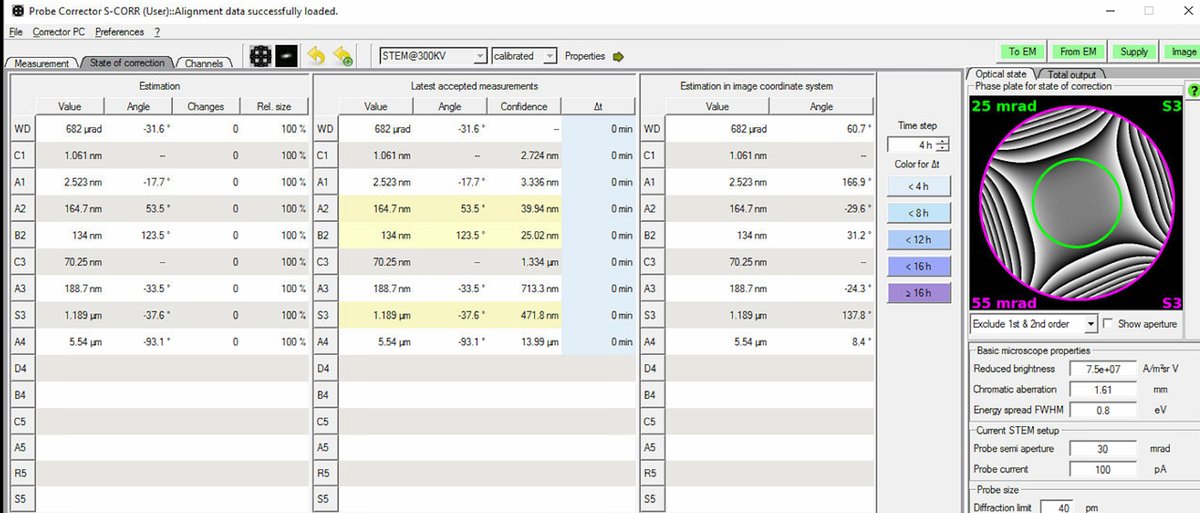
TODO: CONFIRM WITH ANDREW
-
Target: Total D50 of 70-75 pm for high-resolution STEM imaging. The Optimum D50 shows the theoretical best achievable with current aberrations. If these values match closely, corrections are complete.
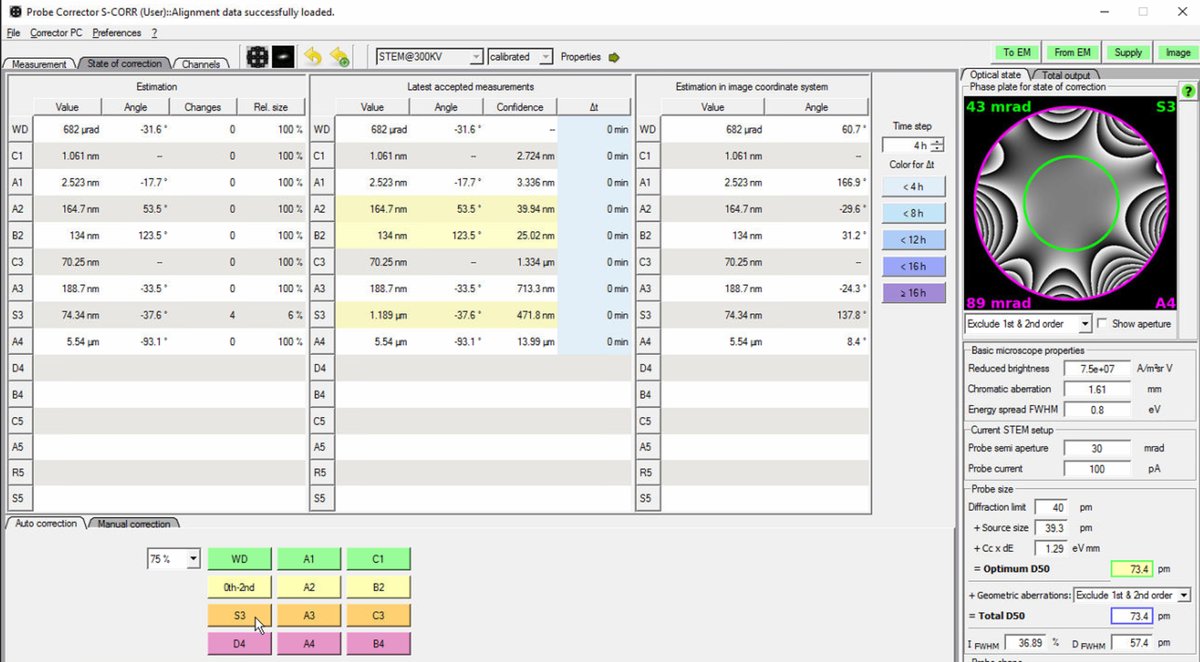
-
If D50 values are significantly higher than target, continue iterating: run another Tableau measurement, apply corrections, then verify with C1A1. The image below shows C1A1 iterations after Tableau corrections:
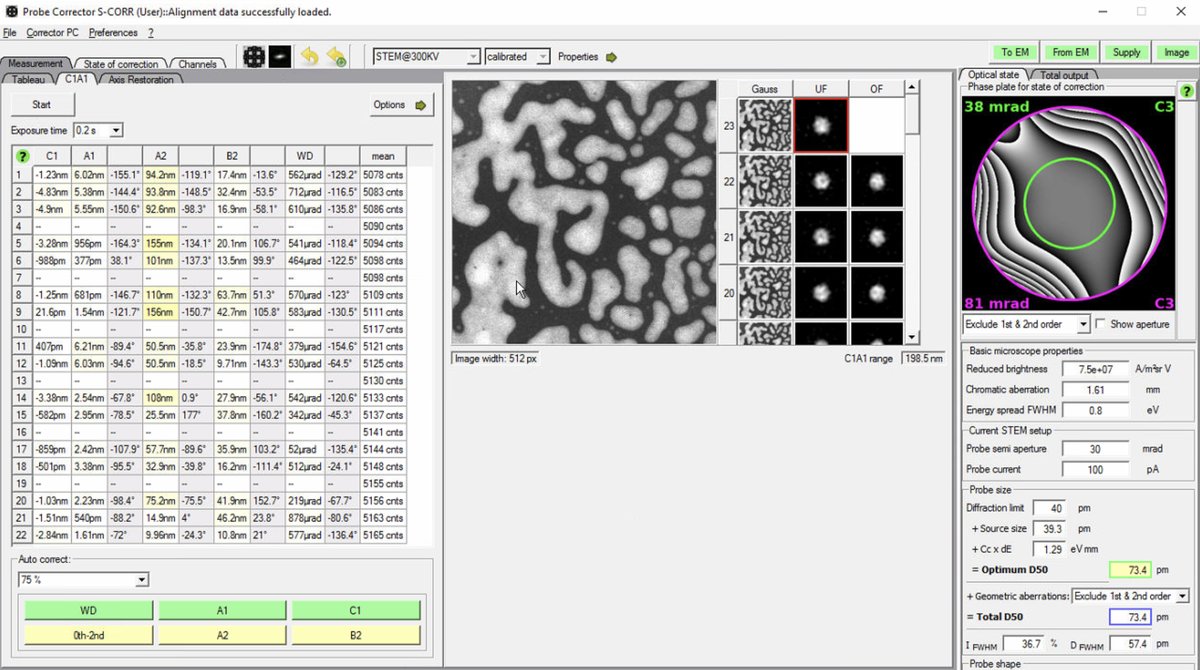
-
-
Return to probe image mode
- Once correction is complete, press the
Diffractionbutton on the hand panel to switch back to probe image mode (STEM scanning). - The system is now ready for high-resolution image acquisition.
- Once correction is complete, press the
Part 3: Imaging
3.1 Acquire images
With aberration correction complete, the system is ready for high-resolution STEM image acquisition. The probe is optimized for atomic-resolution imaging.
-
Acquire HAADF image
-
In
Velox, clickSTEMto enter STEM mode, then clickHAADFto select the HAADF detector. -
Click the play button to start live scanning. Image quality is noticeably improved compared to before correction. A well-corrected probe produces sharper, more detailed images.
-
For initial survey imaging, set resolution to 1024×1024 and dwell time to 500 ns. Fast scanning enables navigation while maintaining image quality.
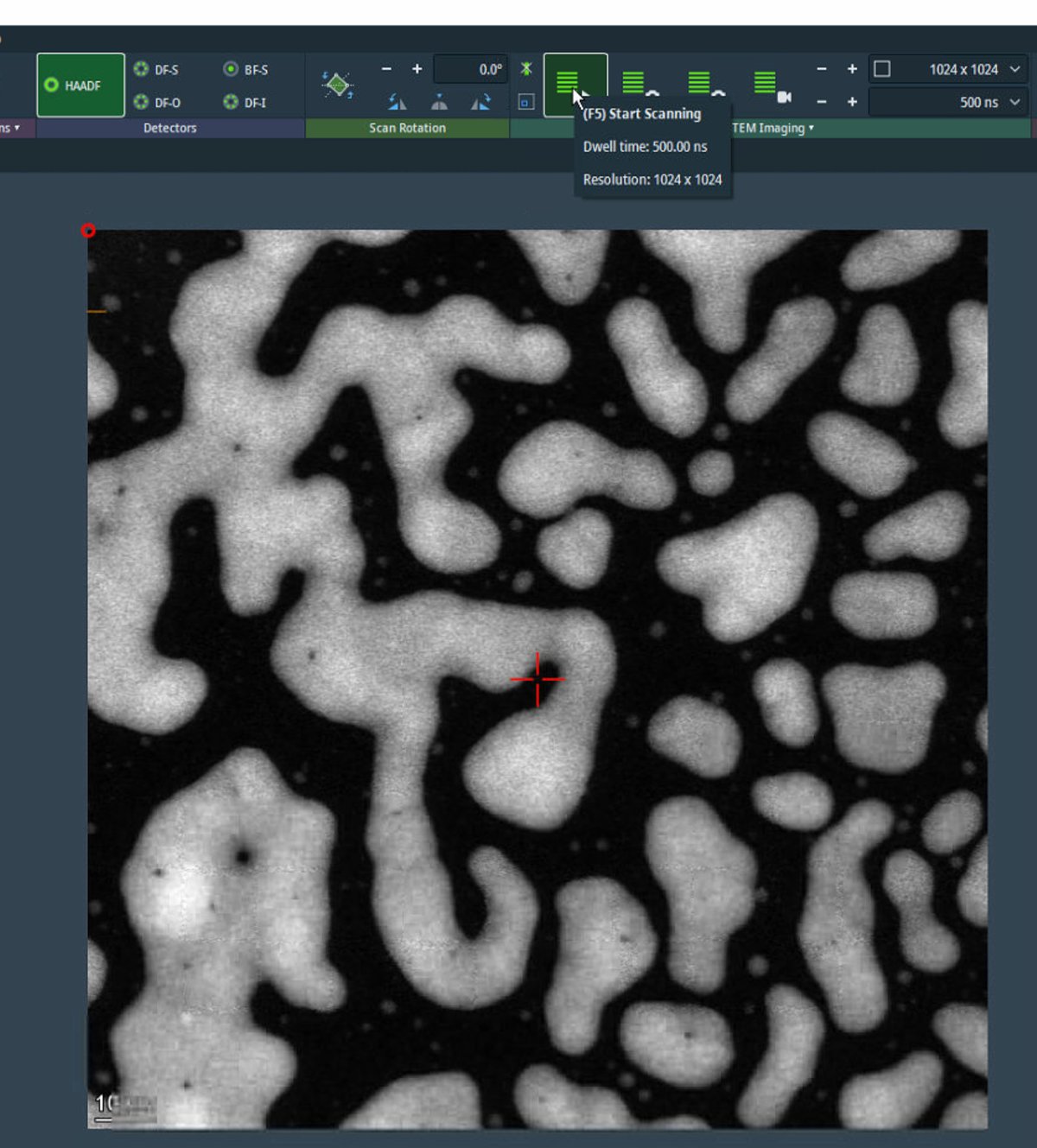
-
-
Navigate to area of interest
-
Use the live scan to find the region of interest. Use the joystick or click on the image to move to different regions. With a well-corrected probe, atomic lattice fringes are visible in crystalline materials.
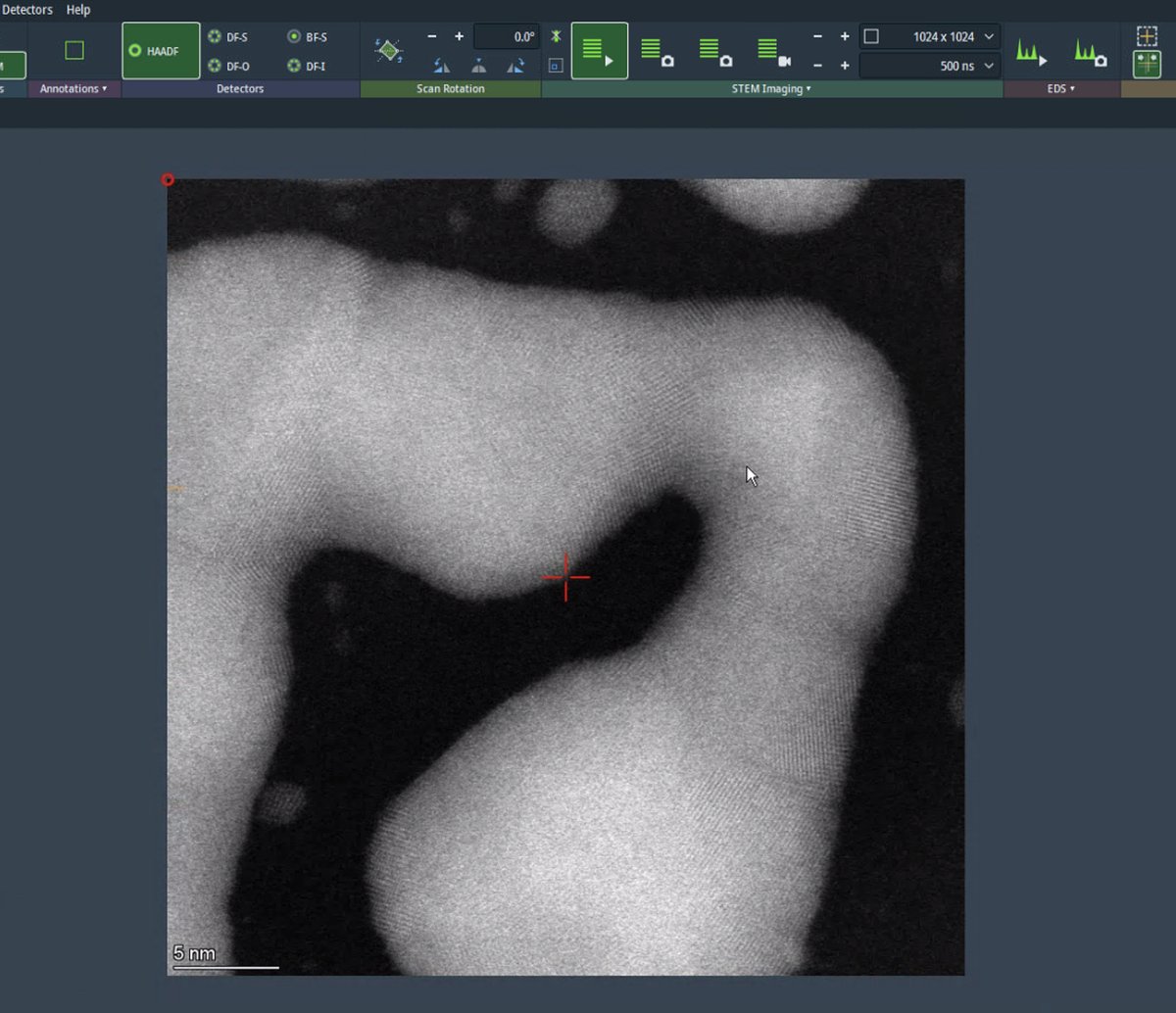
-
Adjust focus using the z-height controls if needed. Small focus changes can significantly affect atomic-resolution contrast.
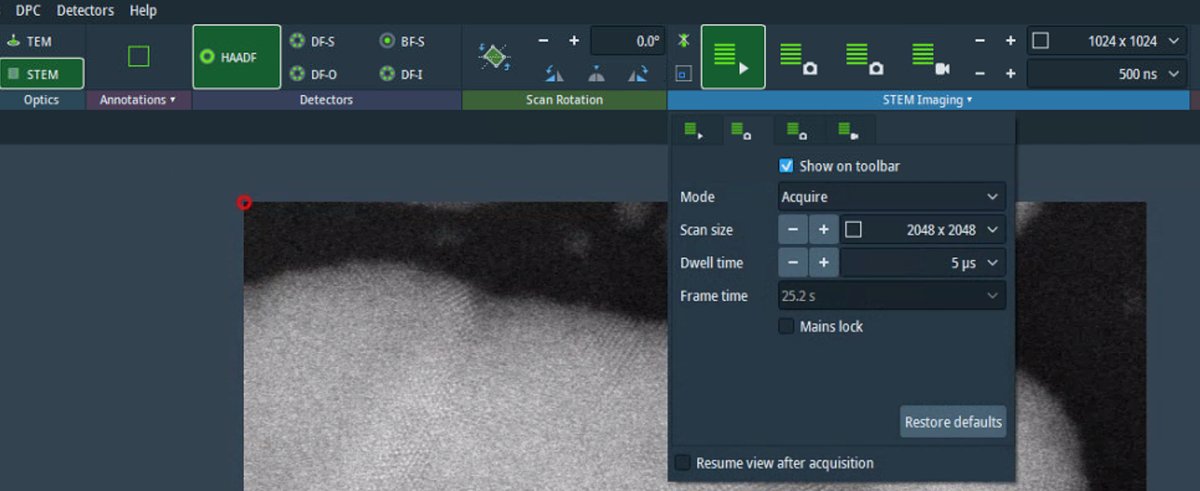
-
-
Capture high-resolution scan
-
Increase the resolution to 2048×2048 or higher. Check the Velox toolbar to verify resolution and dwell time settings before starting the acquisition.
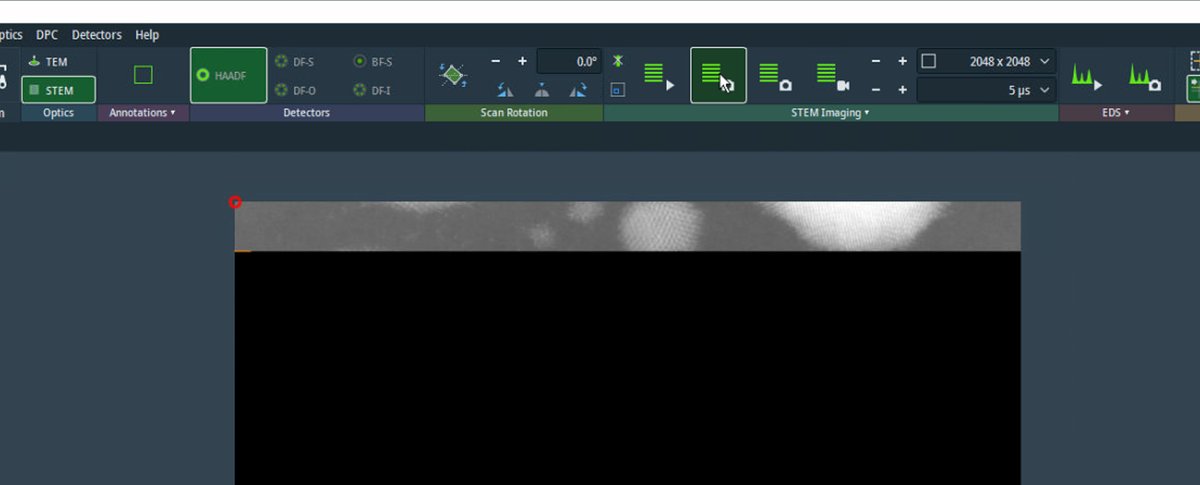
-
Increase the dwell time to 5 µs for better signal-to-noise ratio. Longer dwell times collect more electrons per pixel, reducing noise but increasing total scan time and potential for drift artifacts. After acquisition completes, the beam is blanked automatically to prevent sample damage.
-
3.2 Fine-tuning with Sherpa
Sherpa provides rapid aberration correction that is faster than full Tableau measurement. Use Sherpa for quick refinements after the main alignment, or when aberrations drift during extended imaging sessions.
-
Prepare for Sherpa
-
Before running Sherpa, verify the ronchigram is centered. Press the
Diffractionbutton on the hand panel to switch to diffraction mode (view the ronchigram). -
In
TEMUI, go toDirect Alignmentsand selectDiffraction Shift and Focus alignment: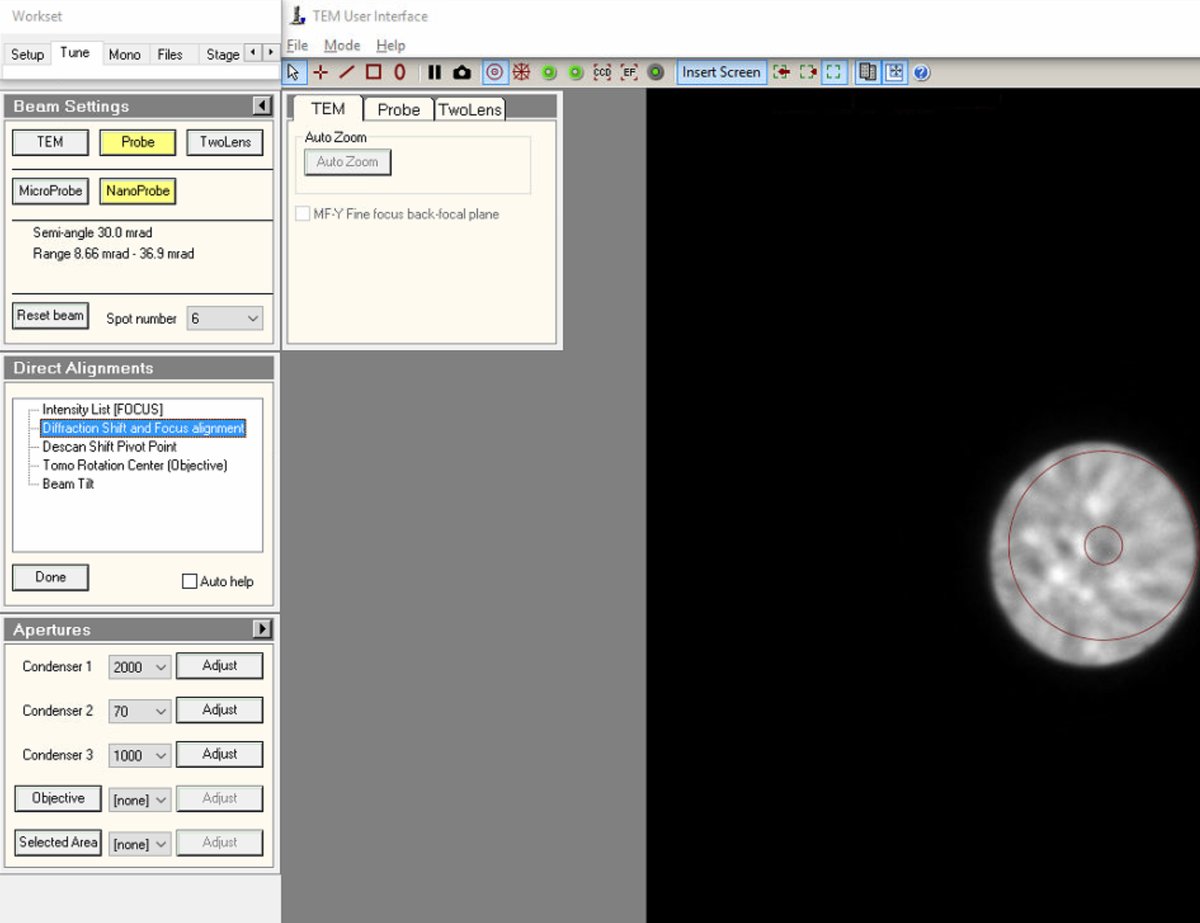
-
Use the
mulXYknobs to center the ronchigram on the display. A centered ronchigram ensures Sherpa measurements are accurate.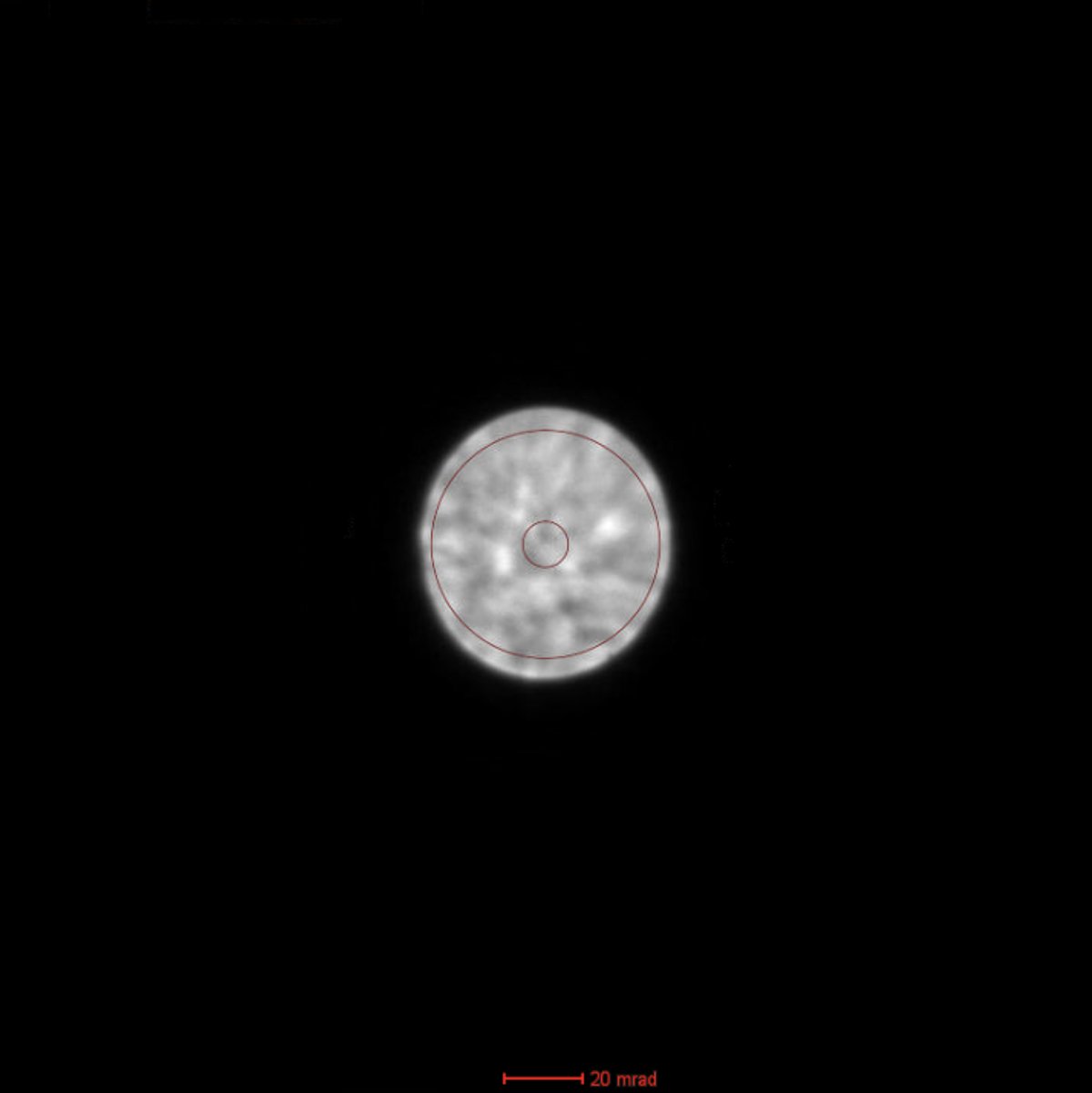
-
-
Adjust C2 aperture (optional)
-
To change C2 aperture size (for example, switching to 50 µm for different probe conditions), locate the
Aperturespanel and change Condenser 2 from 70 to 50 (or the desired size).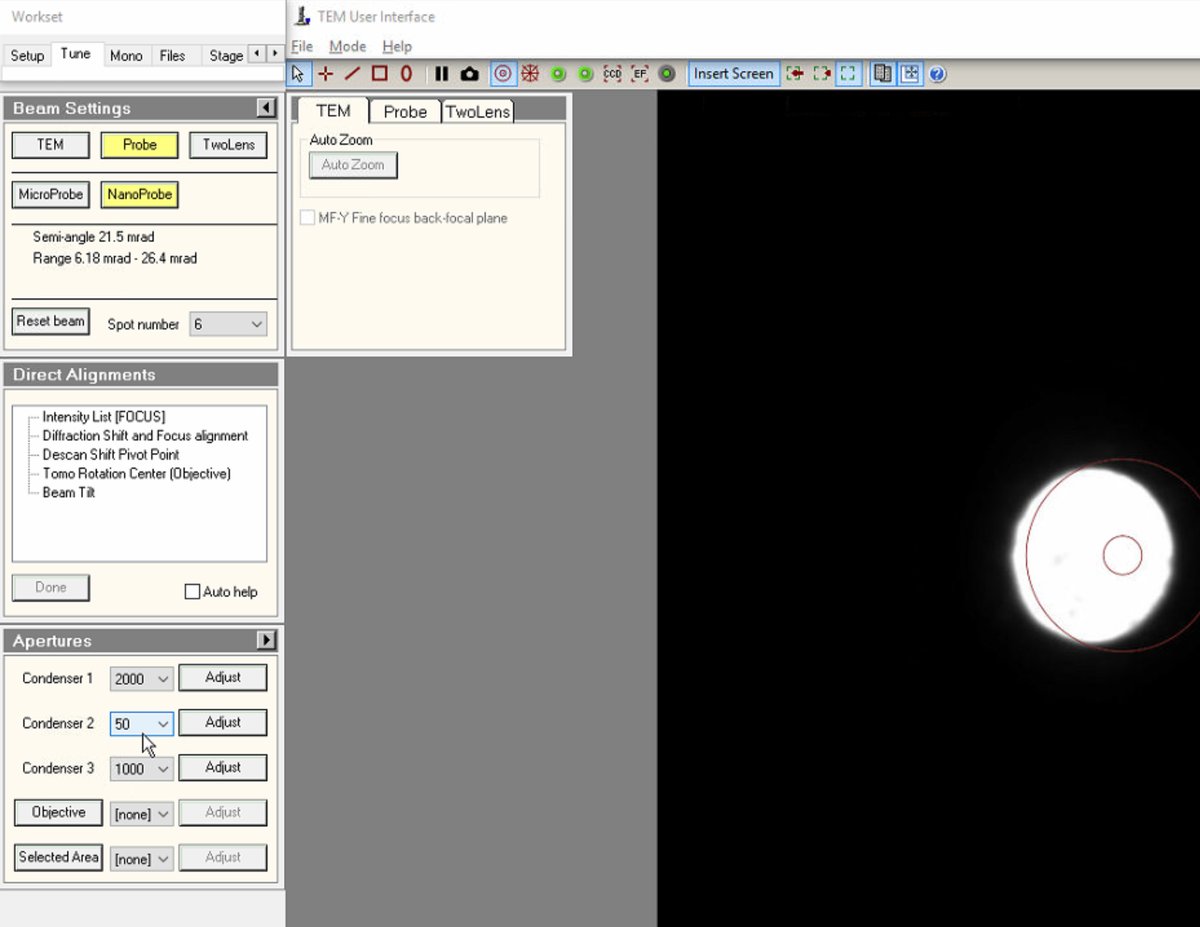
-
Click
Adjustto center the new aperture. The beam remains centered when changing aperture sizes. If not centered, use the adjustment controls to re-center.
-
-
Open Sherpa
-
Open the
Sherpasoftware. Sherpa displays the HAADF image with a crosshair marker indicating the measurement region.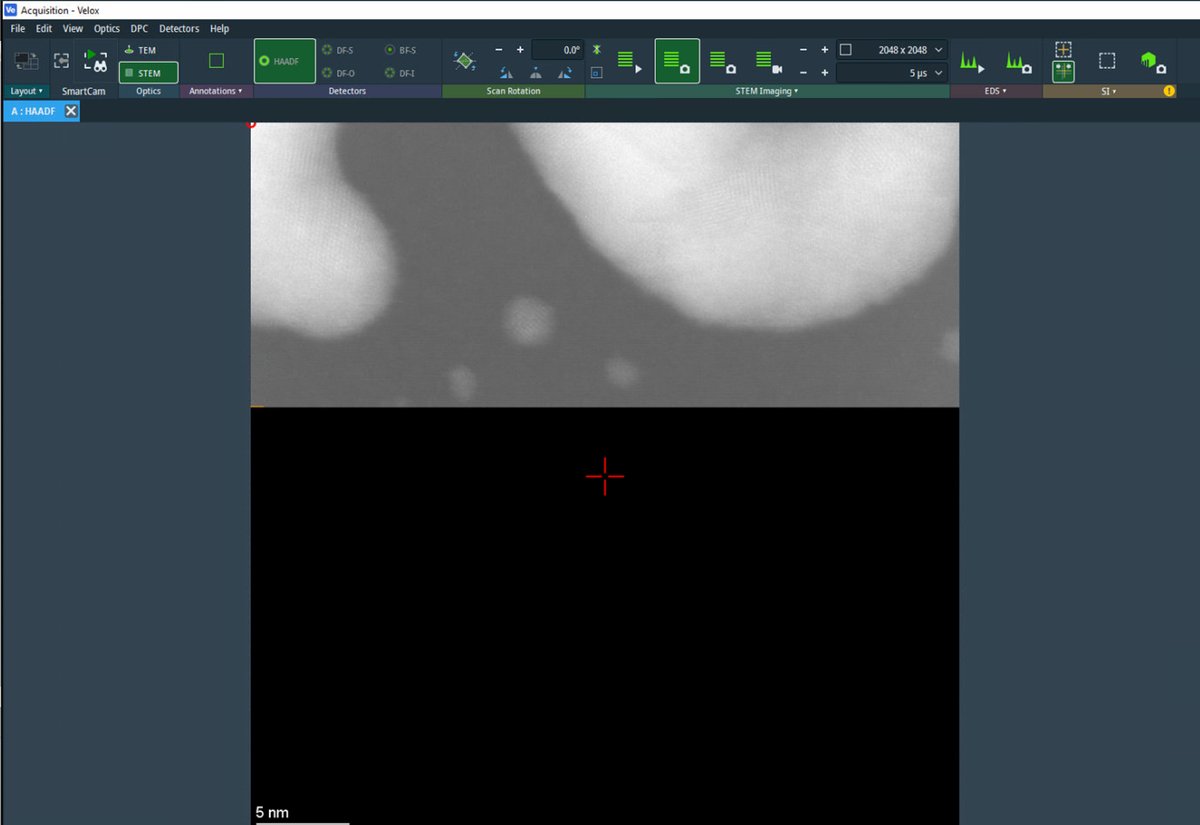
-
Click
C1/A1to run first-order correction (defocus and 2-fold astigmatism).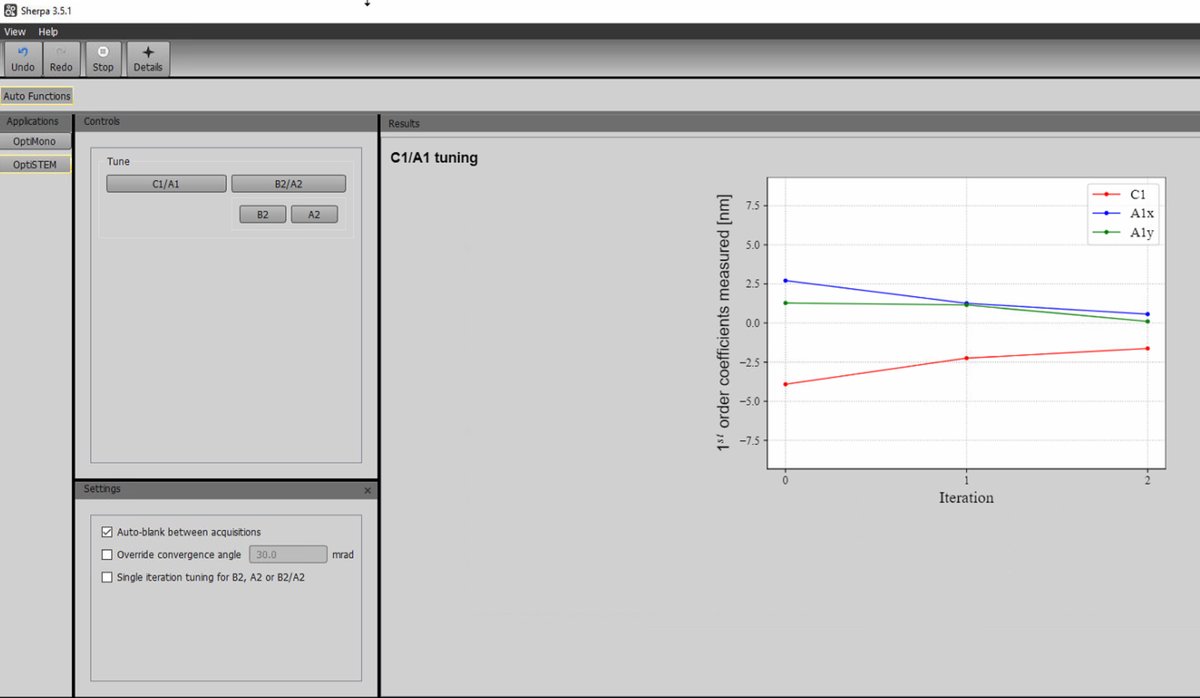
-
-
Run B2/A2 tuning
-
After C1/A1 completes, click the
B2/A2button to correct second-order aberrations (axial coma B2 and 3-fold astigmatism A2). -
Wait for the tuning to complete. Sherpa acquires images and determines optimal corrections.
-
Run multiple iterations if the first pass does not achieve optimal results.
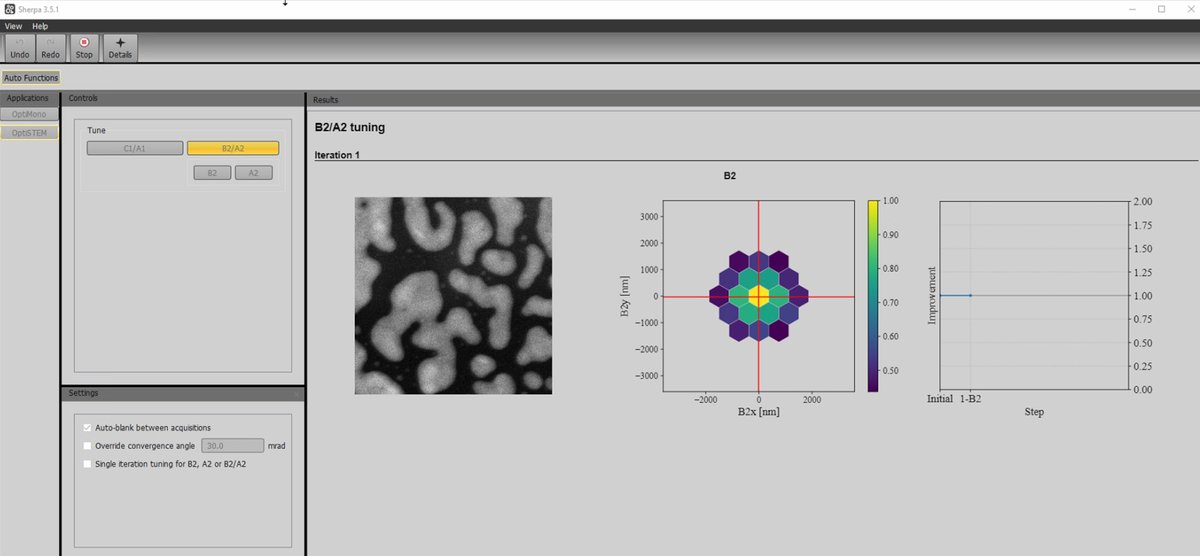
-
-
Review results
-
Sherpa displays the initial image alongside the optimized image for comparison. The corrected image shows improved sharpness and resolution.
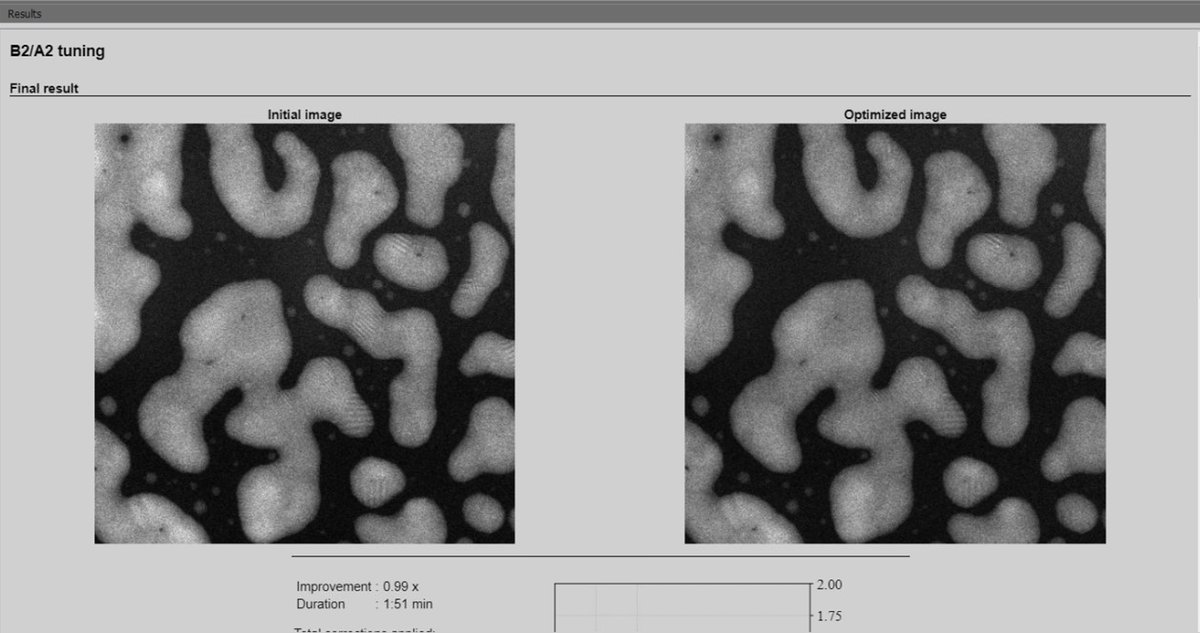
-
3.3 Load your own sample
After completing probe correction on the standard sample, follow the below steps unload the current sample standard and load your own. For holder-specific instructions (single-tilt, double-tilt, tomography), see Sample Loading.
-
Remove the standard sample
-
Put on gloves before handling any holders or samples.
-
Blank the beam and verify the screen is inserted. The screen protects the detectors and cameras below from the beam.
-
Close the column valves by pressing
Column Valves Closed.If a “VCP” error occurs, follow the instructions on the Spectra NEMO page.
-
Reset the holder by clicking
reseton theStagemenu.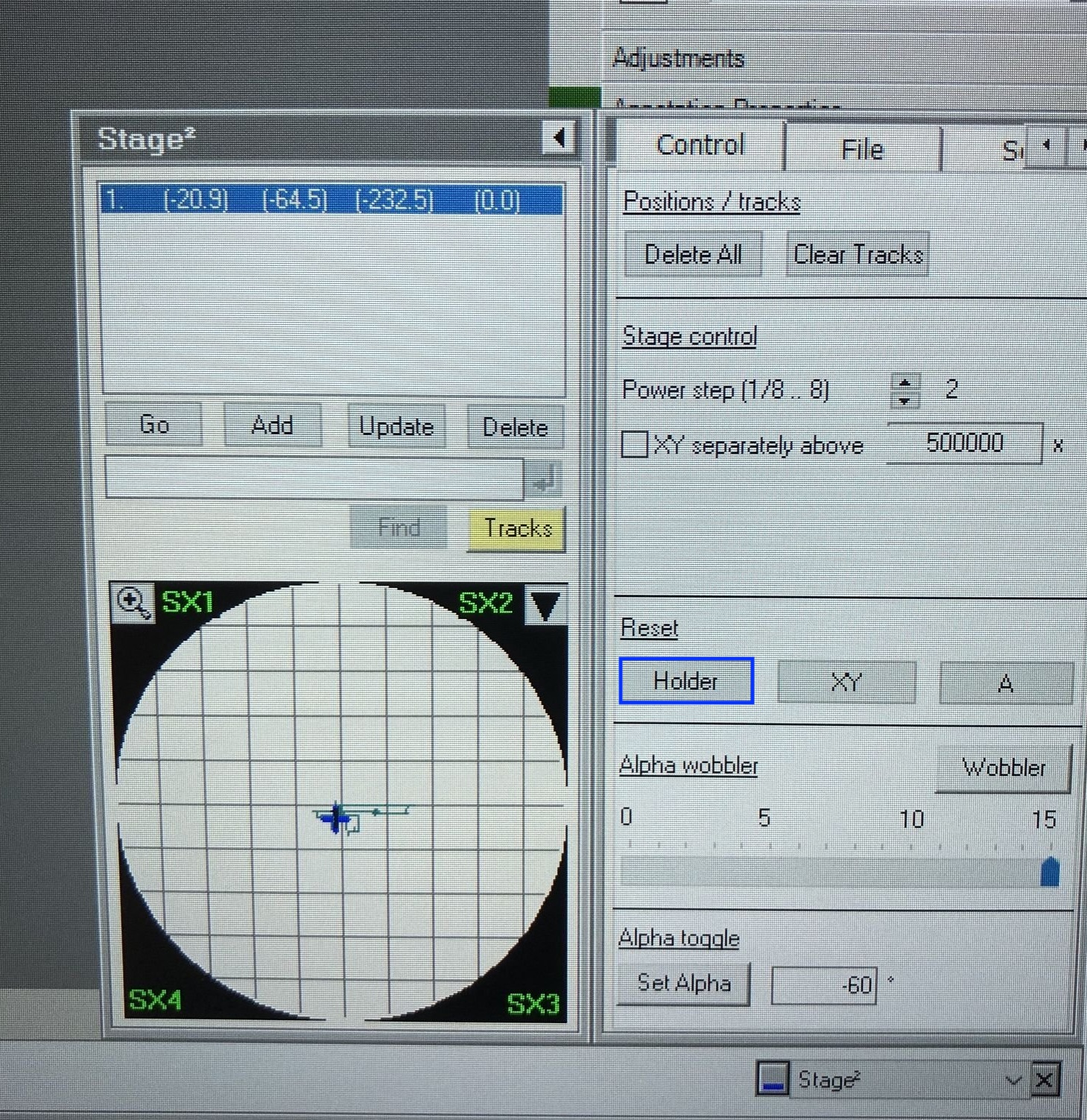
-
Confirm the stage x, y, z values are returning to zero after you reset the holder stage.
-
Pull the holder straight out to the first resistance point. Do not force beyond this point. Turn clockwise, then pull the rest of the holder out continuously.
-
-
Load your sample and insert the holder
IMPORTANT: Do not remove the standard sample from the single-tilt holder. Use a separate holder to load your sample.
-
For holder-specific loading instructions, see Sample Loading.
-
Align the holder with the blue line on the goniometer.
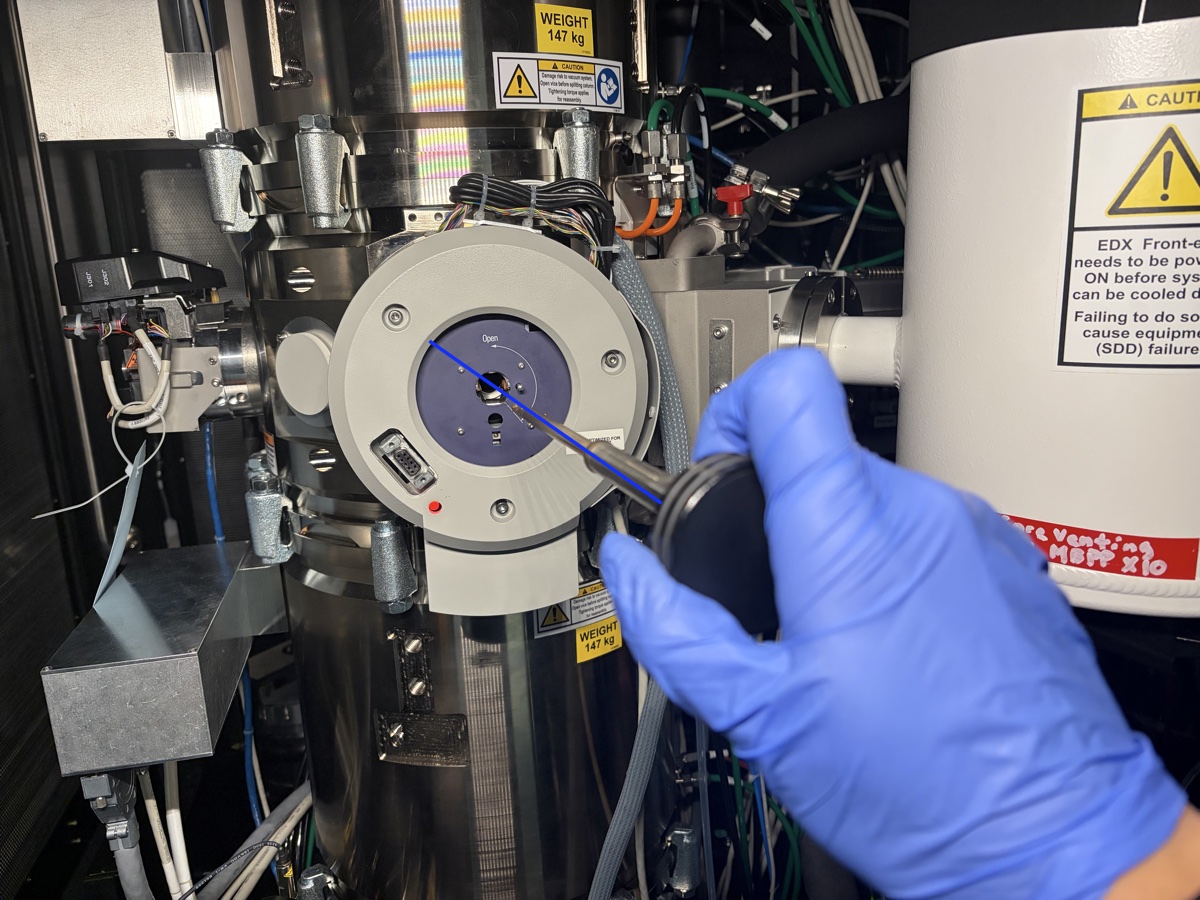

-
Push the holder in until you feel resistance. Do not push all the way in.
-
The turbo pump starts automatically. Wait ~2 minutes for pressure to stabilize. You can monitor the time in
TEMUIor on the screen attached to the Spectra instrument.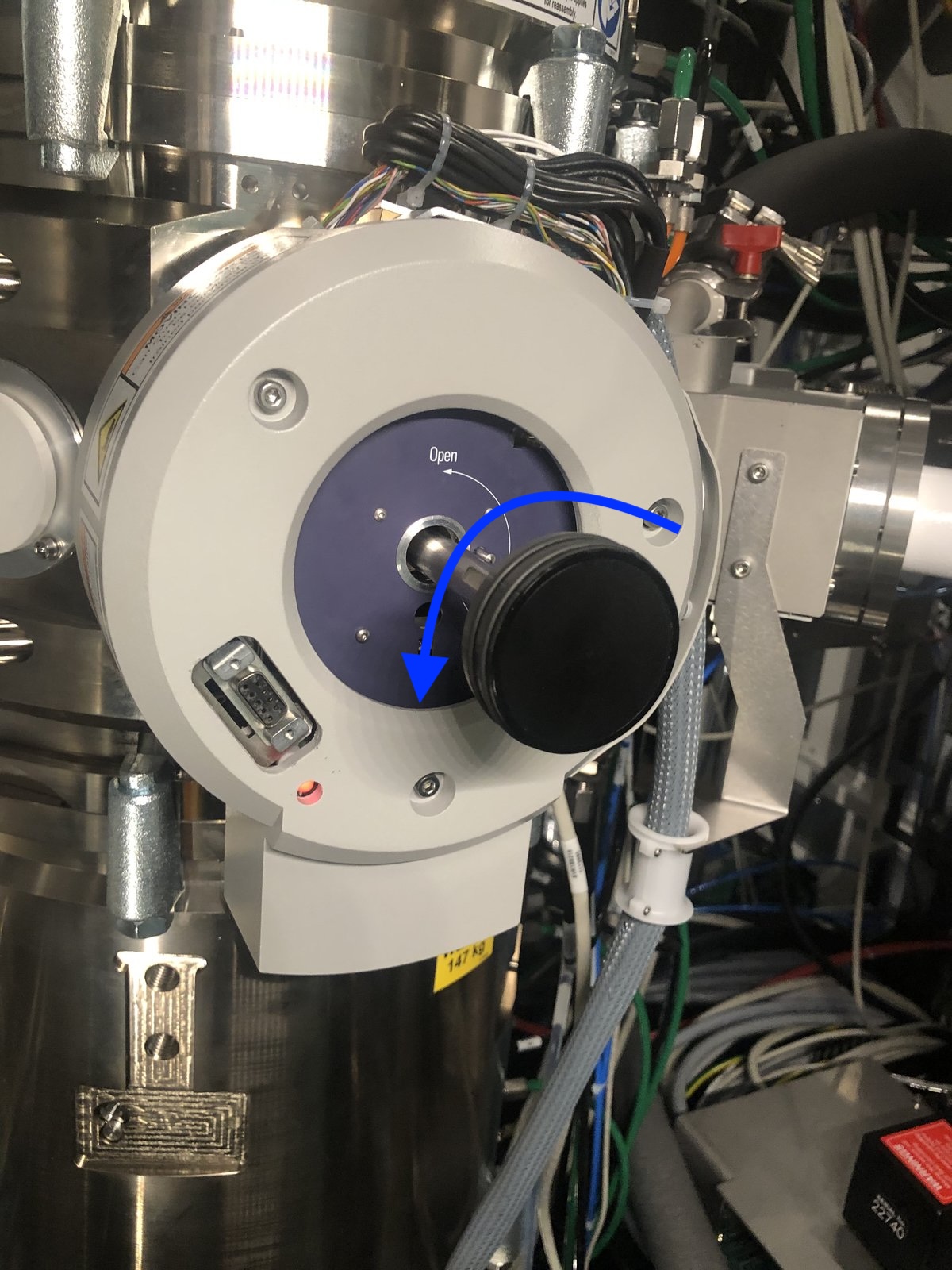
Why wait? The holder insertion opens a small chamber to atmosphere. The turbo pump must evacuate this air before you can insert the holder into the main column. Rushing this step would introduce air into the ultra-high vacuum column, potentially damaging the electron gun and contaminating the system.
-
Turn the holder counter-clockwise until you feel gently stuck, then guide the holder to push in. The holder should move in smoothly.
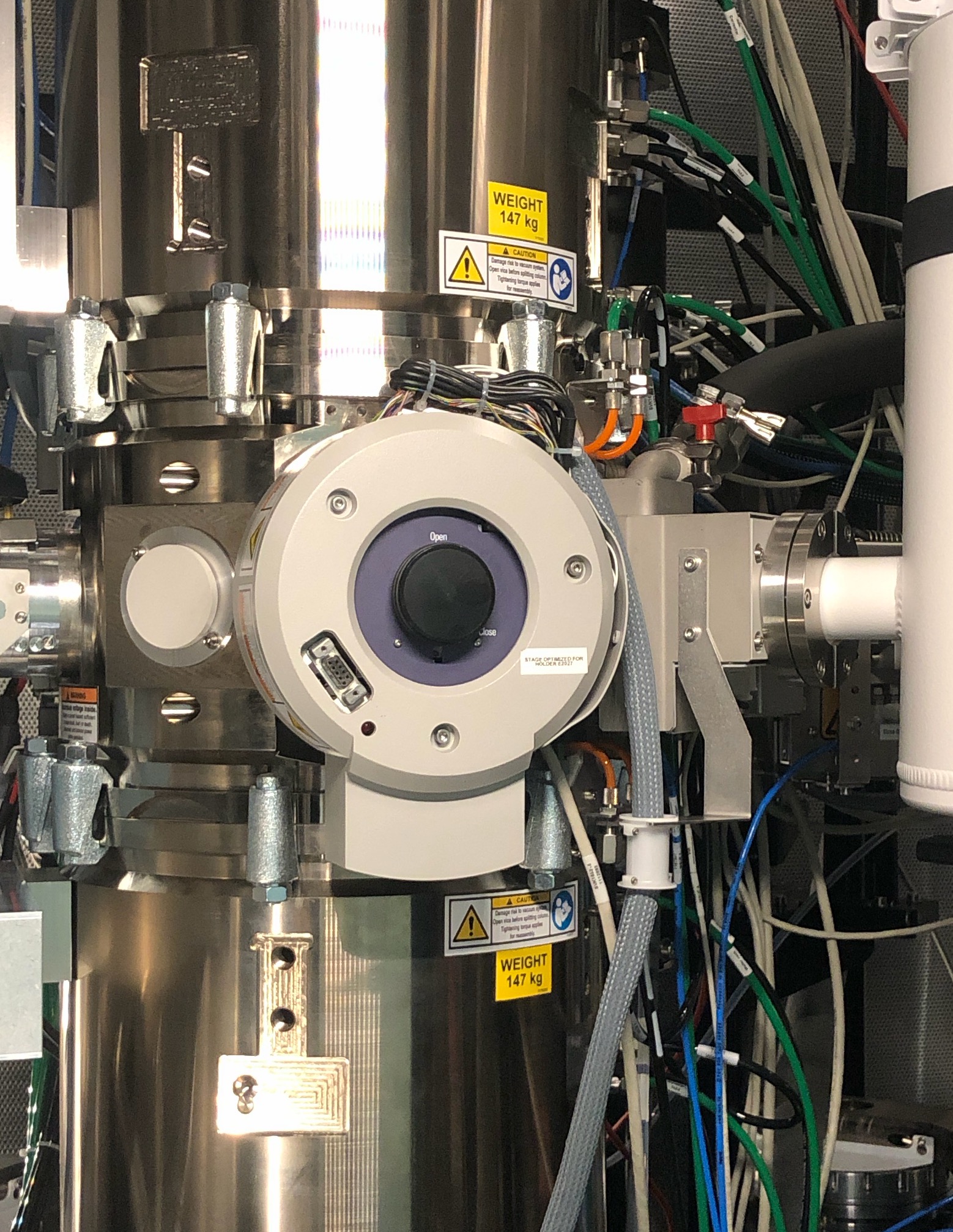

-
In
TEMUI, turn off the turbo pump. Confirm the holder type when prompted.
-
Re-do eucentric height
- Open column valves and re-do eucentric height for your new sample (1.2). Each sample sits at a different physical height in the holder. Find the ronchigram “blow-up” point again so the sample stays centered when tilted and the probe is properly focused.
- Run a quick C1A1 or Sherpa to verify probe correction still holds after the sample change.
3.4 End session
-
Reload the standard sample
-
Put on gloves before handling any holders or samples.
-
Blank the beam and verify the screen is inserted.
-
In
TEMUI, clickColumn Valves Closed. -
Click
Reset Holderunder theStagemenu. Visually verify that the X, Y, and Z stage coordinates are reset after the button is pressed.
-
Pull the holder with your sample straight out to the first resistance point. Do not force beyond this point. Turn clockwise, then pull the rest of the holder out continuously.
-
Set aside your holder and pick up the single-tilt holder with the standard sample.
-
Push the single-tilt holder with the standard sample in until you feel resistance. Do not push all the way in.
-
The turbo pump starts automatically. Wait ~2 minutes for pressure to stabilize.
-
Turn the holder counter-clockwise until you feel gently stuck, then guide the holder to push in.
-
In
TEMUI, turn off the turbo pump. ConfirmSingle tilton TEMUI.
-
Checklist before leaving the Spectra room
- Beam is blanked.
-
Reset Holderhas been pressed and X, Y, Z stage coordinates are verified reset. - Stage is returned to 0° tilt (alpha and beta).
- Arina detector is retracted, verified on the left hand panel.
- Arina detector is turned off, verified on the local Firefox URL.
-
INT SCANphysical button is in pressed state. - Screen is inserted.
- Column valve is closed.
- Turbo pump is turned off.
- Standard sample is loaded.
- All holders are capped and stored in the holder box.
- The sample loading area is tidy.
- Spectra usage is terminated on NEMO.
- Internet accounts (Google, Outlook, etc.) are signed off.
- Fill out the logbook if anything unusual happened during the session.
Troubleshooting
Common problems encountered during STEM sessions.
| Problem | Cause | Solution |
|---|---|---|
| Image drifts when tilting | Eucentric height not set | Re-do eucentric height after loading a new sample |
| C1A1 measurements unstable or fail | Velox is still scanning | Stop live scanning in Velox before running C1A1, then verify the beam is unblanked |
| Aberration values oscillate instead of converging | Overcorrection percentage too high | Start with 100% Auto correct, reduce to 75% as values approach target |
| C1A1 or Tableau shows no signal | Beam is blanked | Click Beam Blank button to unblank before running aberration measurements |
| Good Tableau values but poor image resolution | Missing C1A1 verification step | After Tableau, always run C1A1 again to fine-tune defocus and astigmatism |
| Beam disappears from view | Random adjustments displaced the beam | Go to lower magnification until beam is visible, use joystick to move sample to center, then go to Diffraction Shift and use mulXY to center the beam |
| Lost beam or need to redo alignment | Column misalignment after extended session | Redo eucentric height (1.2) and monochromator tune (1.5). If you cannot find the sample, switch to TEM mode for easier navigation (TEM Spectra) |
FAQ
Beam blanking
When the beam is blanked, the electron beam is deflected away from the sample so no electrons hit it. This prevents unnecessary radiation damage to the sample when not actively imaging. The beam is automatically blanked when scanning stops or after taking a picture. Manual blank/unblank is available via the Beam Blank button on the hand panel or in the software.
Monochromator focus adjustment
The monochromator filters the energy spread of the electron beam by passing it through a narrow slit. Setting Focus = 0 places the beam crossover exactly at the monochromator slit plane. This position maximizes electron throughput while maintaining energy filtering. If the focus is offset from zero, the beam crossover occurs before or after the slit, reducing beam current and degrading energy resolution.
Appendix
Aberration notation
Different notations exist for aberrations in the literature. The table below shows the Krivanek notation (used in Probe Corrector software), the alternative notation (used in this guide), and descriptions.
| Krivanek | Alt | Description | Krivanek | Alt | Description |
|---|---|---|---|---|---|
| \(C_{10}\) | \(C_1\) | Defocus | \(C_{41}\) | \(B_4\) | 4th order coma |
| \(C_{12}\) | \(A_1\) | 2-fold astigmatism | \(C_{43}\) | \(D_4\) | 3-lobe aberration |
| \(C_{21}\) | \(B_2\) | Axial coma | \(C_{45}\) | \(A_4\) | 5-fold astigmatism |
| \(C_{23}\) | \(A_2\) | 3-fold astigmatism | \(C_{50}\) | \(C_5\) | 5th order spherical |
| \(C_{30}\) | \(C_3\)/\(C_s\) | Spherical | \(C_{52}\) | \(S_5\) | 5th order star |
| \(C_{32}\) | \(S_3\) | Star aberration | \(C_{54}\) | \(R_5\) | Rosette |
| \(C_{34}\) | \(A_3\) | 4-fold astigmatism | \(C_{56}\) | \(A_5\) | 6-fold astigmatism |
Changelog
- Mar 1, 2026 - Add pre-session checklist, sample loading section with glove requirement, end session with explicit reload steps, fix image paths
- Feb 28, 2026 - Add prerequisite link to TEM Alignment guide; add lost-beam troubleshooting
- Jan 31, 2026 - Initial draft: instructions by Sangjoon Bob Lee, screenshots by Andrew Barnum during Spectra 300 hands-on training
TEM (Spectra)
This guide covers optional TEM alignment on the Spectra 300: column setup, eucentric height, aperture alignment, and image correction. TEM mode is useful for fast sample navigation and for users who need TEM-specific data (HRTEM, diffraction patterns). Most users will proceed directly to STEM (Spectra).

Prerequisite: The sample is already loaded and the holder is inserted into the Spectra 300. For sample loading and end session procedures, see STEM (Spectra).
Acronyms:
mulXY- Multifunction X/Y knobs on hand panelTEMUI- TEM User Interface (software)
Workstation layout:
| Monitor | Software | Purpose |
|---|---|---|
| Bottom left | TEMUI | Microscope control, vacuum, alignments |
| Bottom right | Velox | Live imaging, acquisition |
| Top left | ImageCorrector | Aberration measurement & correction |
| Top right | Velox image gallery | Captured images from Velox |
Overview
This guide covers two main phases:
| Phase | Procedures | Time |
|---|---|---|
| Part 1: Column alignment | Vacuum check, beam setup, eucentric height, monochromator, C2 aperture, condenser stigmatism, beam tilt, rotation center | 10-15 min |
| Part 2: Image correction | Capture image, C1A1 correction, Tableau measurement, save settings | 10-15 min |
Part 0: Safety check
Complete the pre-session checklist in STEM (Spectra) before proceeding. Do not skip this step.
Part 1: Column alignment
1.1 Open column valves
Before imaging, verify that the vacuum system is ready and the column valves can be safely opened.
-
Verify vacuum pressure
-
In
TEMUI, check the vacuum pressure values on the log scale (lower = better):Gauge Log Value Why Important Gun 1 Highest vacuum needed for stable electron emission Liner <10 Prevents electron scattering along beam path Octagon 1 Protects sample from contamination and oxidation Projection <30 Maintains image quality in projection system Buffer tank <50 Ensures stable pumping performance Backing line <80 Turbo pump pushes compressed gas into the backing line
-
-
Open column valves
-
In
TEMUI, clickCol Valves Open. The status changes to indicate the column valves are open and the turbo pump is off.
-
-
Set condenser apertures
-
In
TEMUI, go to theTunetab, thenApertures. Set Condenser 1, 2, 3 to 2000, 70, 1000.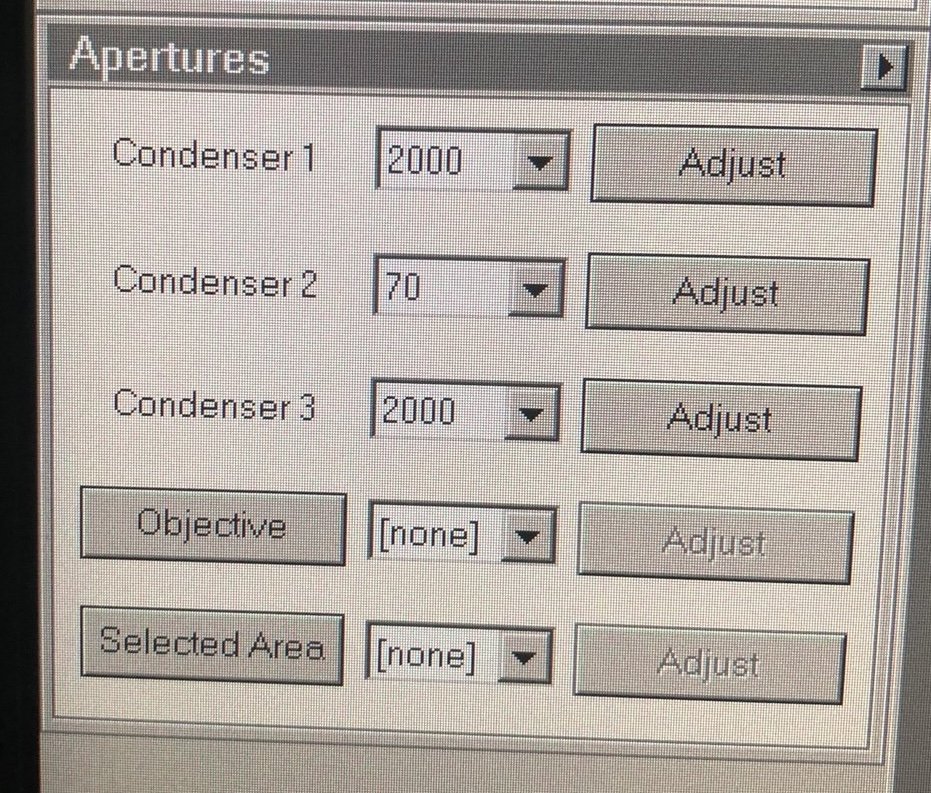
-
1.2 Beam setup
Configure the beam parameters for initial navigation and sample finding.
-
Enter TEM mode
- On the
Veloxsoftware (bottom right monitor), verify TEM mode is active. If not, click theTEMbutton.
- On the
-
Set spot size
- Set Spot Size 3 by pressing the
L3orR3button on the hand panel. As spot size decreases, screen current increases and the image gets brighter. - If the image is too bright, turn the intensity knob to decrease the screen current to around 2 nA and press
Linearmode to see better contrast.
- Set Spot Size 3 by pressing the
-
Find sample region
-
Set ~500x magnification by adjusting the magnification knob.
-
Locate the gold (dark) and amorphous carbon boundary by driving the joystick on the hand panel. This contrast boundary serves as a visual marker to identify the region of interest across various magnifications.
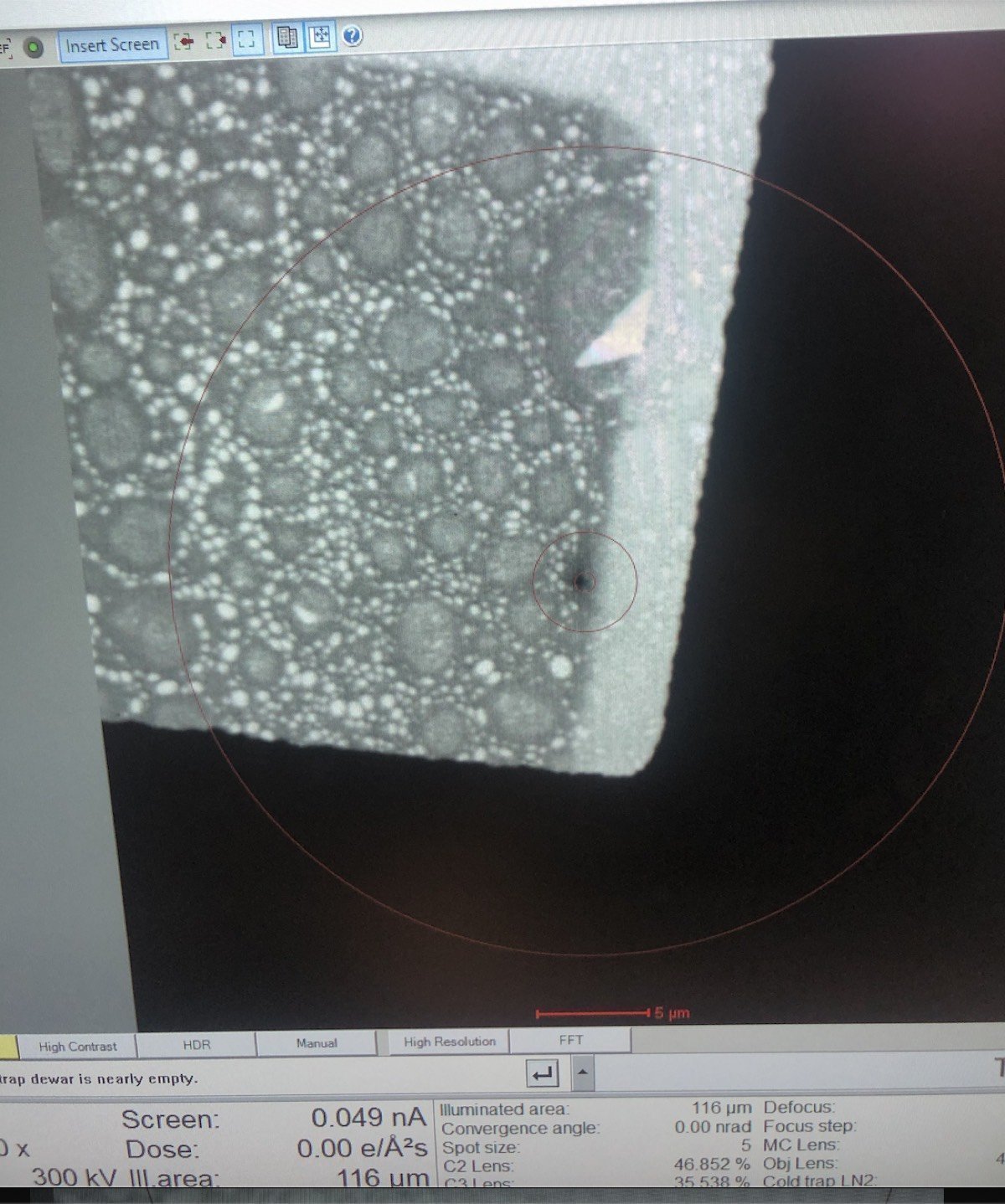
-
1.3 Eucentric height
At eucentric height, the sample remains stationary when tilted. This is essential for accurate imaging and aberration correction. Complete eucentric height alignment after loading each sample. Do not skip this step.
-
Adjust z-axis
-
Set ~7,500x magnification by adjusting the magnification knob.
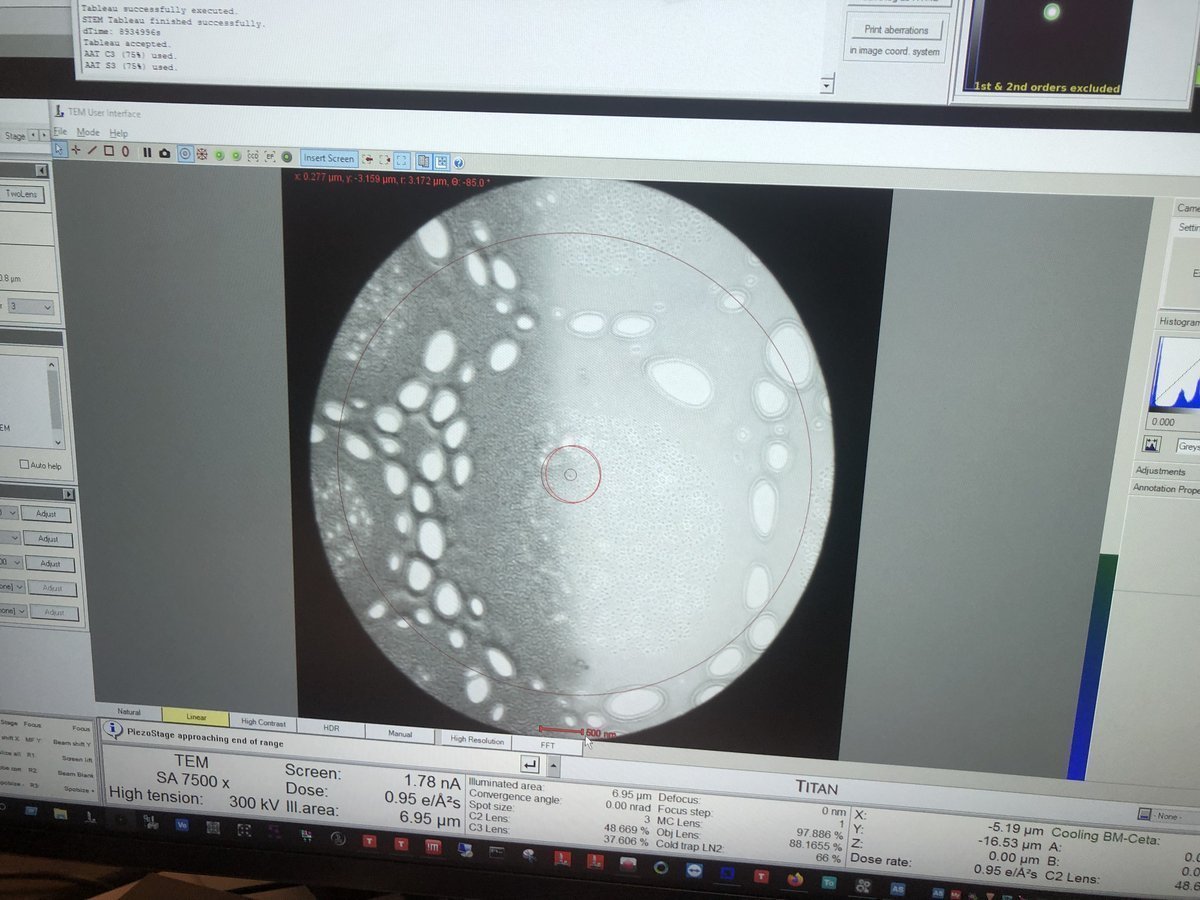
-
Press
z-axisup or down on the hand panel. Watch the image contrast change as the sample moves through focus. At eucentric height, the contrast is minimized (the image appears most “washed out”).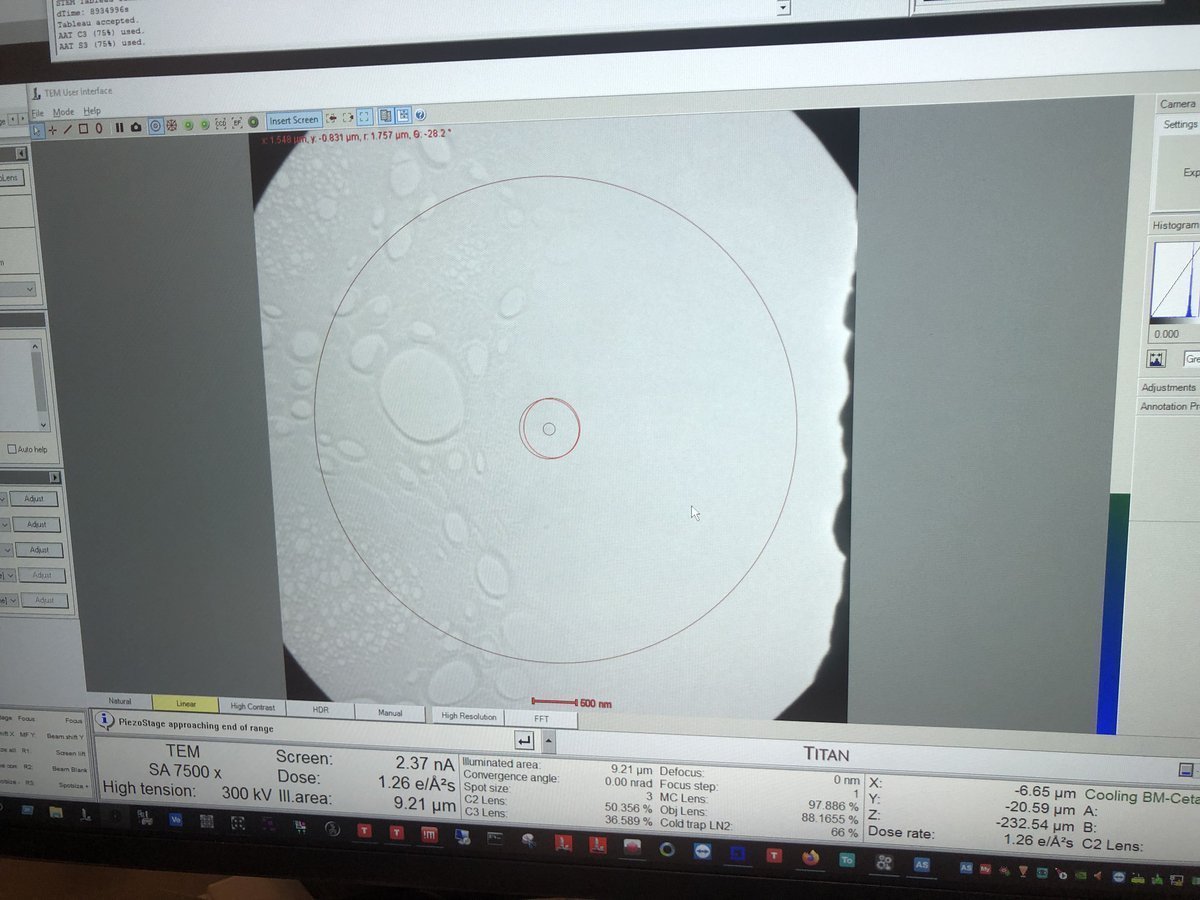
-
1.4 Monochromator tune
The monochromator selects a narrow energy range from the electron beam. If the beam edge looks jagged, the monochromator needs alignment.
-
Check beam edge
- Do you see a jagged area along the beam edge in the previous step? If not, skip this section. Otherwise, follow the steps below.
-
Adjust monochromator
-
In
TEMUI, go to theMonotab, then openMonochromator Tune (Expert)and clickShift.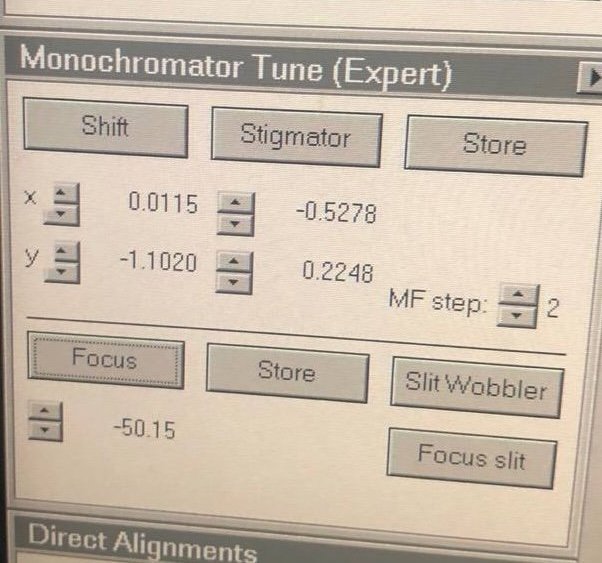
-
Adjust
mulXYknobs until the jagged area disappears.
-
1.5 C2 aperture alignment
The C2 aperture controls convergence angle and beam size. It blocks off-axis electrons — only electrons within a certain angular range pass through. A user must center this aperture on the optical axis so that the beam expands and contracts symmetrically.
-
Enter two-lens mode
-
In
TEMUI, go to theTunetab, thenBeam Settings, and clickTwolens. In two-lens mode, C3 is turned off, so the beam behavior on screen is purely from C2. This makes it straightforward to detect and correct any C2 aperture misalignment. In three-lens mode, C3 reshapes the beam after C2, masking the misalignment.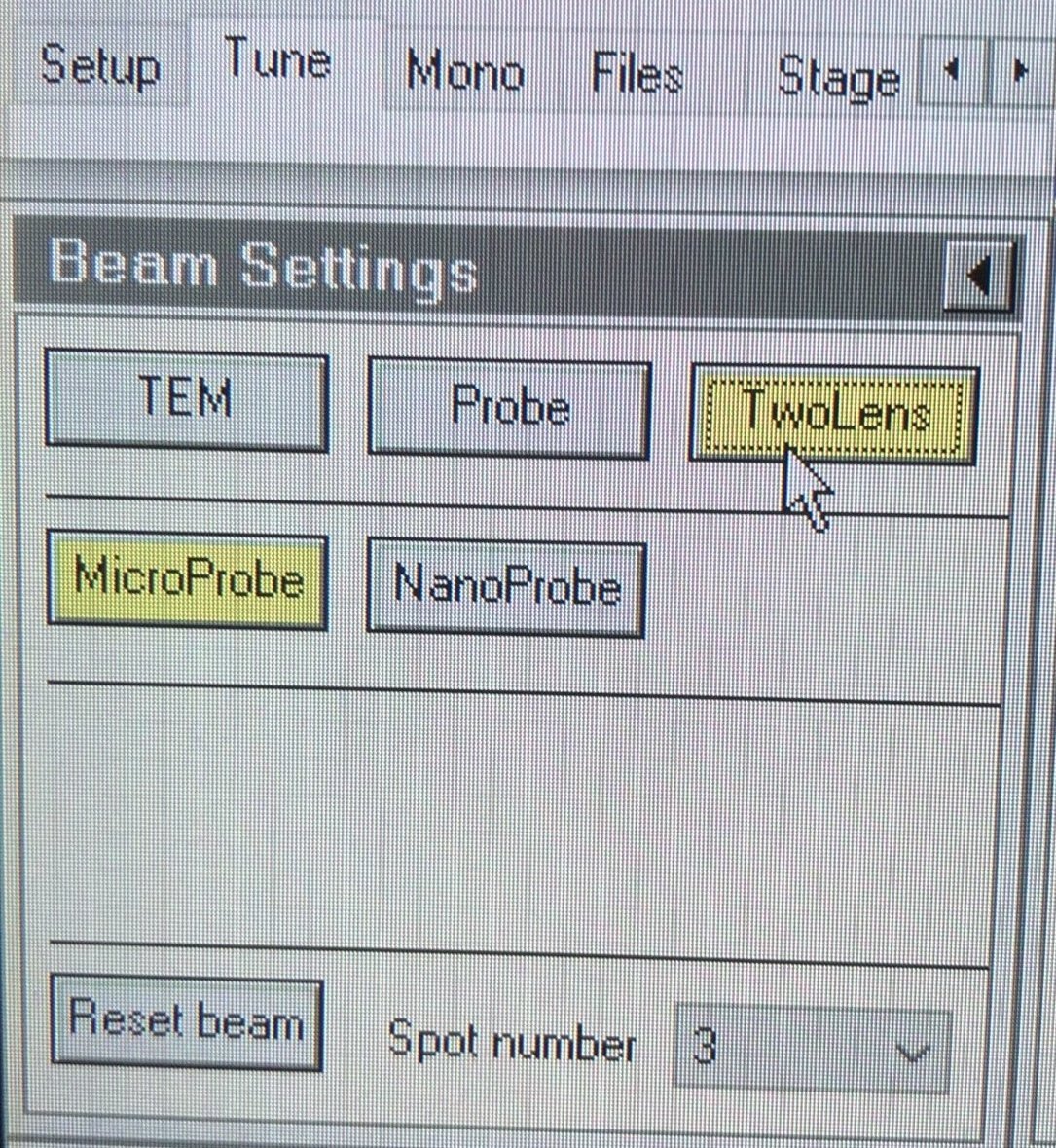
-
-
Center and align C2 aperture
-
Center the beam by rolling the hand panel ball.
-
Converge the beam by varying the intensity knob.
-
Vary beam size by turning the intensity knob counterclockwise and clockwise. Notice the beam expansion is not concentric — this indicates the C2 aperture is off-center.
-
Make the beam concentric: go to
Apertures, clickAdjustnext toCondenser 2, then adjust themulXYknobs until the beam expands and contracts concentrically.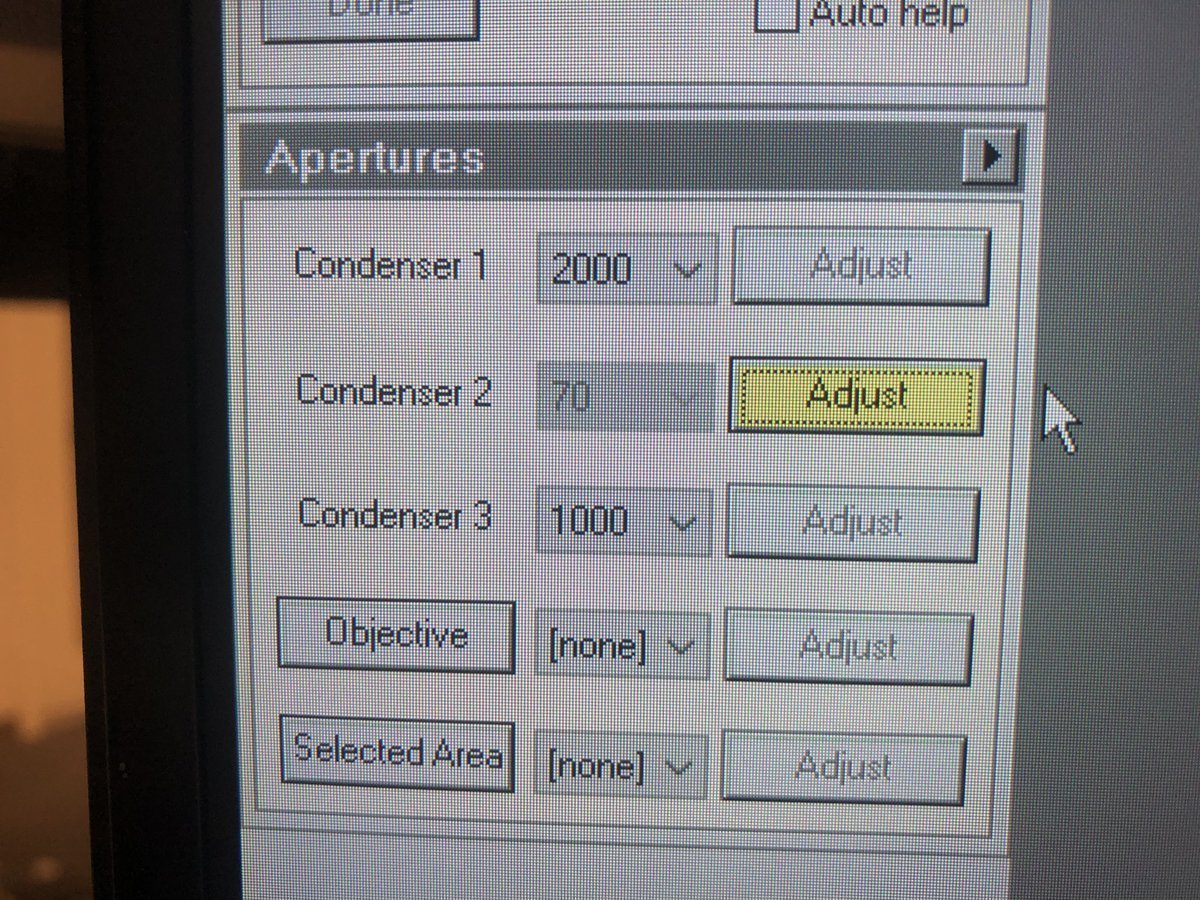
-
-
Return to three-lens mode
-
In
TEMUI, go toBeam Settingsand clickTEMto return to three-lens mode.
-
Verify the beam is centered and concentric.
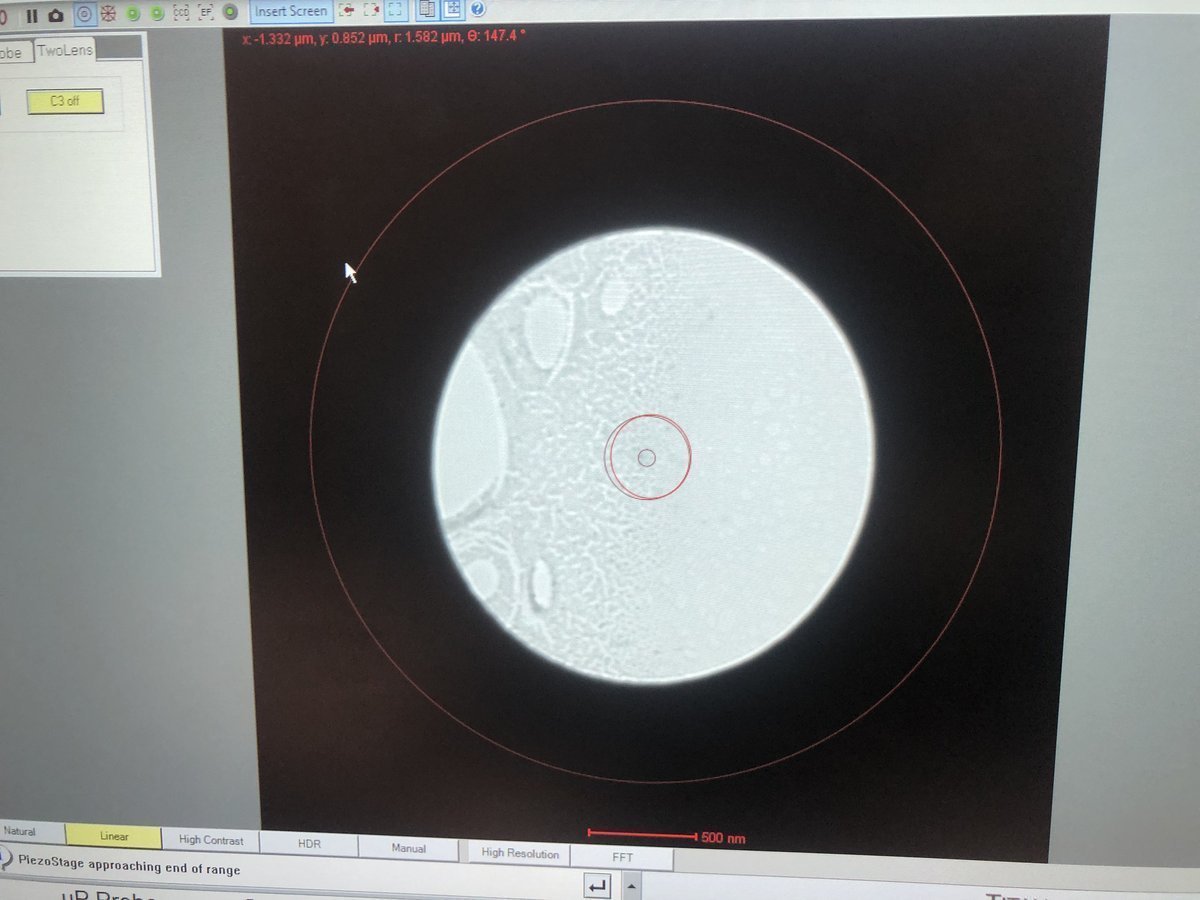
-
1.6 Condenser stigmatism
Condenser astigmatism causes the beam to appear elliptical instead of round. Correcting this ensures a symmetric probe.
-
Increase magnification
- Set ~200kx magnification by adjusting the magnification knob.
- If the beam has shifted from center, go to
Tunetab, thenDirect Alignment, clickBeam Shift, and adjust themulXYknobs to re-center.
-
Correct stigmatism
-
Enlarge the beam by adjusting the intensity knob.
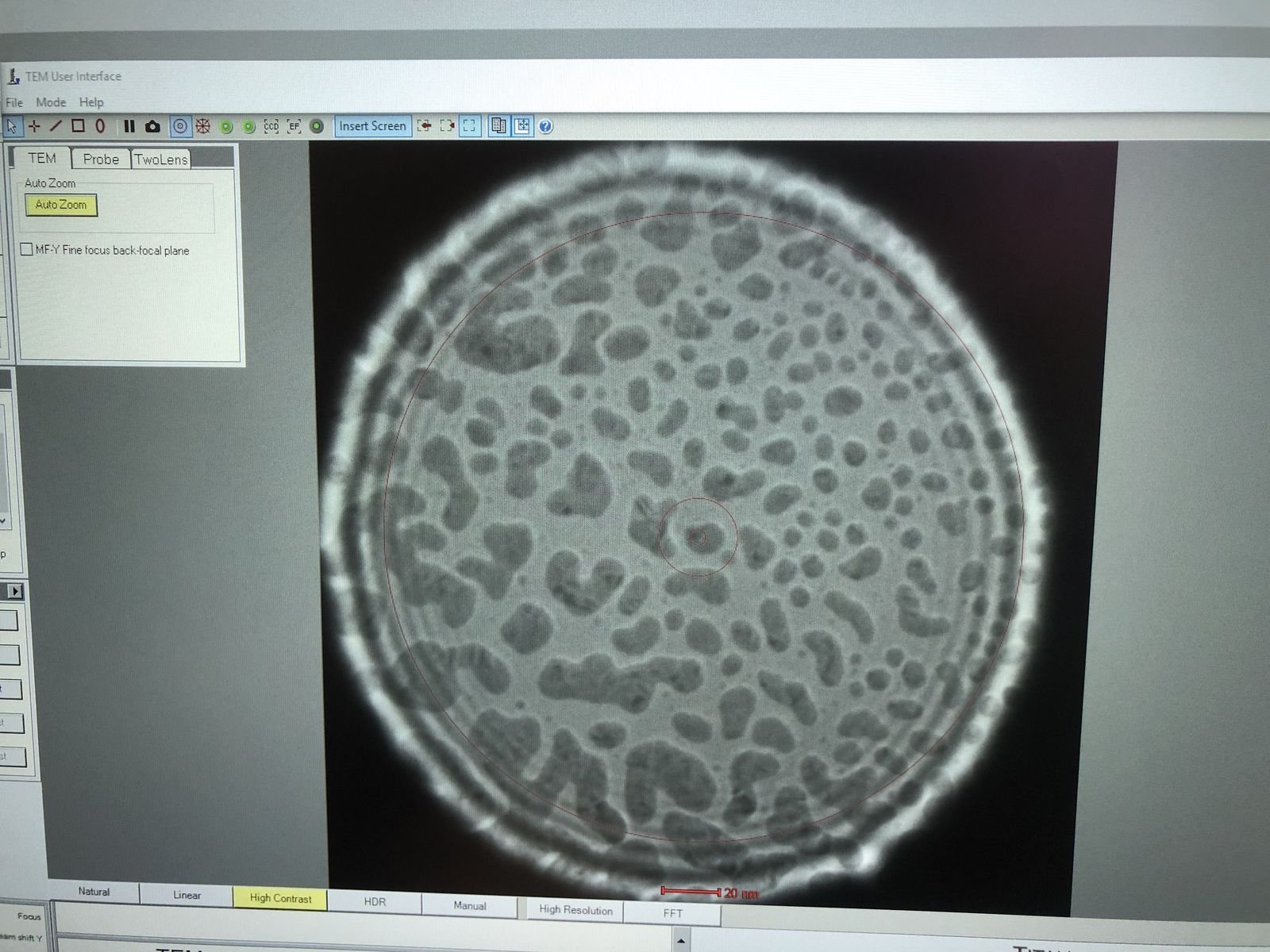
-
In
TEMUI, clickStigmator, thenCondenser. Adjust themulXYknobs to make the beam as round as possible. The beam should remain circular as you vary the intensity knob. PressNonewhen done.
-
1.7 Beam tilt
Beam tilt alignment minimizes lateral beam shift when the beam angle changes. Proper alignment ensures the beam tilts around a single point without drifting.
-
Align beam tilt
- In
TEMUI, go toDirect Alignmentand clickBeam tilt pp X. Adjust themulXYknobs to minimize the lateral jiggle. - Repeat for
Beam tilt pp Y. - If the beam center has shifted, click
Beam Shiftand adjust themulXYknobs to re-center.
- In
1.8 Rotation center
Rotation center alignment ensures the image rotates around the center of the field of view when focus changes.
-
Align rotation center
- In
TEMUI, go toDirect Alignmentand clickRotation Center. The image pulses in and out of focus. - Adjust the
mulXYknobs to minimize lateral movement. The pulsing should appear concentric (expanding and contracting from the same point) with no side-to-side drift.
- In
Part 2: Image correction
2.1 Capture image
Before running the image corrector, a user must set up live imaging in Velox and find a suitable sample region.
-
Prepare for imaging
- Find a flat area with a distribution of particle sizes and no holes.
- Important: Enlarge the beam to cover the entire fluorescent screen before lifting it. When the screen is raised, the camera and detectors below are exposed to the beam. A concentrated beam can permanently damage them.
- Press
R1on the hand panel to lift the fluorescent screen.
-
Start live imaging
-
In
Velox(right monitor), click the play button to start live imaging.
-
Do not change the intensity knob while the screen is lifted. The screen is lifted when
TEMUIshows a black display with dose reading “Unavail”. -
Gold nanoparticles should be visible on screen.
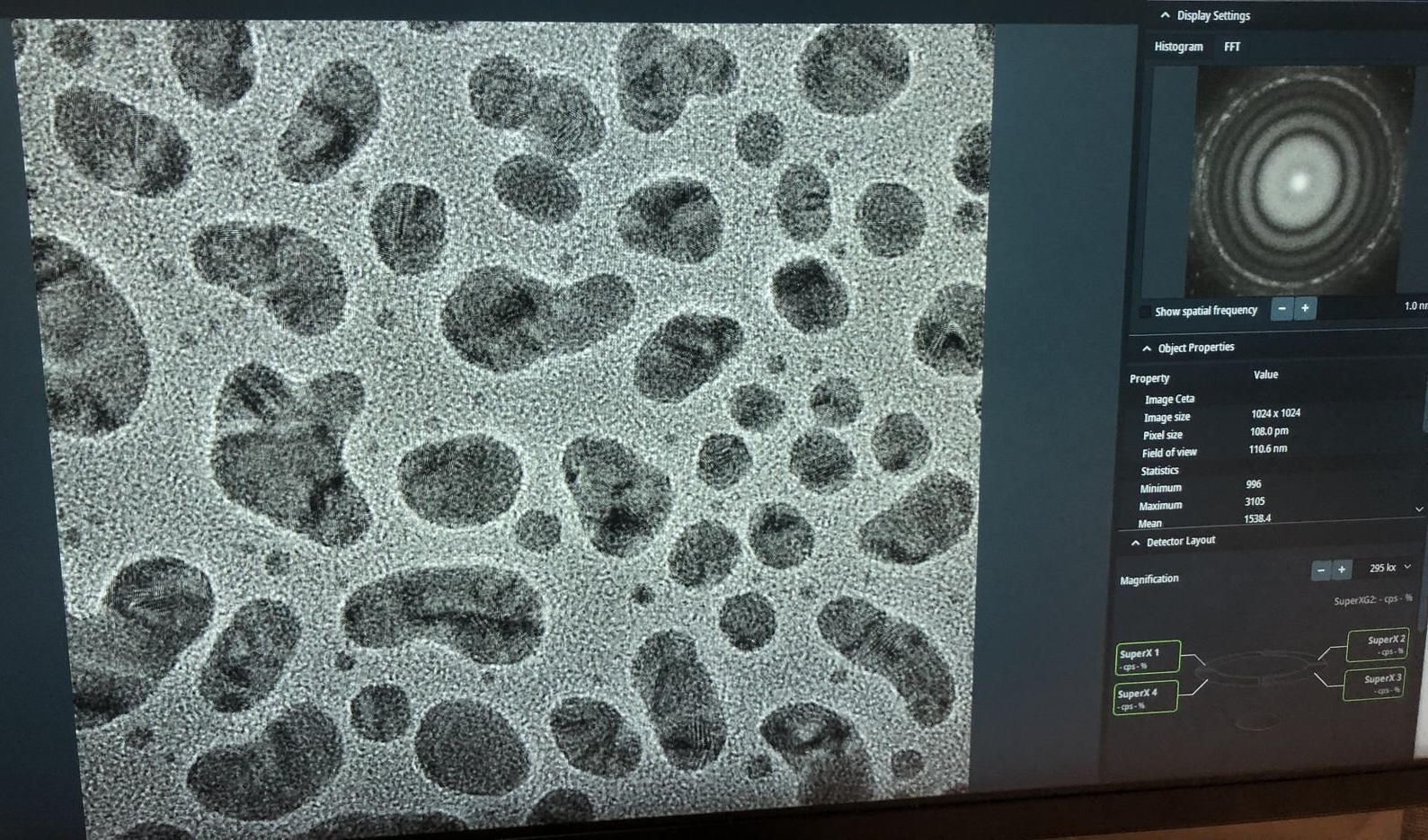
-
-
Explore focus (optional)
-
Press the
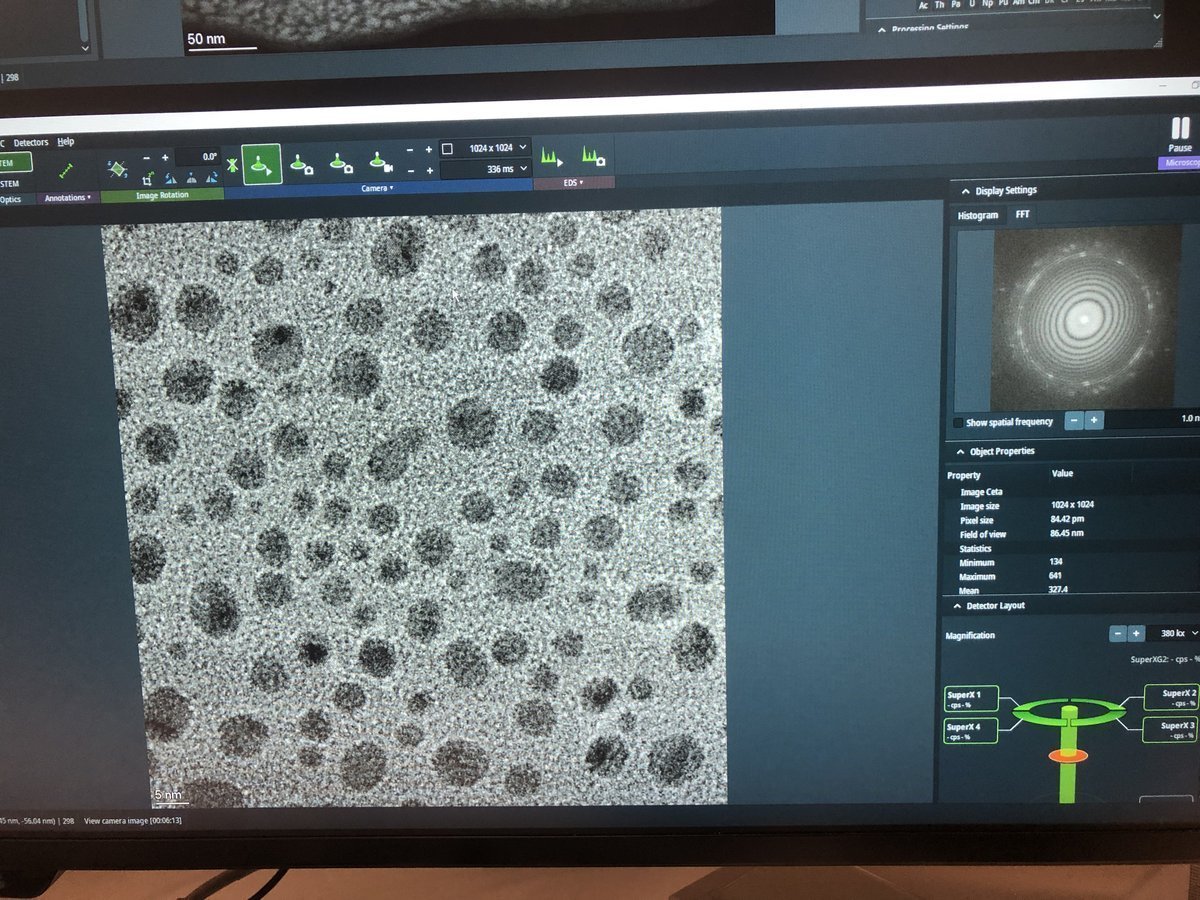
z-axisbuttons to observe how focus affects the image.Underfocus — edges appear bright with white Fresnel fringes:
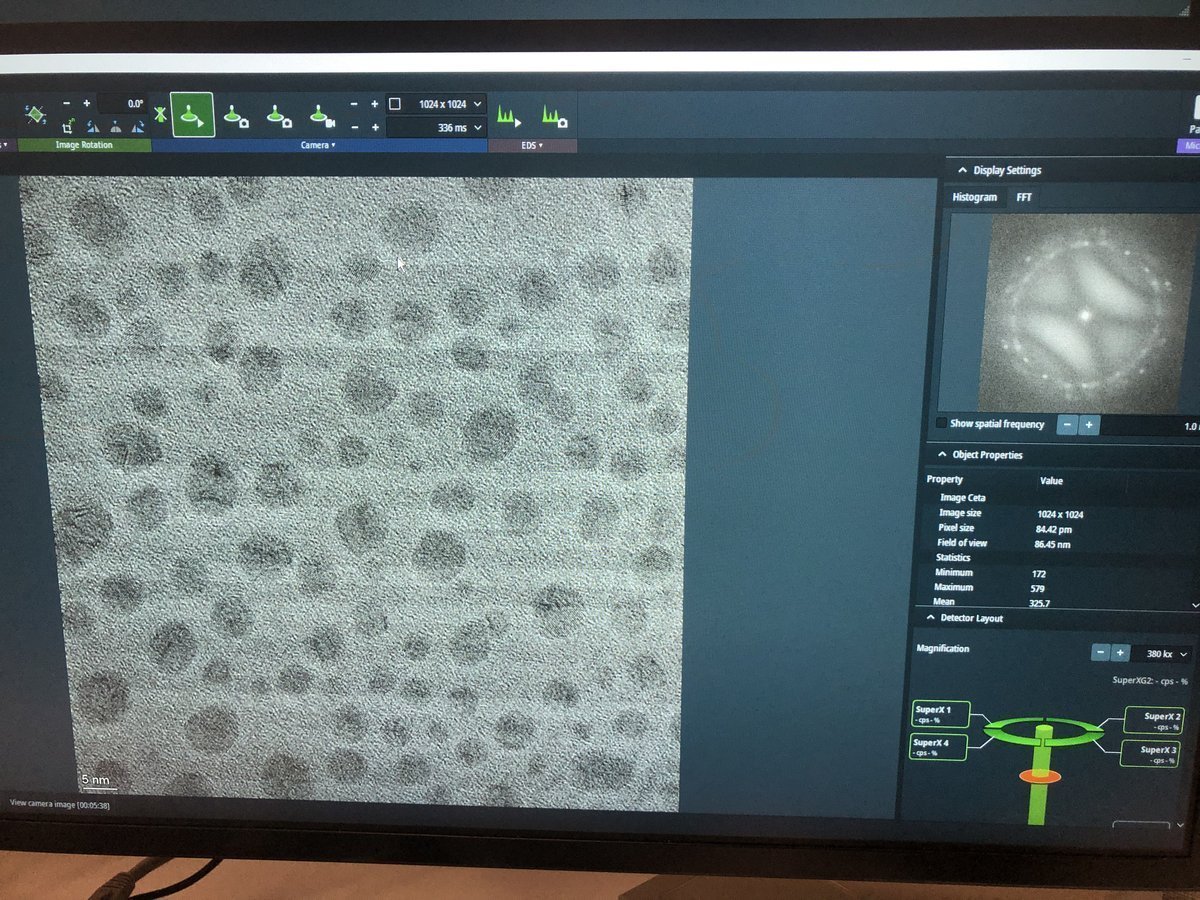
On focus — minimal fringe contrast:
Overfocus — contrast inverts, dark fringes at edges:
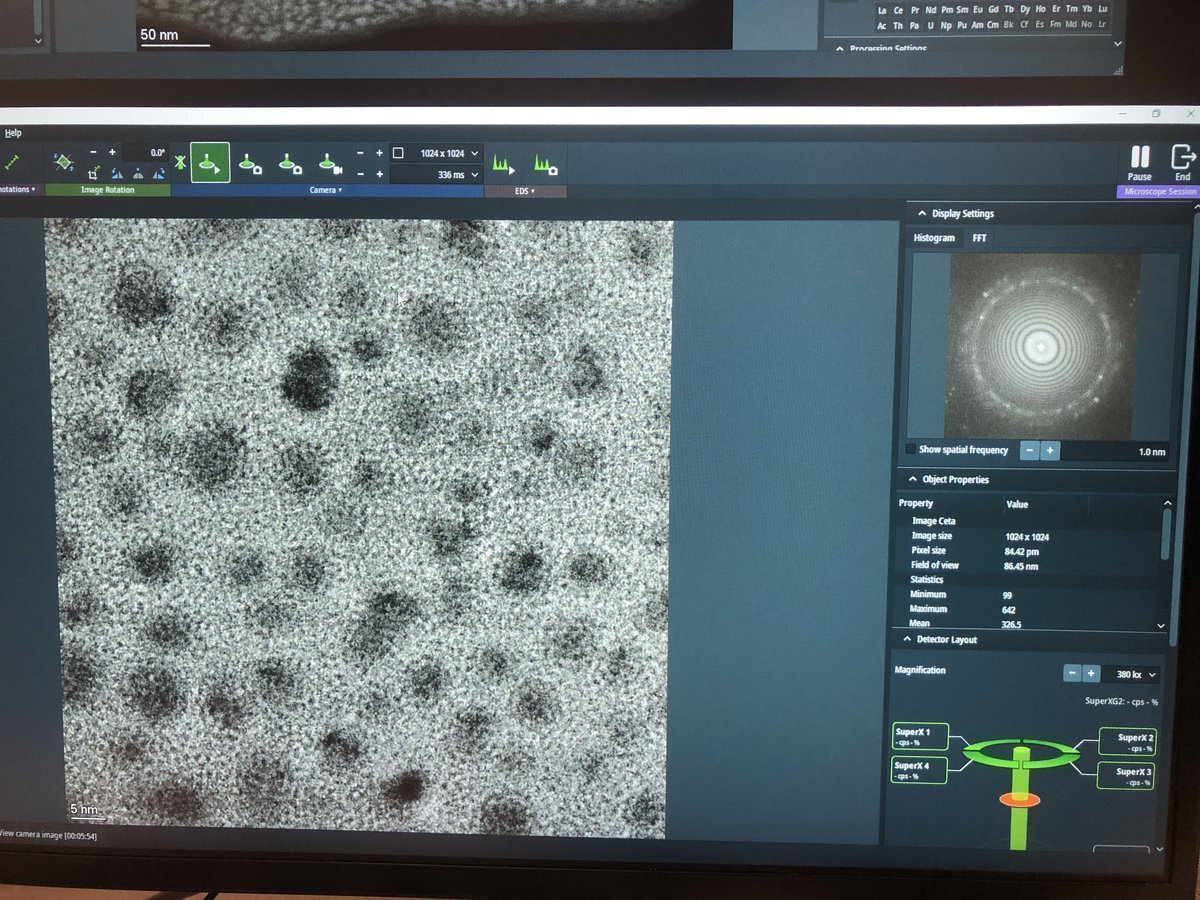
-
2.2 C1A1 correction
C1A1 corrects first-order aberrations in the image-forming lenses: defocus (C1) and 2-fold astigmatism (A1).
-
Set underfocus
-
Press
Z-axisdown until you see 4-5 rings in the FFT (slight underfocus). The rings indicate Thon rings from the amorphous carbon, which the corrector software uses for aberration measurement.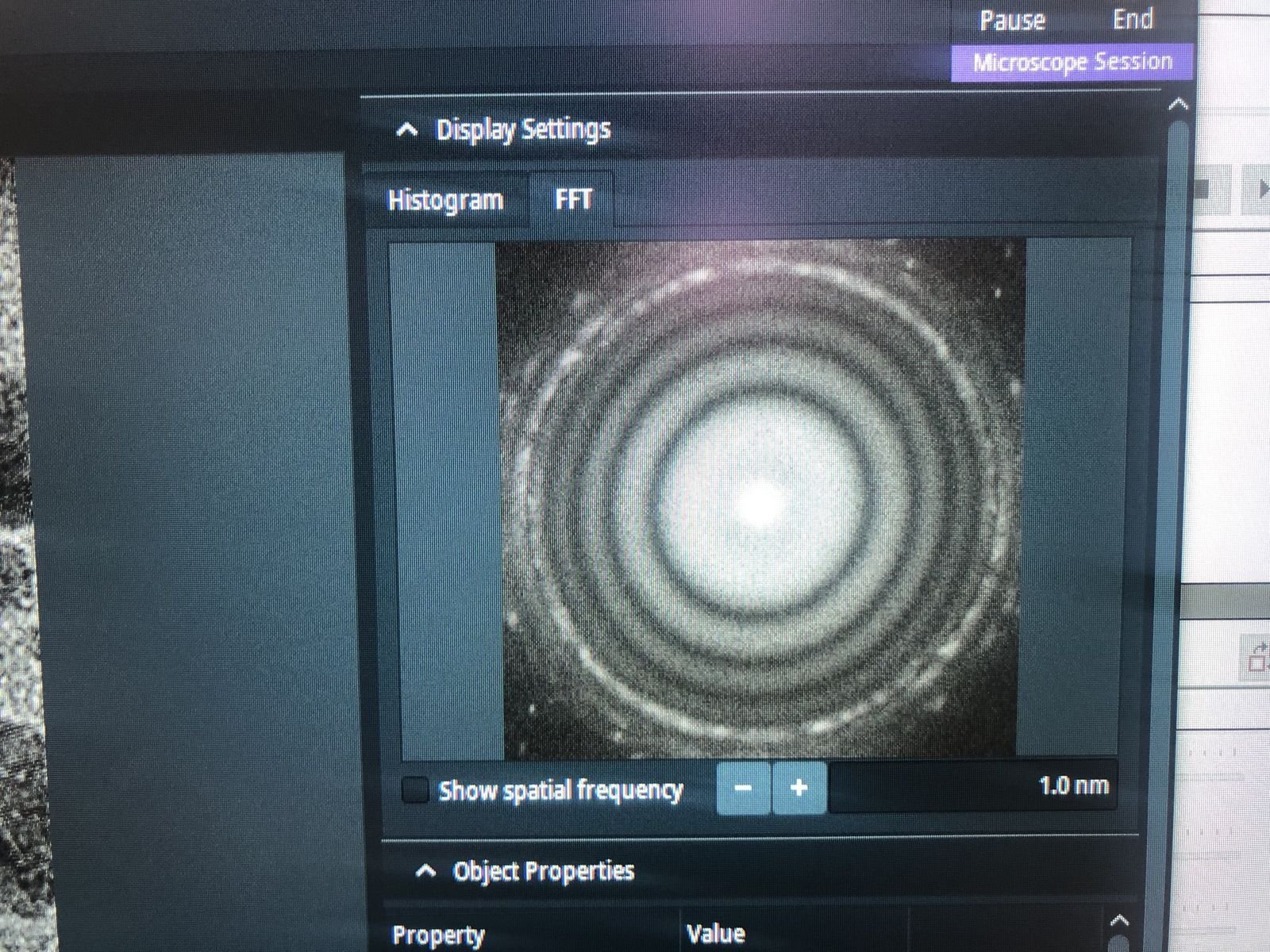
-
-
Reset stigmator values
-
Stop live scanning by clicking the play button in
Velox. -
In
TEMUI, go to theStigmatorquick tab. ResetObjectiveandImage A1to zero. If non-zero, right-click each button to reset, then clickDone.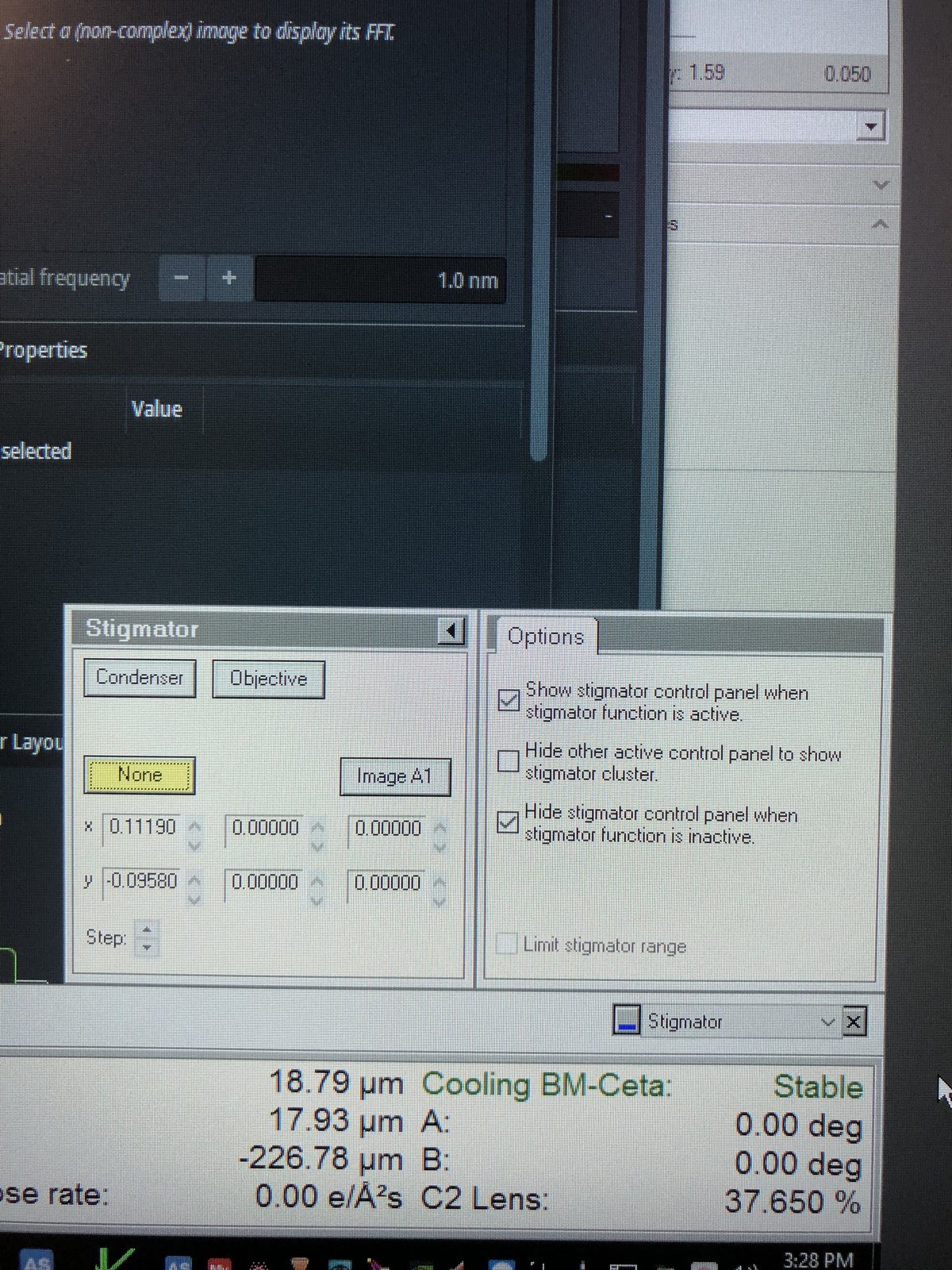
-
-
Run C1A1
-
Open the
ImageCorrectorsoftware (top left monitor). -
Set exposure time to 0.3s.
-
Go to the
C1A1tab and clickStart. The microscope wobbles the focus up and down (changing objective lens current). The FFT is captured and its ring symmetry, angular distribution, and ring spacing are analyzed. -
During the iteration, carefully set intensity to 800–900 counts by adjusting the intensity knob so the corrector has enough signal.
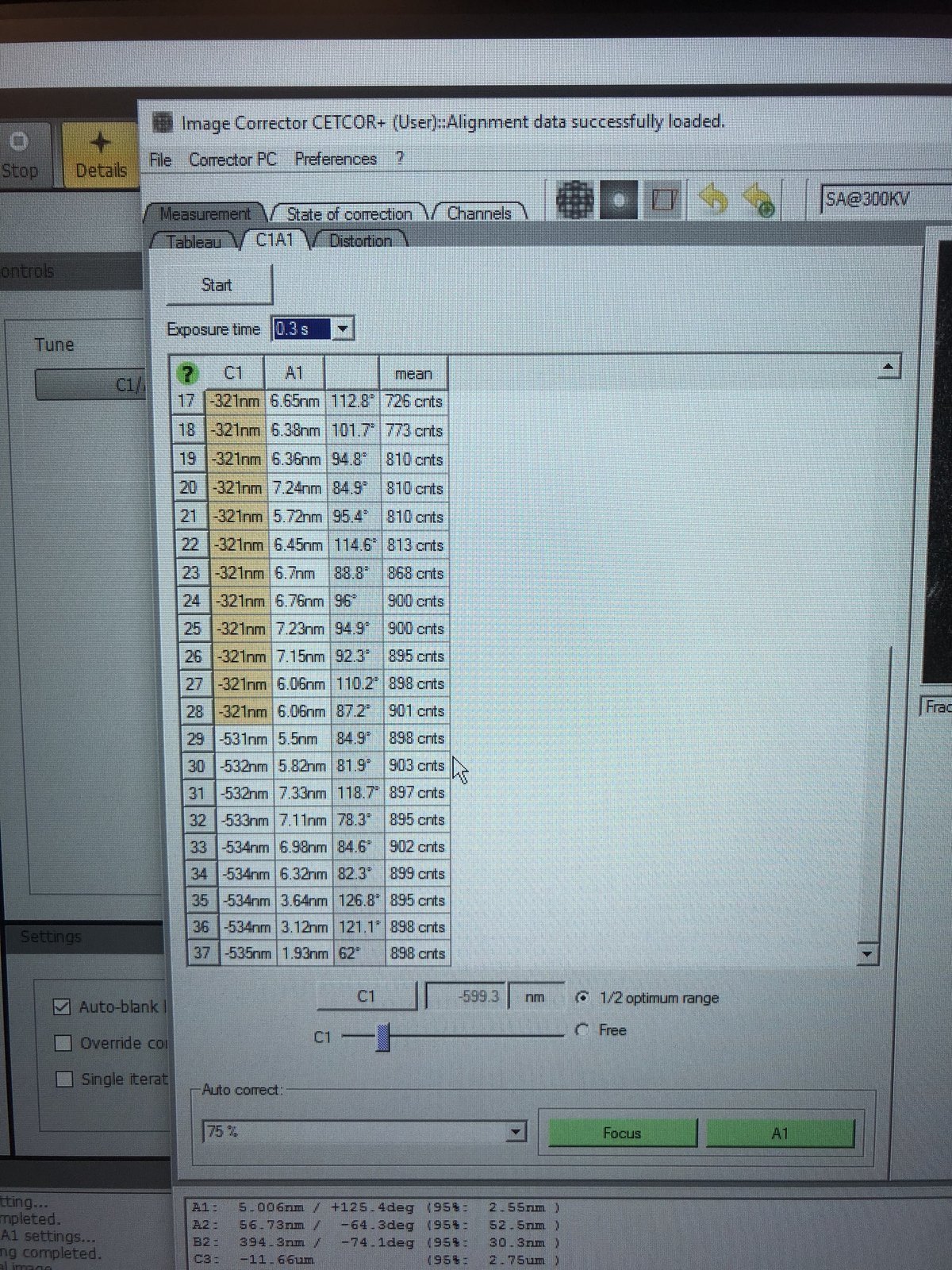
-
Under
Auto correct, set to75%, then pressFocusandA1during the iteration to apply corrections. -
Aim for
A1< 5 nm. IfC1shows orange, manually adjust the Z-axis during the iteration.C1should be close to the suggested value (in the image above, the software suggests C1 of −599.3 nm).
-
2.3 Tableau measurement
Tableau measures higher-order aberrations by acquiring images at multiple beam tilts. This is necessary for sub-angstrom resolution.
-
Run Tableau
-
In
ImageCorrector, go to theTableautab, selectStandardnext toTableau type, then clickStart.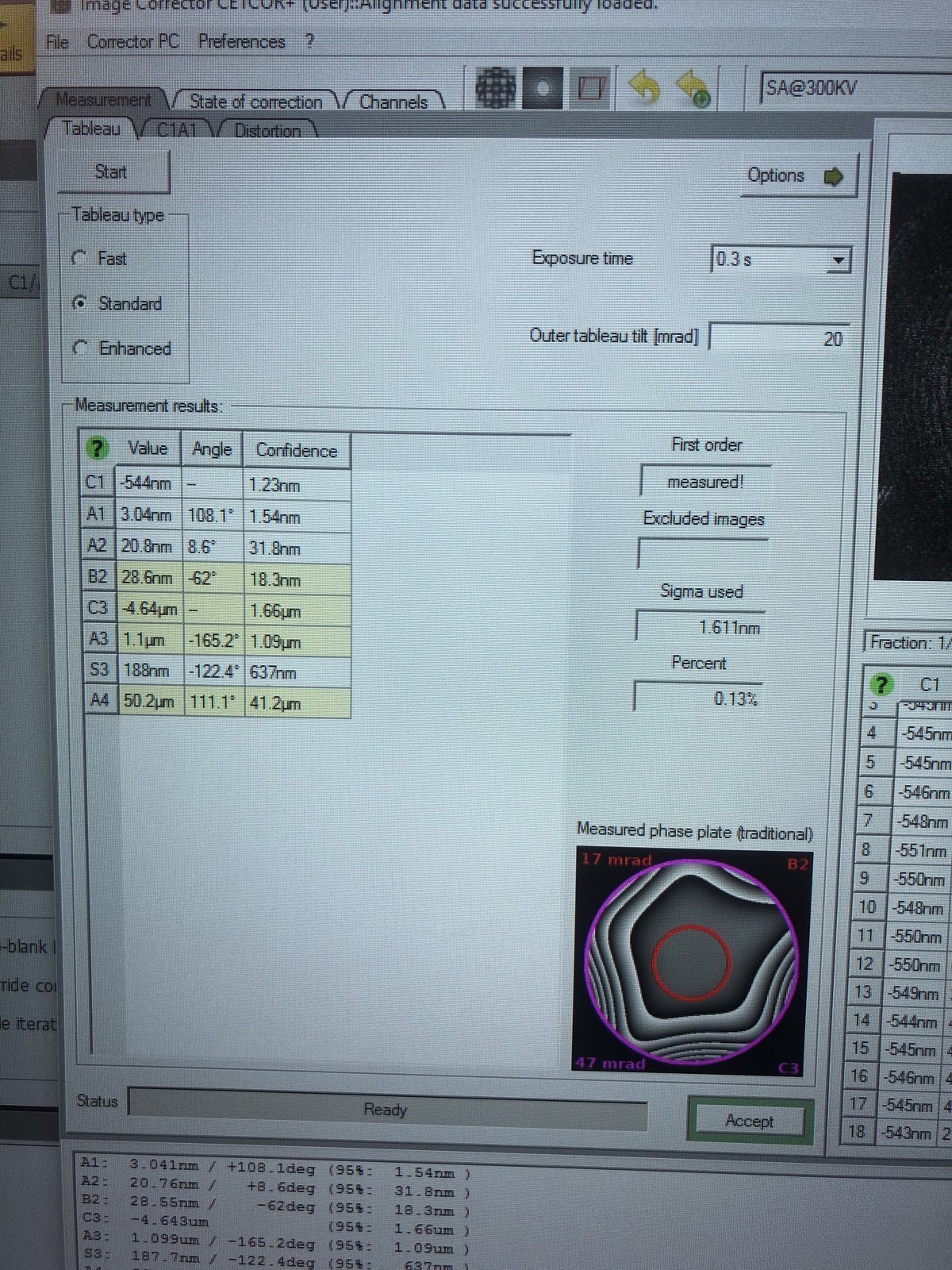
-
-
Verify results
-
After the iteration completes, verify the aberration values match the targets below, then click
Accept:Parameter Resolution < 0.10 nm (20 mrad) Resolution < 0.08 nm (24 mrad) A1 < 5 nm < 5 nm A2 < 100 nm < 50 nm B2 < 100 nm < 50 nm C3 ~ −8 μm ~ −8 μm A3 < 5 μm < 1.5 μm S3 < 5 μm < 1 μm -
In
Velox, click the camera button to capture an image and verify improvements.
-
2.4 Save optics settings
-
Save register
-
In
TEMUI, go toFiles, thenSBL FEG Registers. -
Add name
300KV-TEM-<NAME>and clickAdd.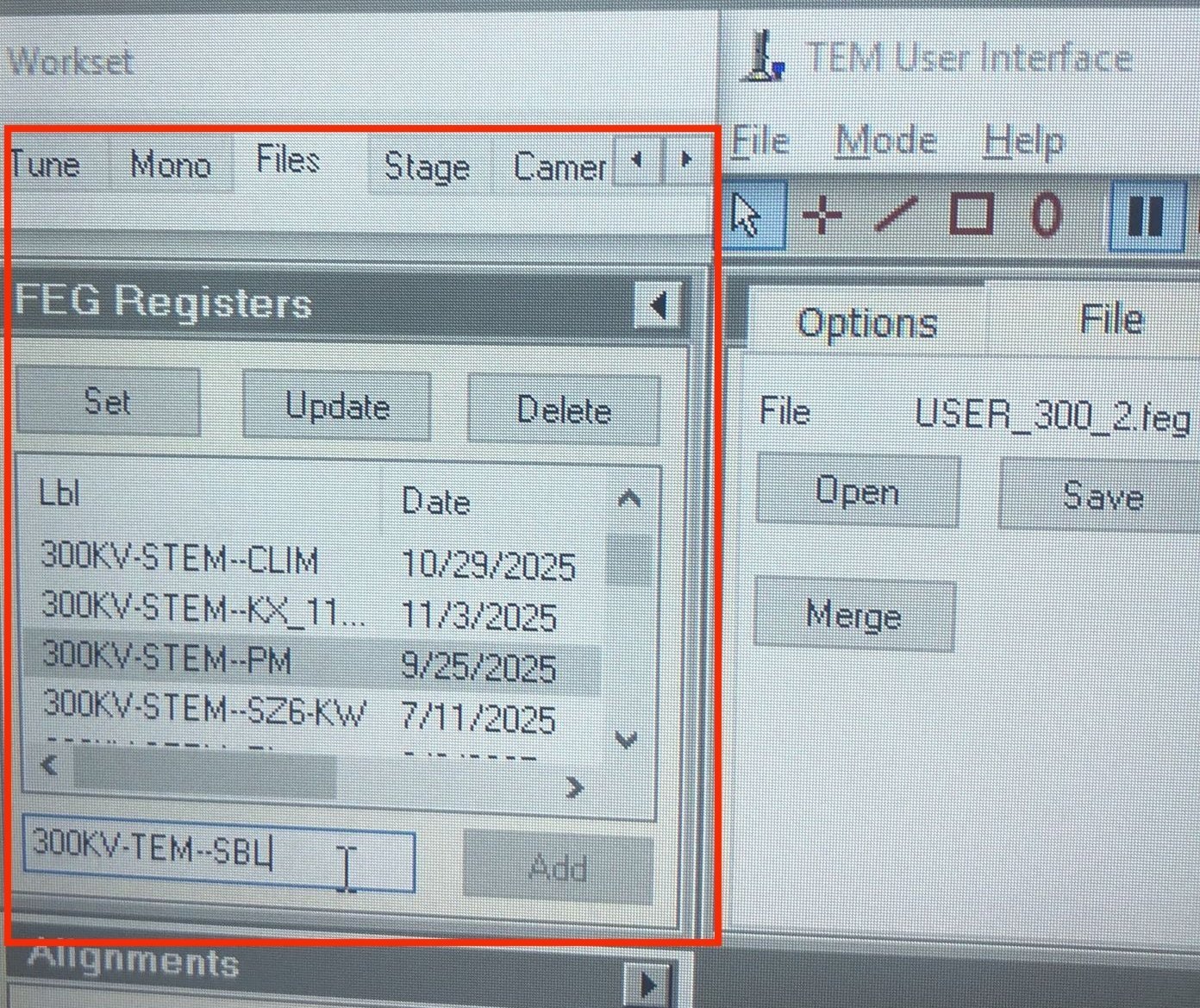
-
-
Verify corrected image
- In
Velox, click thePlaybutton to start live imaging and verify the aberration-corrected image quality. - Done. You are now ready for STEM probe alignment.
- In
Appendix
Save file to USB
Plug your USB into the following computer:

Reference images (click to expand)
Gray colors during C1A1 probe correction:
Seeing gray colors like below?
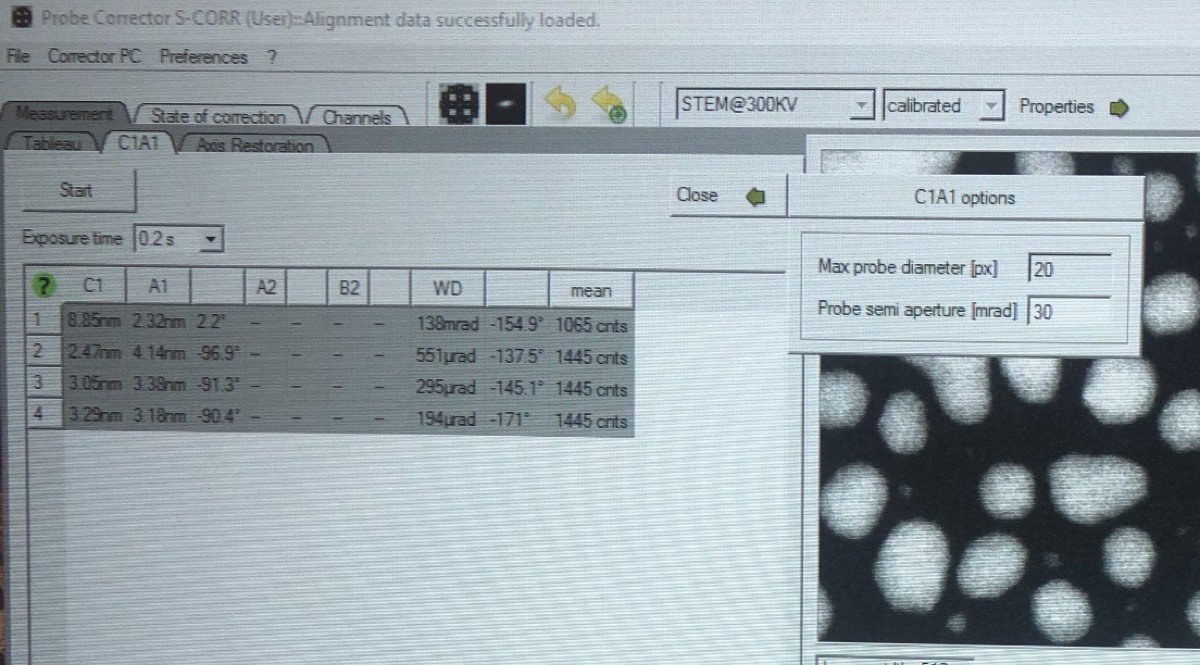
In Velox, click Auto-tune. Increase the signal until it touches the red and blue dotted lines:
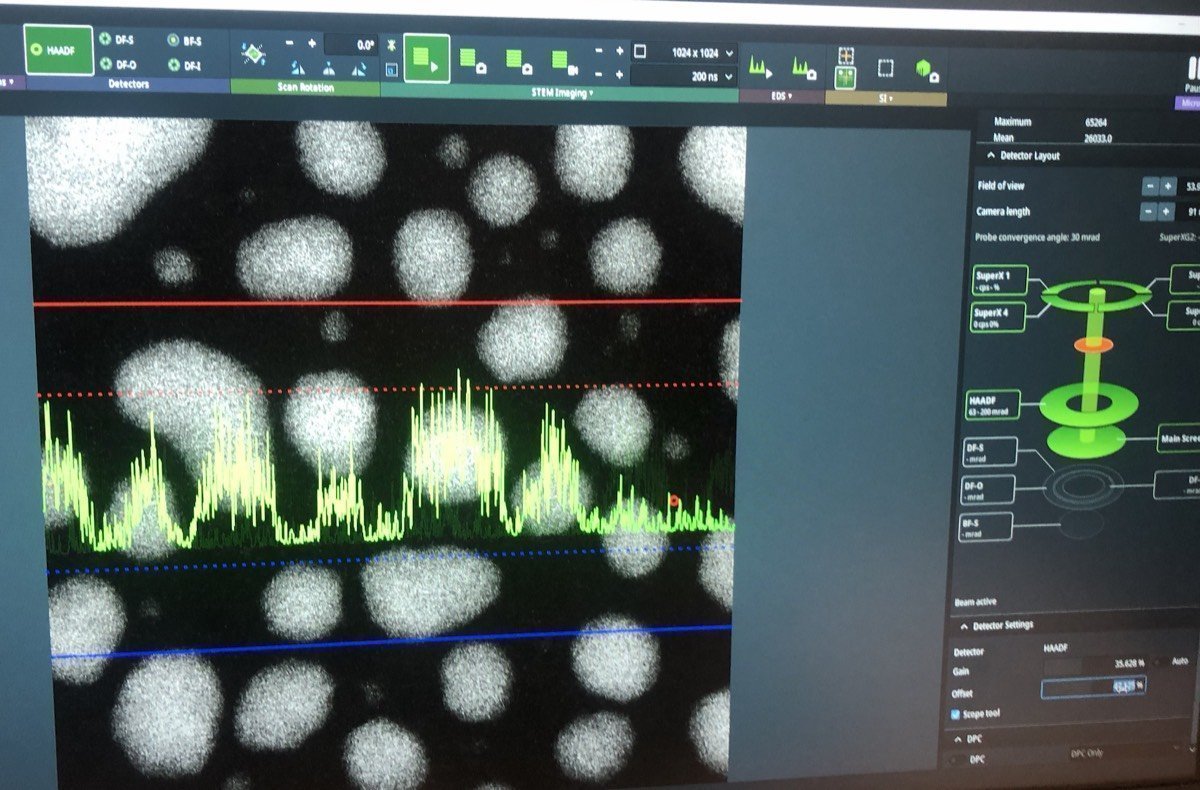
Hand panel R1, R2, R3 values:
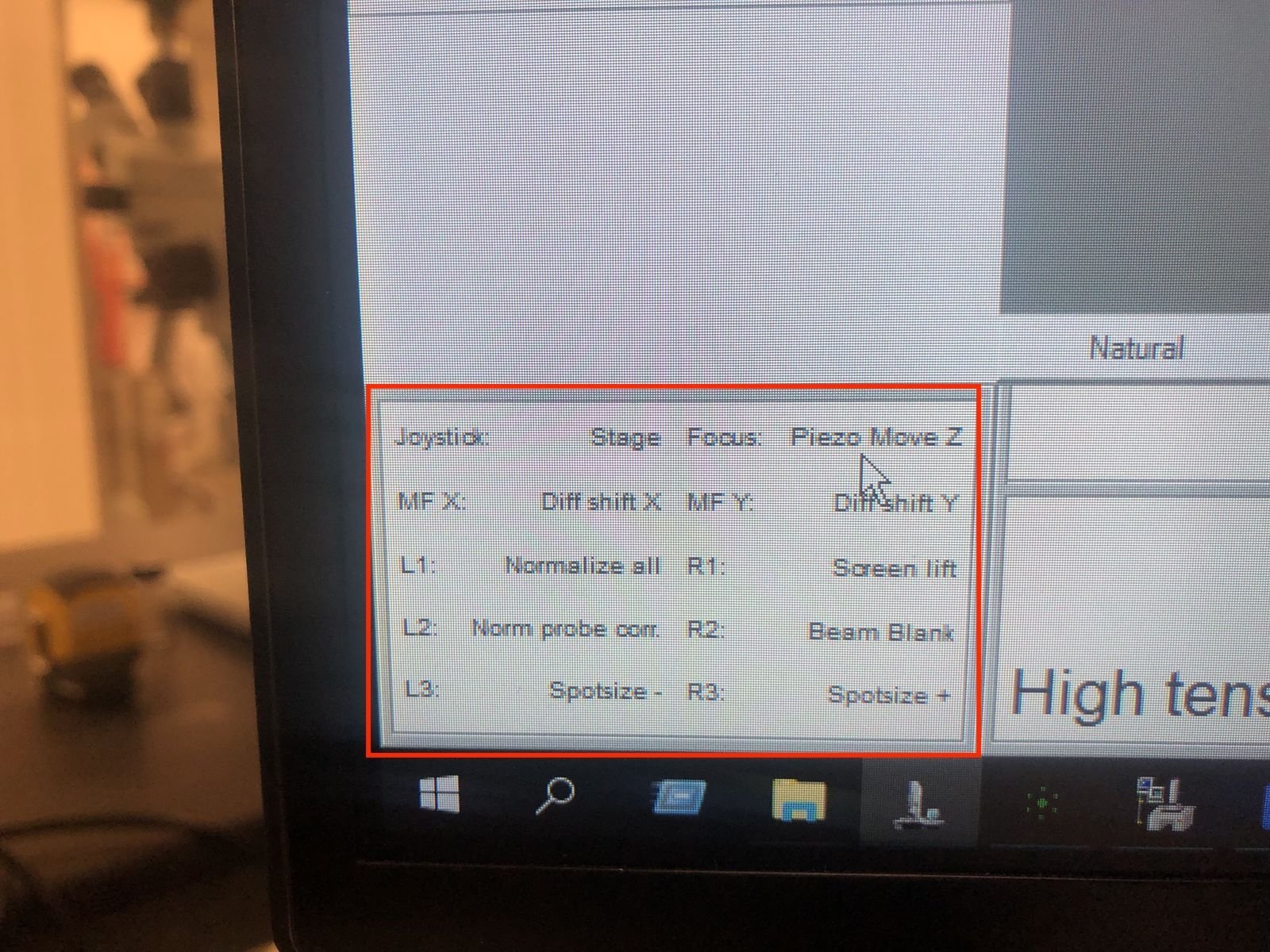
Stage position and coordinates:
Dose rate and TEM mode display:
HAADF detector on TEMUI:
Samples with holes:
Wobbler to check eucentric height:
At eucentric height, tilting the holder should induce minimal shift.
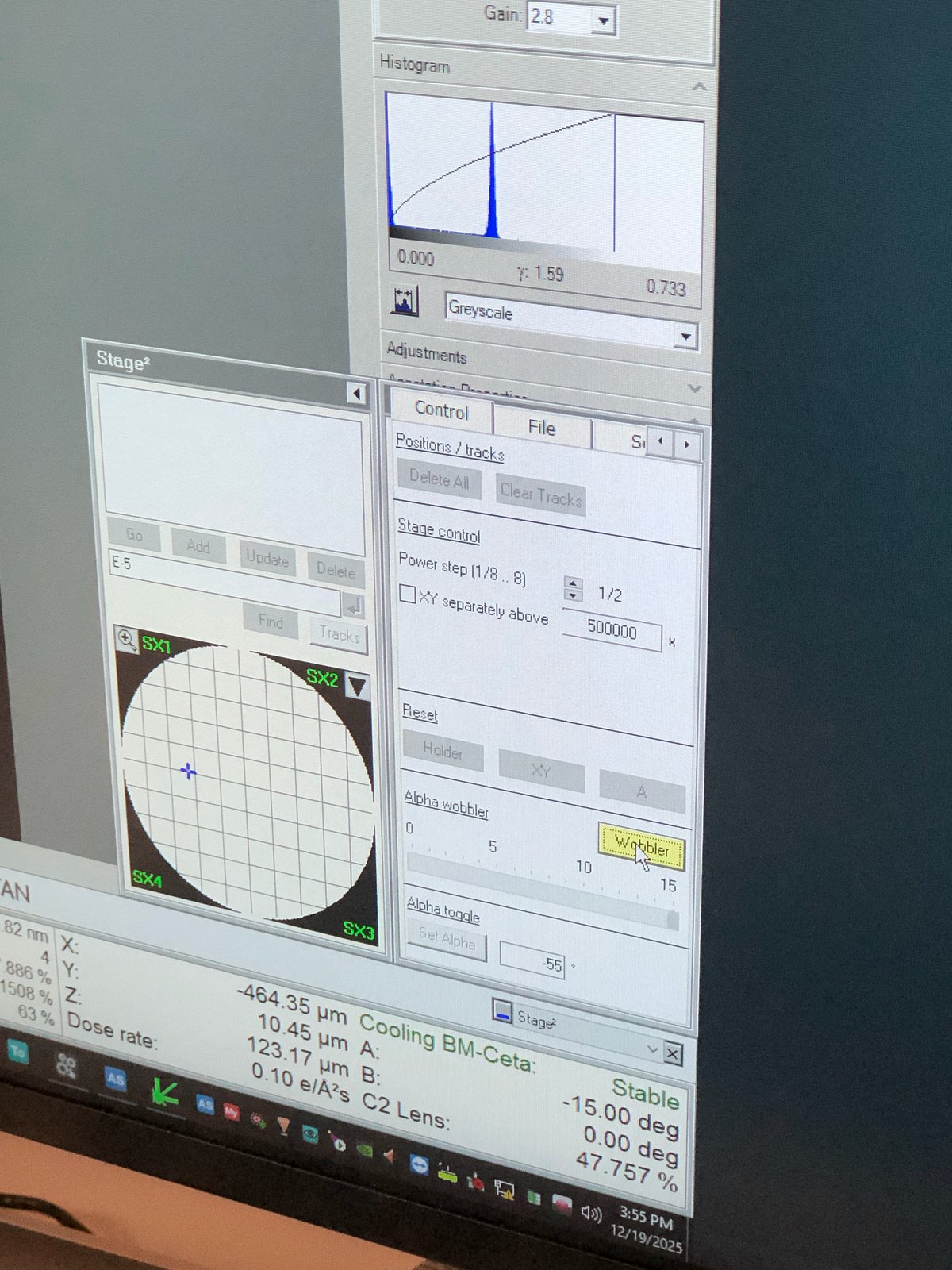
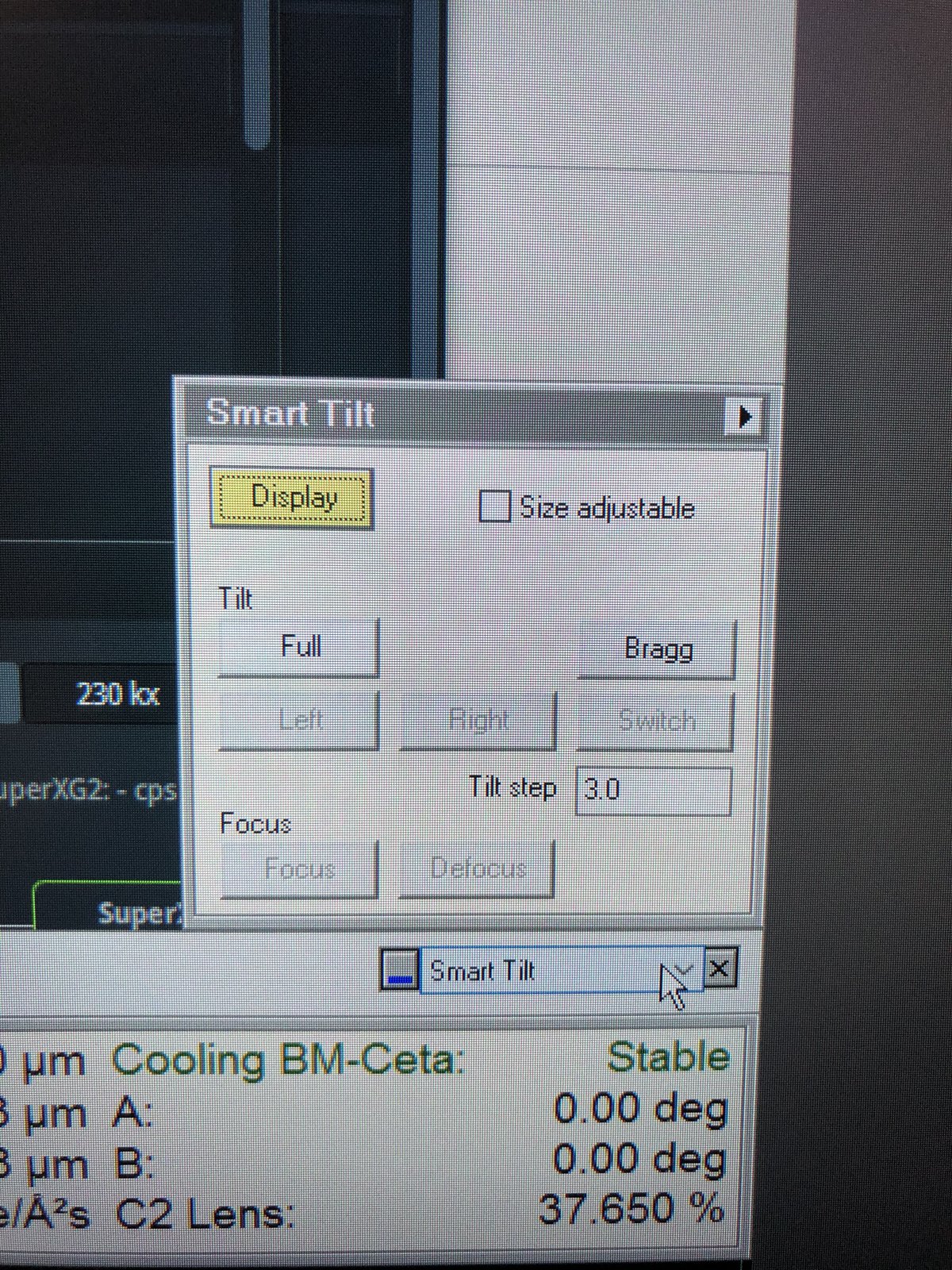
Smart tilt:
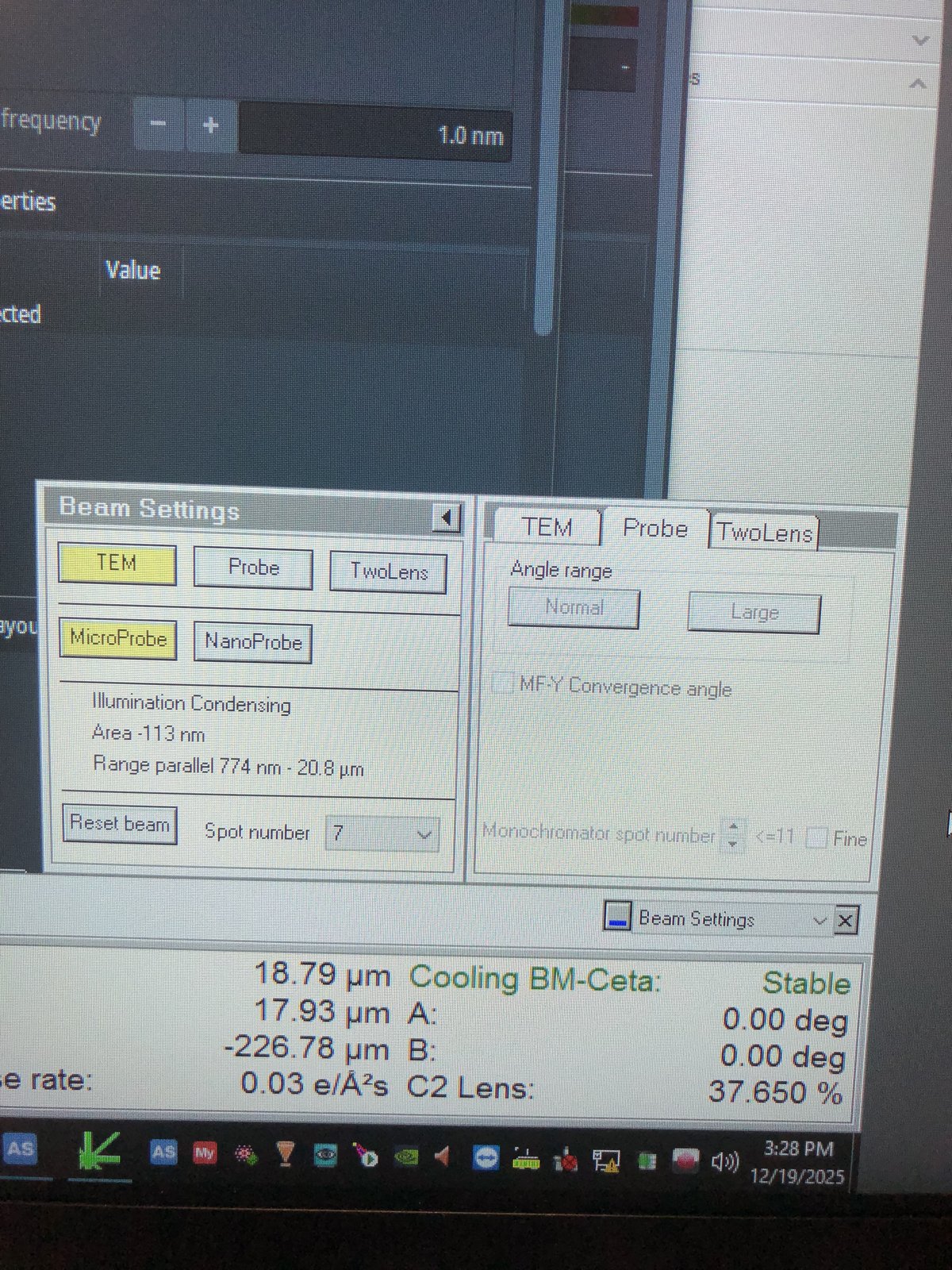
Beam setting in Quick tab:
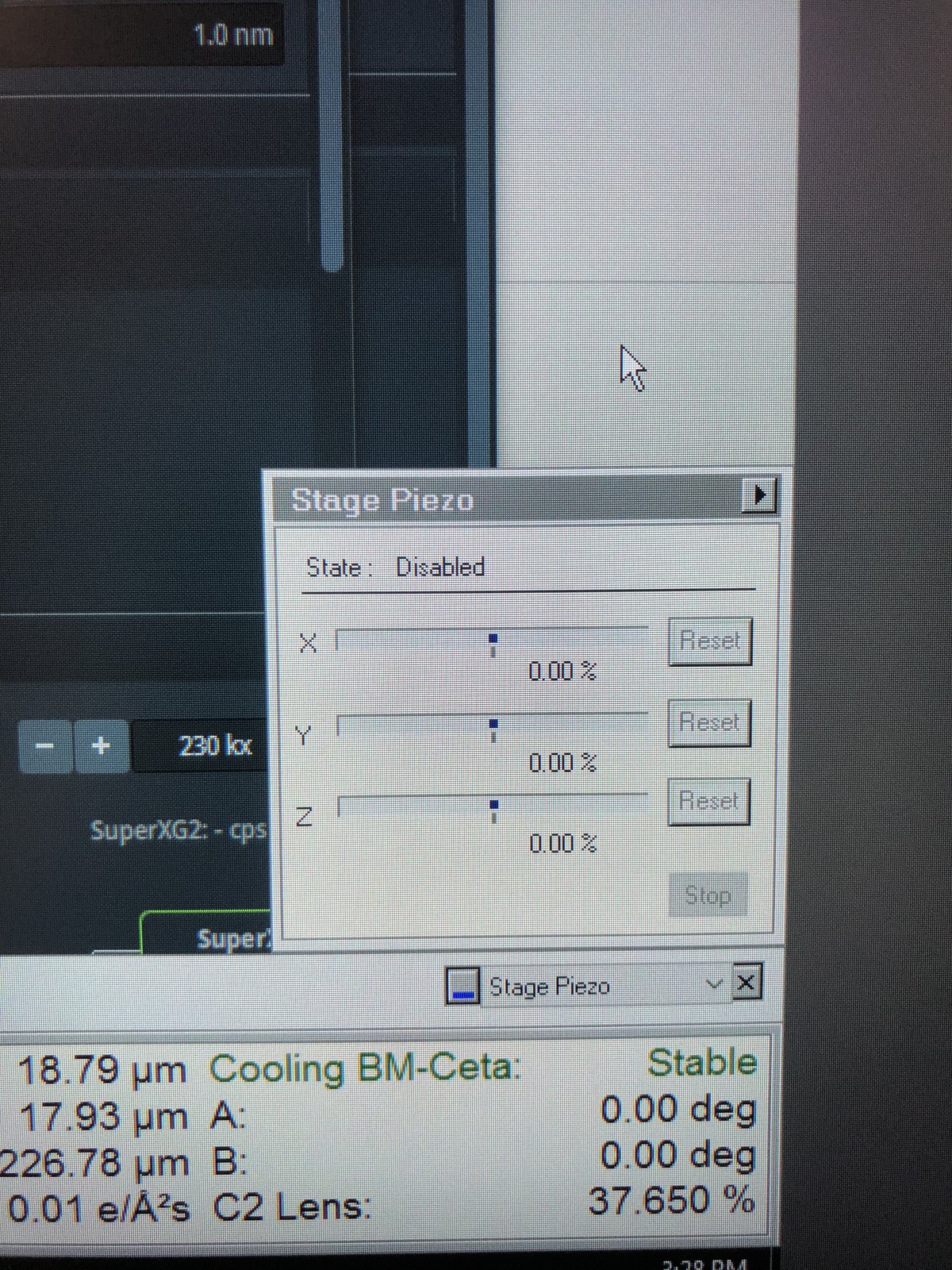
Stage piezo in Quick tab:
Stage tab:
Troubleshooting
Common problems encountered during TEM sessions.
| Problem | Cause | Solution |
|---|---|---|
| Beam is not round after C2 alignment | Condenser astigmatism | Go to Stigmator, then Condenser, adjust mulXY knobs |
| Beam shifts when changing magnification | Beam Shift not set | Use Direct Alignment, then Beam Shift to store center position |
| Image drifts when tilting | Eucentric height not set | Re-do eucentric height (1.3) |
| C1A1 shows orange for C1 | Focus too far from target | Manually adjust Z-axis during iteration |
| Tableau values outside specification | Higher-order aberrations uncorrected | Run additional Tableau iterations, reduce Auto correct to 75% |
| Gray image in Velox during C1A1 | Intensity too low for corrector | Adjust intensity knob to 800-900 counts during iteration |
| No beam visible after opening column valves | Beam is blanked or screen not inserted | Check beam blank status, verify screen position |
FAQ
Convergence angle: In TEMUI, go to Beam Setting, then Probe, and use the mulXY knobs to adjust.
Tableau and C1A1: Tableau measures aberrations visually across multiple tilt angles. C1A1 corrects first-order aberrations (defocus and astigmatism). Run C1A1 first, then Tableau for higher-order corrections.
Underfocus direction: Counterclockwise on hand panel, Z-axis down.
Eucentric height: The z-position where tilting does not shift the sample. At eucentric height, defocus = 0 and probe size is smallest relative to the sample.
Beam Shift vs hand panel ball: Beam Shift stores the center position internally, so the beam stays centered when changing magnification. The hand panel ball moves the beam but does not save the position.
Underfocus vs overfocus: Underfocus produces bright white Fresnel fringes at edges. Overfocus inverts the contrast with dark fringes.
Monochromator: Filters the electron beam to select a narrow energy range, improving energy resolution for EELS and reducing chromatic aberration.
Two-lens vs three-lens mode: Two-lens mode (C1+C2) turns off C3, providing a simpler beam path for C2 aperture alignment. Three-lens mode (C1+C2+C3) is the standard operating mode for TEM imaging.
Objective lens in TEM: In TEM, the objective lens sits below the sample and forms the first magnified image. In STEM, it sits above the sample and focuses the probe.
C2 aperture purpose: The C2 aperture blocks off-axis electrons, controlling the convergence angle and beam current. It must be centered on the optical axis for symmetric beam expansion.
References
Changelog
- Mar 1, 2026 - Restructure to match STEM guide format with subsections, checklists, and troubleshooting table
- Dec 15, 2025 - Add pre-probe corrector with STEM Direct Alignment steps by @bobleesj
- Dec 12, 2025 - Add STEM training images by Guoliang Hu
- Dec 8, 2025 - First draft of Spectra training by @bobleesj
4DSTEM
This guide covers 4DSTEM data acquisition using the Dectris Arina detector on the Spectra 300 at Stanford SNSF. 4DSTEM records a full convergent beam electron diffraction (CBED) pattern at every scan position, producing a 4-dimensional dataset (2D scan x 2D diffraction). Screenshots recorded by Guoliang Hu during training; instructions written by Sangjoon Bob Lee.
Prerequisite: Complete TEM (Spectra) column alignment and STEM (Spectra) probe correction before starting this guide.
Acronyms:
mulXY- Multifunction X/Y knobs on hand panelTEMUI- TEM User Interface (software)CBED- Convergent Beam Electron Diffraction
Overview
| Phase | Procedures | Time |
|---|---|---|
| Part 1: Detector setup | Retract CETA, initialize Arina, connect remote software | 5 min |
| Part 2: Beam configuration | Set convergence angle, apertures, camera length (optional) | 5 min |
| Part 3: Acquisition | Insert detector, acquire diffraction data | varies |
| Part 4: End session | Retract and power off the Arina detector | 2 min |
Part 1: Detector setup
1.1 Retract CETA detector
Before inserting the Arina detector, a user must retract the CETA camera. Both detectors occupy the same physical space below the column. If the CETA is not retracted, inserting the Arina will crash both detectors. Do not skip this step.
-
On the bottom left computer, open the blanker/shutter software (red square icon with white T).
-
Click the CETA icon to retract the CETA detector.
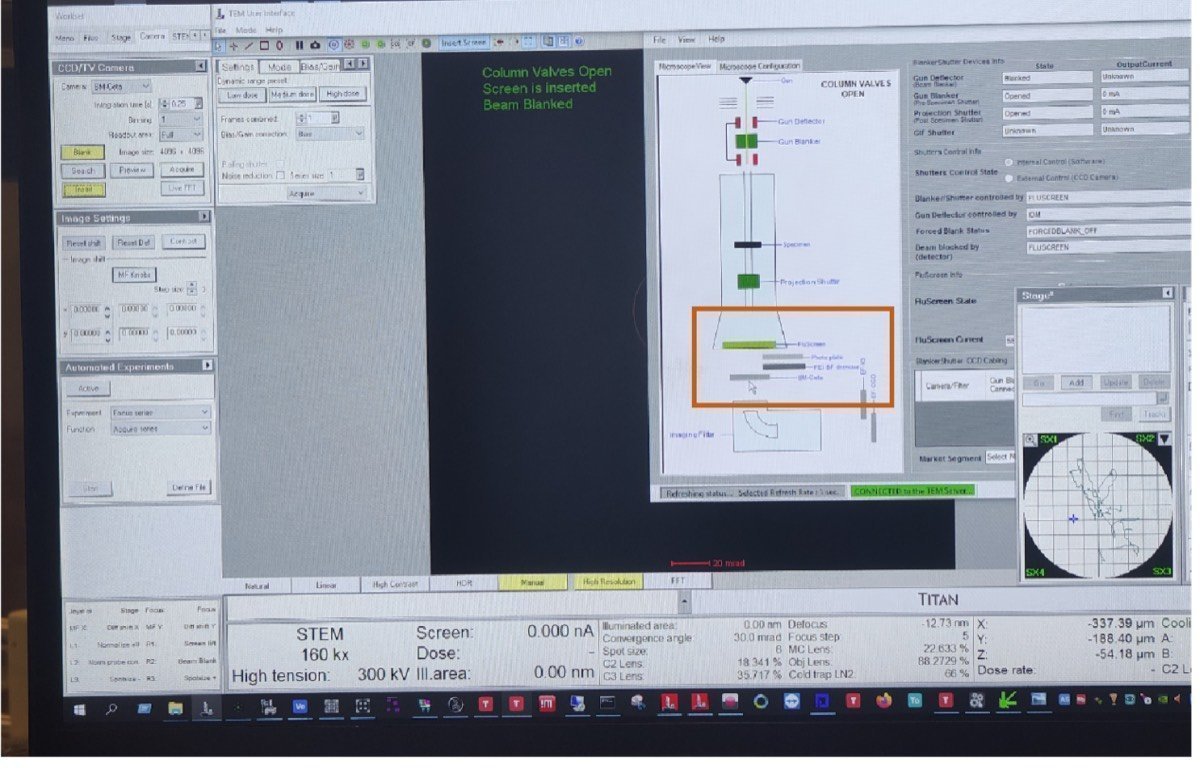
-
Visually verify the CETA camera position is retracted from the diagram.
-
In
TEMUI, locate theSTEM Detector (User)panel and verify all detectors are retracted:- HAADF: Retracted
- BF-S (Bright Field): Retracted
- DF-S (Dark Field): Retracted
1.2 Initialize the Arina detector
-
Open the instrument enclosure on the Spectra 300.

-
Locate the Dectris Arina detector unit inside the enclosure.

-
Press and hold the button below the Arina detector (blue indicator light) for 10 seconds. When powered on, the button stays pressed in and the blue light is illuminated.
1.3 Connect remote software
-
On the control workstation, open Firefox and click the remote connection bookmark.

-
Enter the detector IP address:
192.168.12.73. -
Click
Initialize detector. Wait for initialization to complete before proceeding. The interface shows a progress bar while the detector initializes.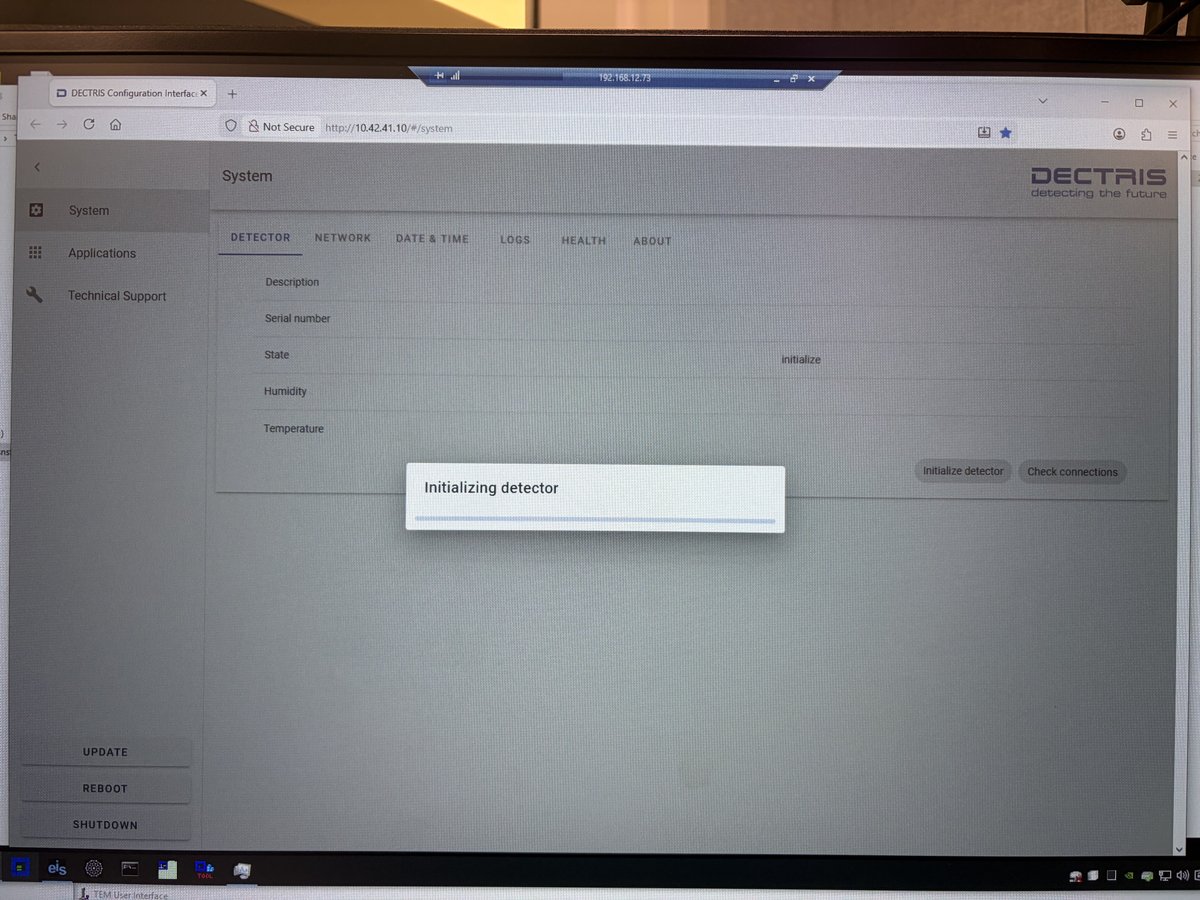
1.4 Configure file saving
-
Open the NOVENA detector software.
-
Click
Save Imagesand select the destination folder.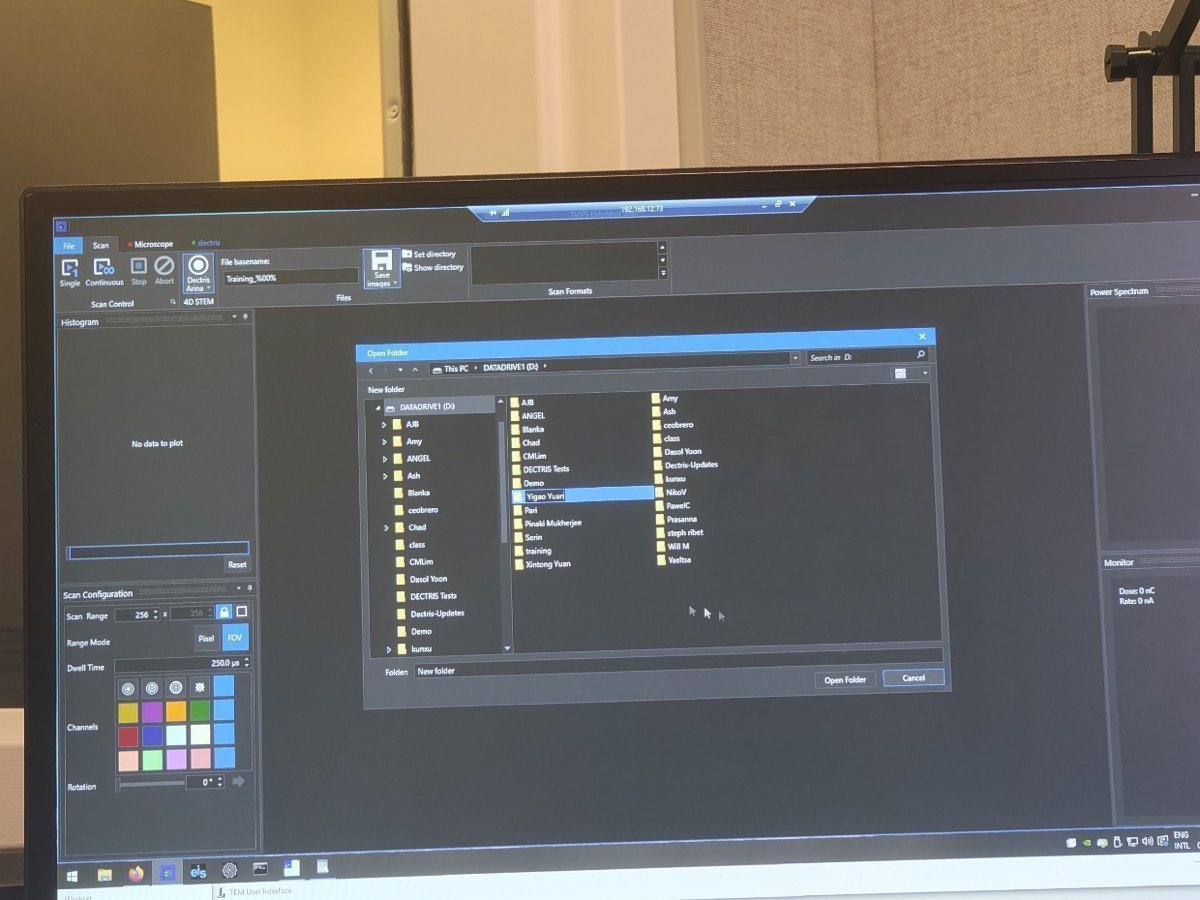
-
Set the filename format to
(name)_%00%. The%00%placeholder auto-increments the frame number. -
Use
Continuousfor live streaming (preview) andSingleto record and save a dataset.
Part 2: Beam configuration (optional)
The STEM (Spectra) guide sets 30 mrad convergence angle by default. If that is suitable for your experiment (e.g., ptychography), skip this section and go directly to Part 3: Acquisition. If you need a different convergence angle (e.g., nanobeam diffraction), follow the steps below.
Why change the convergence angle for 4DSTEM?
The convergence angle depends on the type of 4DSTEM experiment. For ptychography, 30 mrad (the same as standard STEM) works well because overlapping disks are part of the reconstruction. For nanobeam diffraction, where separated Bragg disks are needed to index reflections, a much smaller angle is used, typically 1 to 10 mrad depending on the material and the required disk separation.
Interactive demo: Explore how convergence angle affects the CBED pattern at bobleesj.github.io/electron-microscopy-website/cbed
2.1 Enable descan
- In
TEMUI, locate theSTEM Imaging (Expert)panel and enableDescan.
2.2 Configure beam for nanobeam diffraction
The default STEM setup uses C2 = 70 and 30 mrad convergence angle. For nanobeam diffraction, reduce both to get separated Bragg disks. The table below shows typical values:
| Parameter | STEM default | Nanobeam diffraction |
|---|---|---|
| C2 aperture | 70 | 50 |
| C3 aperture | 1000 | 30 |
| Convergence angle | 30 mrad | ~10 mrad |
| Beam current | ~0.150 nA | ~0.032 nA |
| Camera length | 91 mm | 230 mm |
Why change the C2 aperture?
The C2 aperture limits the angular range of electrons entering the probe-forming optics. A smaller aperture (50 vs 70) blocks more off-axis electrons, producing a more coherent beam with cleaner diffraction patterns at each probe position. The C2 aperture size and convergence angle are proportional (approximately 7:1 ratio, e.g. C2 = 70 gives ~10 mrad).
TODO: Confirm the C2 aperture to convergence angle ratio with staff
-
In
TEMUI, go toTunetab, thenApertures. Change C2 from 70 to 50, and C3 from 1000 to 30.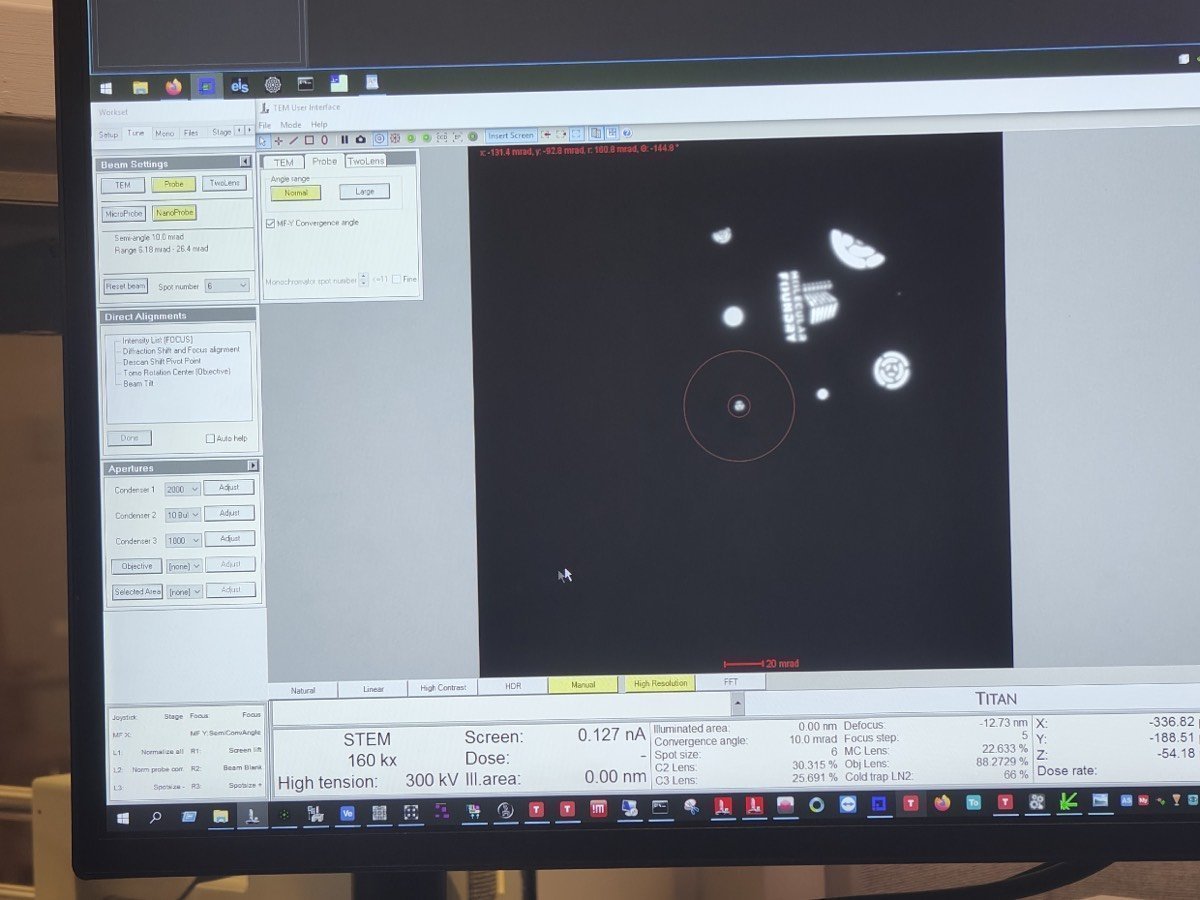
-
In
Beam Setting, clickMF-Y Convergence Angle. Use themulYknob to adjust the convergence angle to 10 mrad, then clickMF-Y Convergence Angleagain to deselect.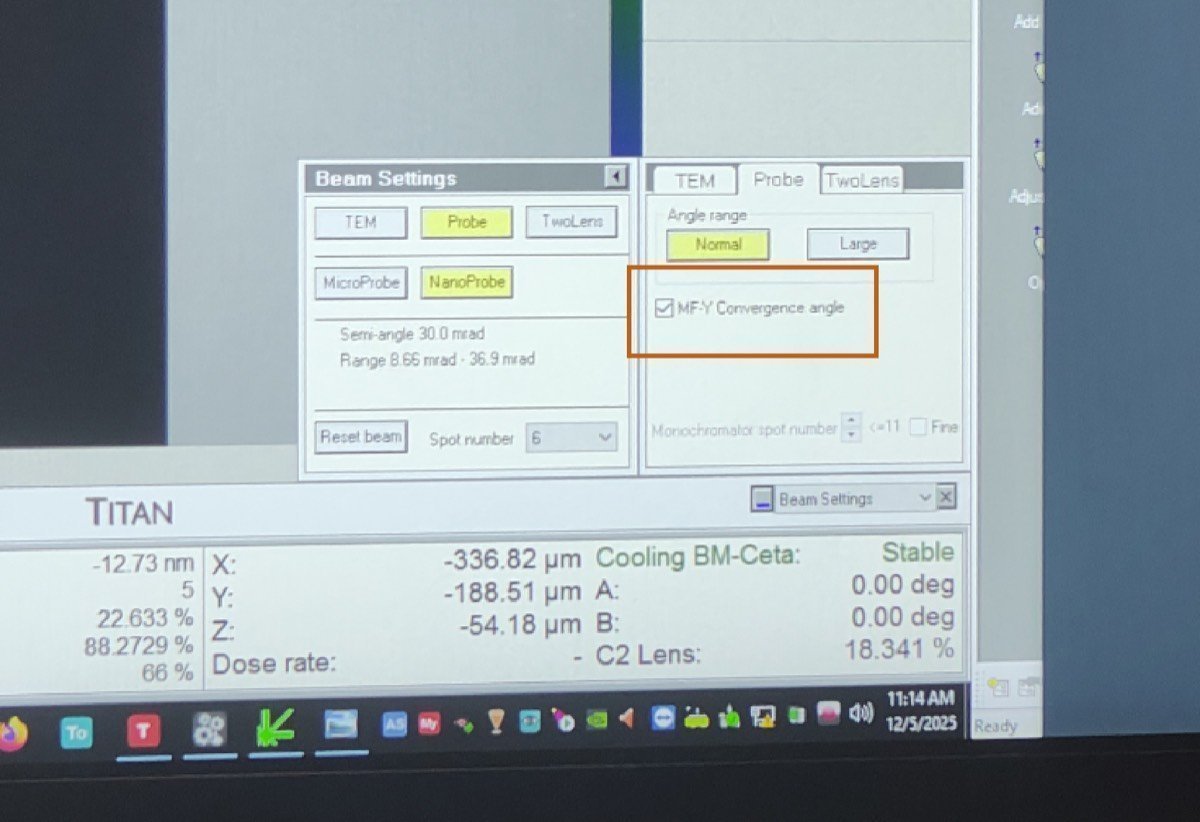
-
Adjust beam current: in
TEMUI, go toMono, clickFocus, and use the intensity knob to set the current to ~0.032 nA. -
Set the camera length to 230 mm (or 285 mm depending on the required angular range for your material).
2.5 Retract HAADF
- In
TEMUI, retract the HAADF detector. The HAADF ring would block electrons from reaching the Arina detector below.
Part 3: Acquisition
3.1 Insert detector and configure scan
-
On the Arina hand panel, press
Insertto move the detector into position. The green “Inserted” light confirms the detector is in place. -
On the scan control box, press
EDS Scan.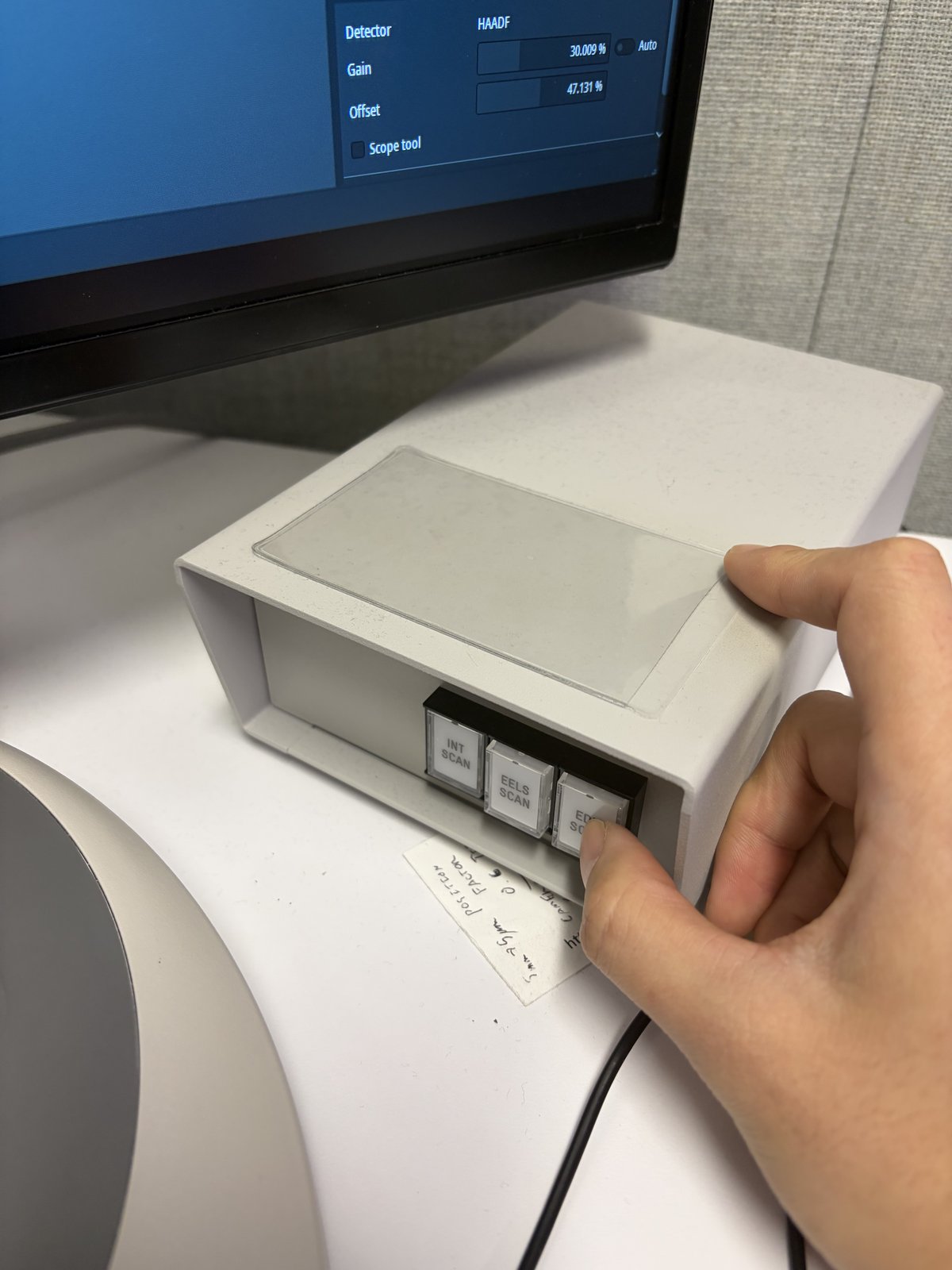
-
Press
R1on the hand panel to lift the fluorescent screen. The Arina detector sits below the screen.
3.2 Acquire data
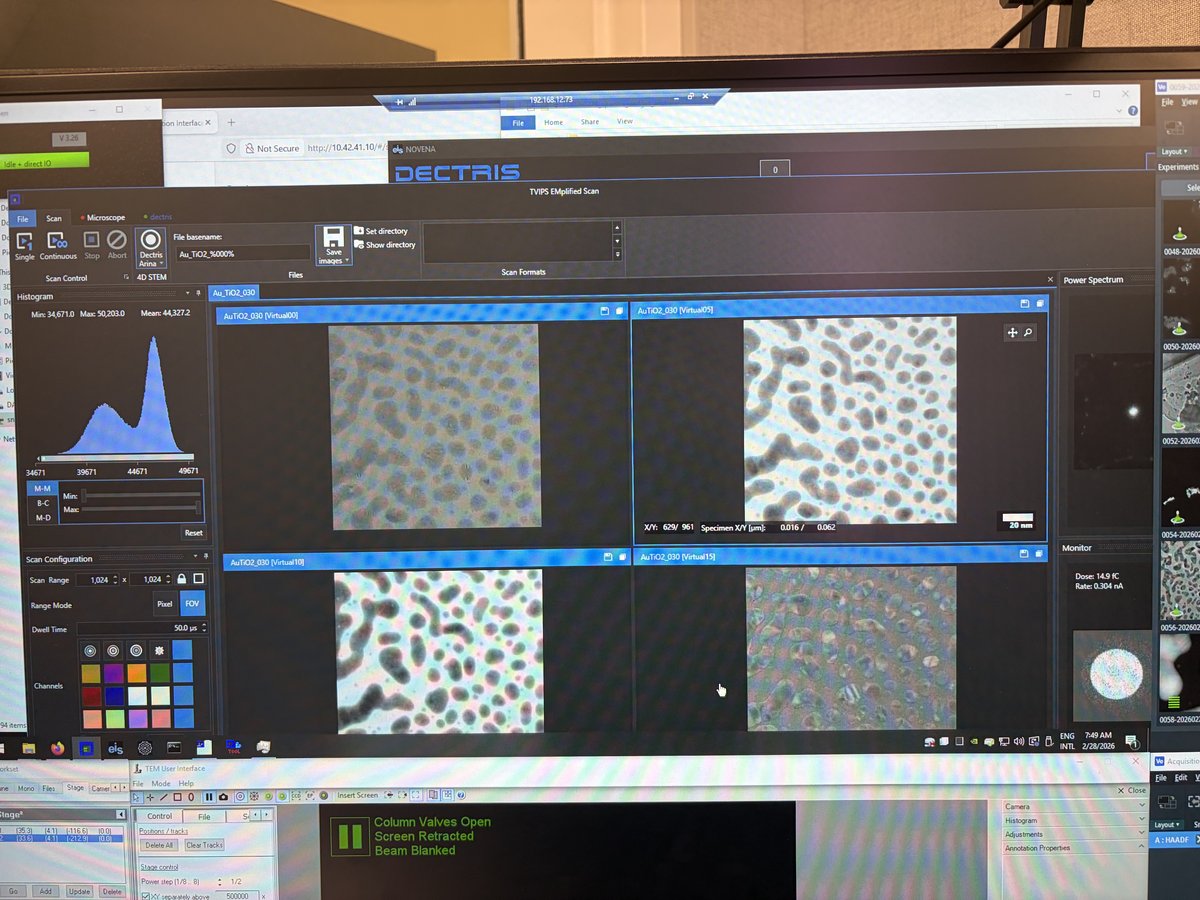
-
In the NOVENA software, click
Scan, thenContinuousto start a live preview. Verify the central beam is centered on the detector. -
If the beam is off-center, use the
mulXYknobs with diffraction shift to center it. -
Once centered, click
Stop, then clickSingle Scanto acquire and save the dataset.NOTE: Each scan produces a 4D dataset: a CBED pattern at every pixel in the scan area. File sizes can be large depending on scan resolution and detector binning.
3.3 Quick analysis
- In the NOVENA software, use
RebinandReprocessfor a quick check of the acquired data. For detailed analysis, export the data for processing with external software (py4DSTEM, etc.).
Part 4: End session
4.1 Retract the Arina detector
-
On the Arina hand panel, press
Retractto move the detector out of the beam path. The green “Retracted” light confirms the detector is clear.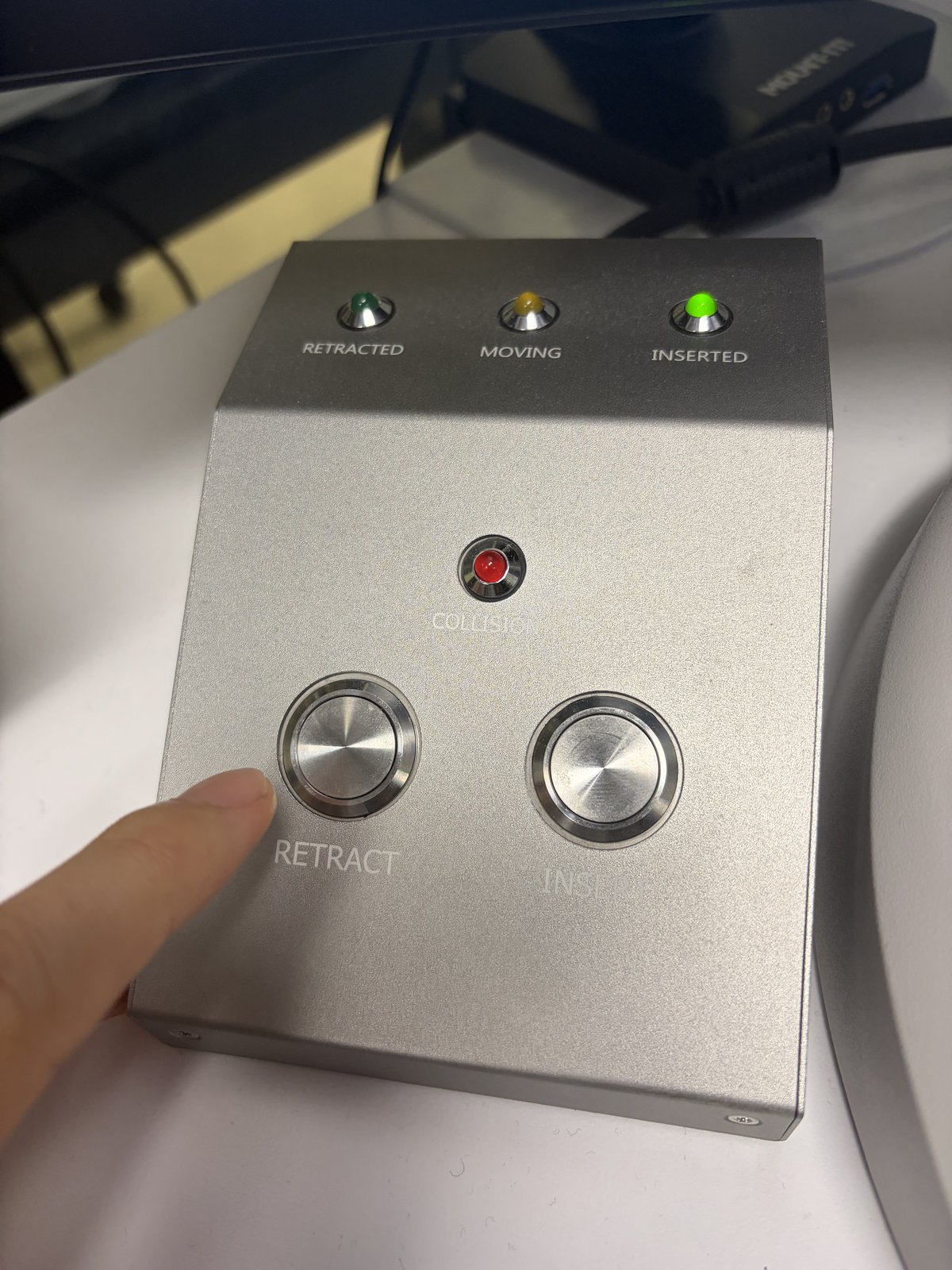
4.2 Power off the detector
- Open the Spectra 300 instrument enclosure.
- Press and hold the button below the Arina detector for 10 seconds. The button releases and the blue indicator light turns off.
4.3 Close session
Follow the steps in End session.
Changelog
- Feb 28, 2026 - Rewrite SOP by Sangjoon Bob Lee with full procedural instructions, inline FAQ dropdowns, and new images
- Dec 10, 2025 - First draft and images shared by Guoliang Hu
EELS
Caution
VERY ROUGH DRAFT - @bobleesj and Guoliang Hu took notes and pictures during training. This document will be updated with more detailed steps and images.
TODO:
- Add image for “EELS Scan” button (Step 2, Part 3)
- Better photo for STEM SI button
- Show what auto gain looks like
- Clarify zero loss extraction steps
- Add comparison images for beam not centered
This guide covers Electron Energy Loss Spectroscopy (EELS) on the Spectra 300. The process has two parts: calibration in TEM mode and STEM EELS spectrum imaging.
Prerequisite: Complete the STEM alignment procedure before starting.
Acronyms:
GIF- Gatan Imaging FilterEFTEM- Energy Filtered Transmission Electron MicroscopyZLP- Zero Loss PeakSI- Spectrum ImagingmulXY- Multifunction X/Y knobs on hand panel
Part 1: Calibration
Use a vacuum or thin amorphous carbon area for calibration.
-
Find a calibration area
- Locate a vacuum region or thin amorphous carbon area on the standard sample
-
Open DigitalMicrograph
-
Open
DigitalMicrographon the left monitor -
If you see any dialog box, click
OKto dismiss
-
Click
EFTEM(Energy Filtered Transmission Electron Microscopy)
-
-
Open FilterControl
-
Go to
Help→User Mode→Power User -
Go to
Window→Floating Window→Filter Control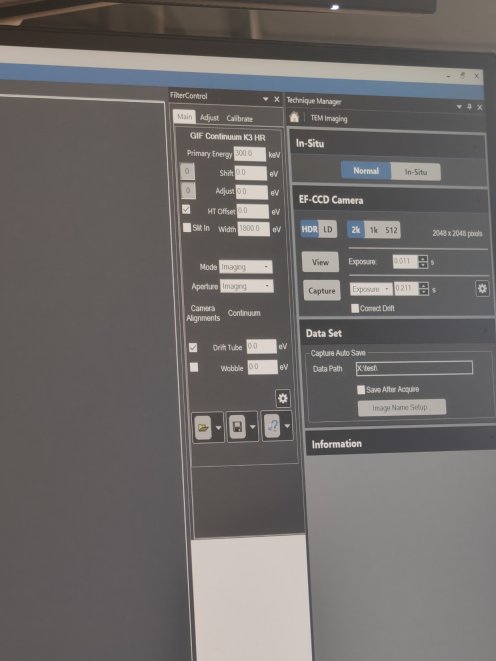
-
Notice the green circle in
TEMUIshowing EELS detector is active
-
-
Set beam intensity
-
Converge the beam by adjusting the intensity knob
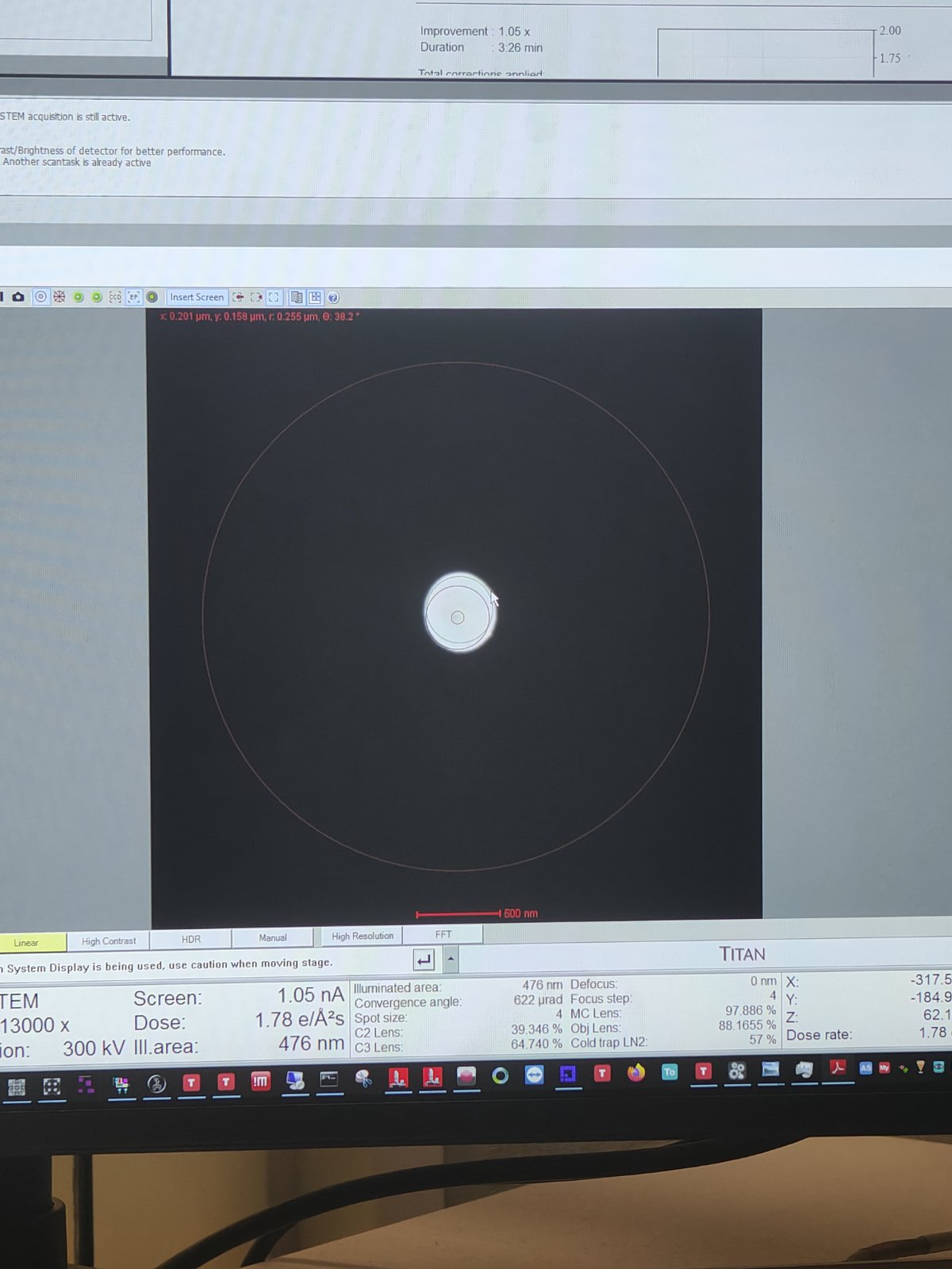
-
Lift the fluorescent screen by pressing
R1 -
Click
Viewin DigitalMicrograph -
Go to
Filter Control→Aperture→Maskto verify beam positionCorrect intensity:
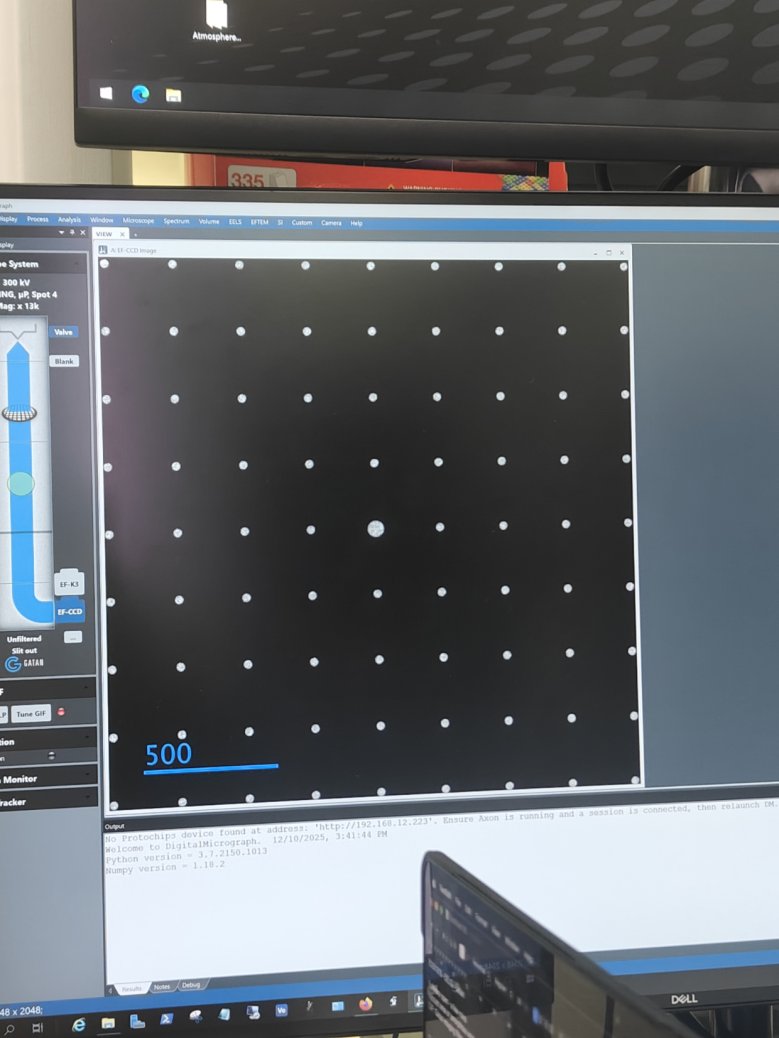
Too high intensity (oversaturated):
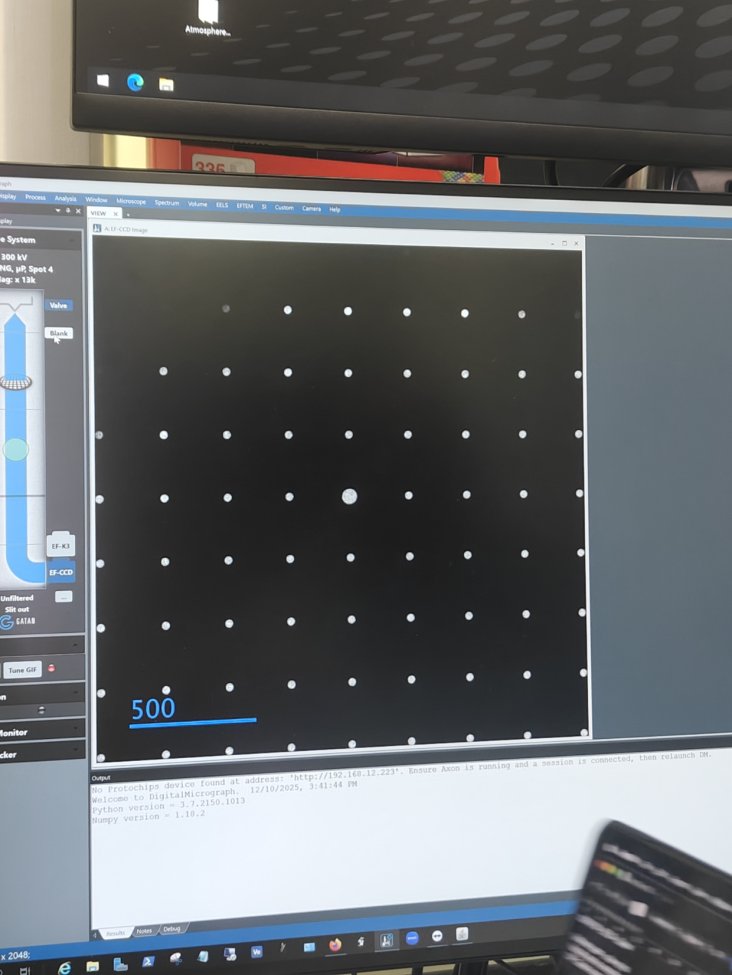
-
-
Center and tune the GIF
-
Click
Center ZLPin Filter Control -
Click
Tune GIF. Notice the message appears:
-
Click
OKto confirm
-
Part 2: TEM EELS acquisition
-
View the sample
-
Set magnification to ~17,000x
-
Click
Viewin DigitalMicrograph -
Select
EF-CCD Camera→Viewto see image real-time
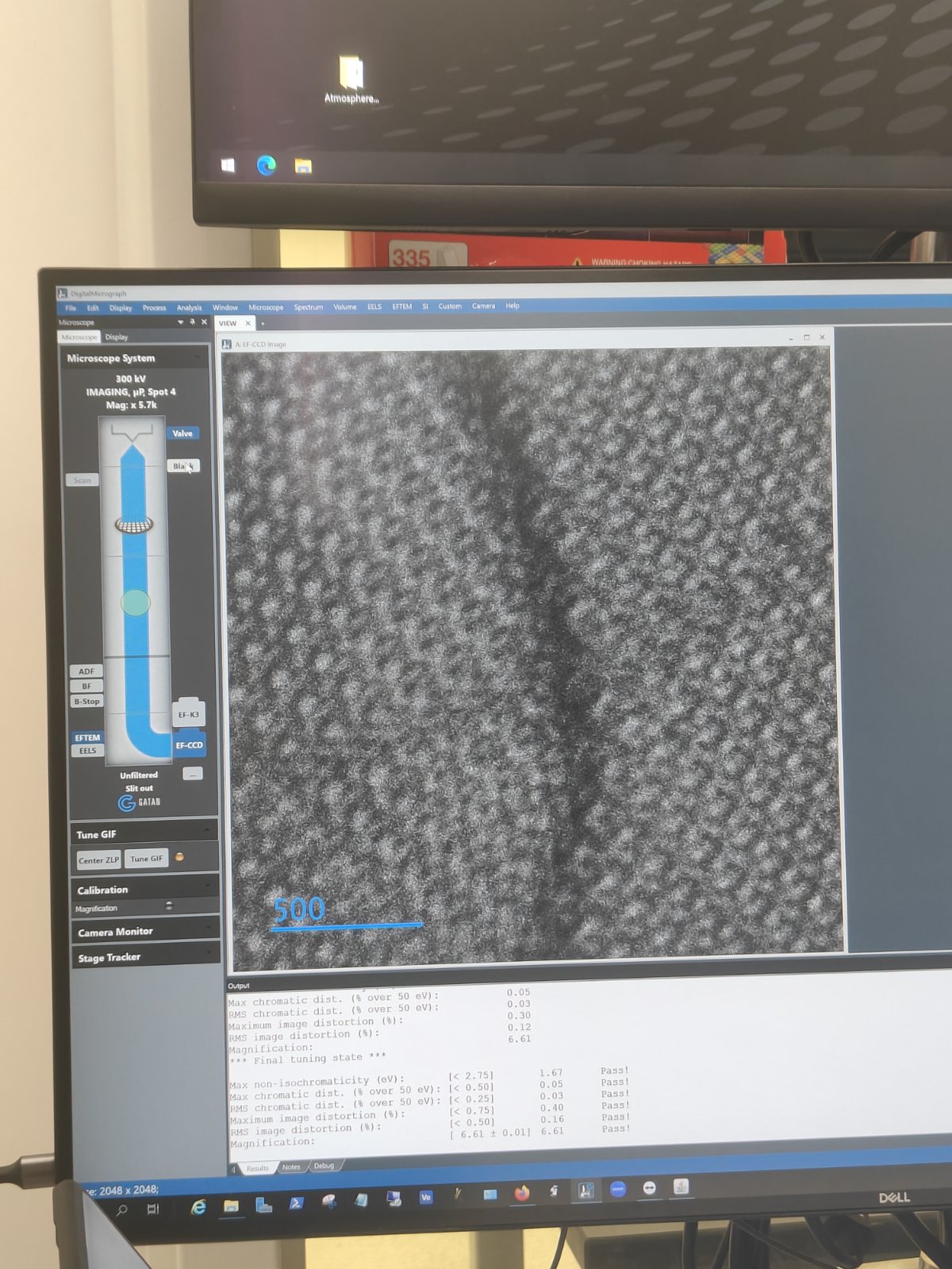
-
-
Acquire zero loss image
-
Go to
SingleMap→ clickZero Loss Image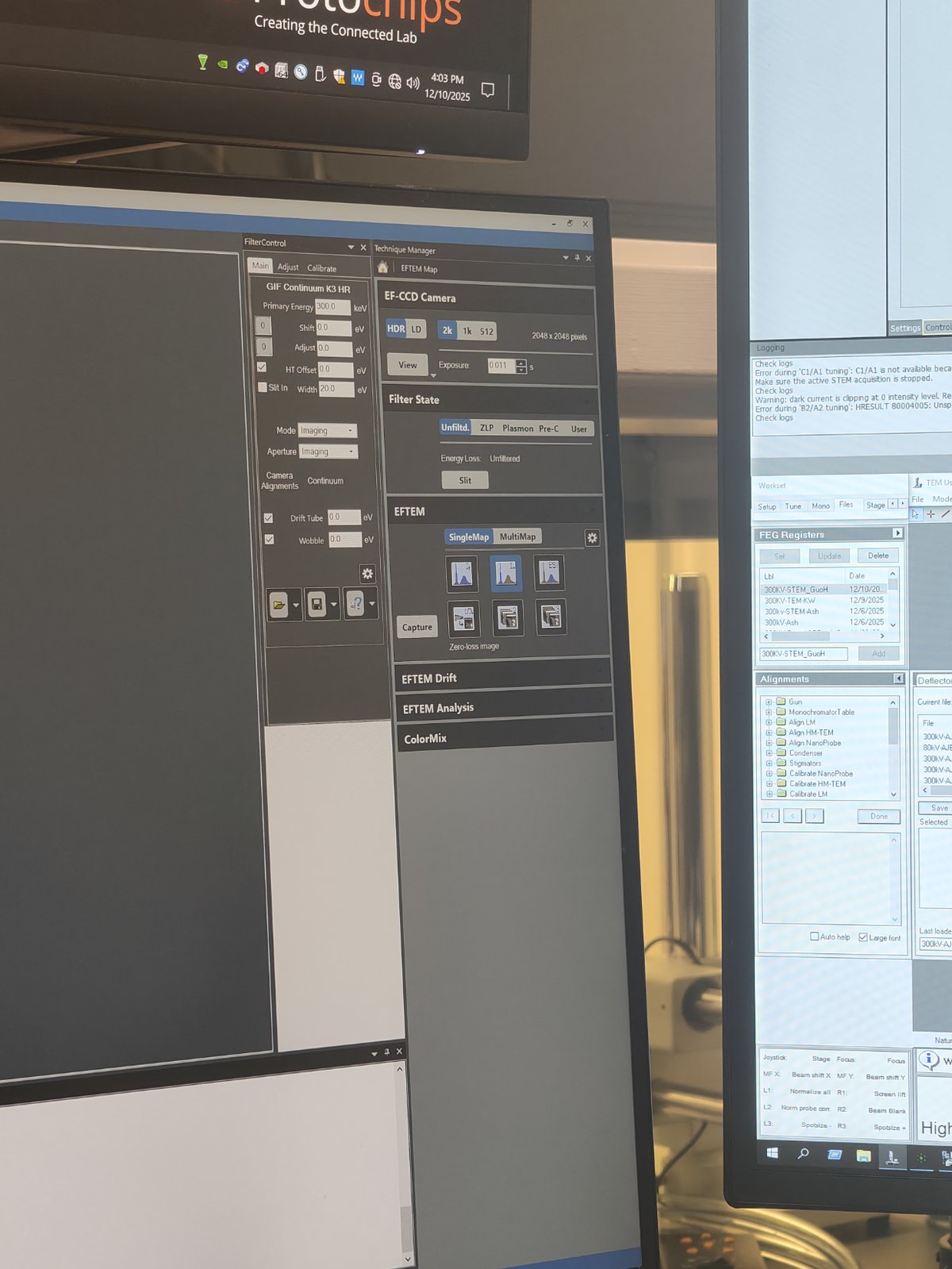
-
-
Switch to EELS mode
-
Click
EELSbutton to switch modes
FIXME: use image where EELS is clicked
-
Notice the 2D EELS spectrum. Observe the plasma peak near the zero loss peak.
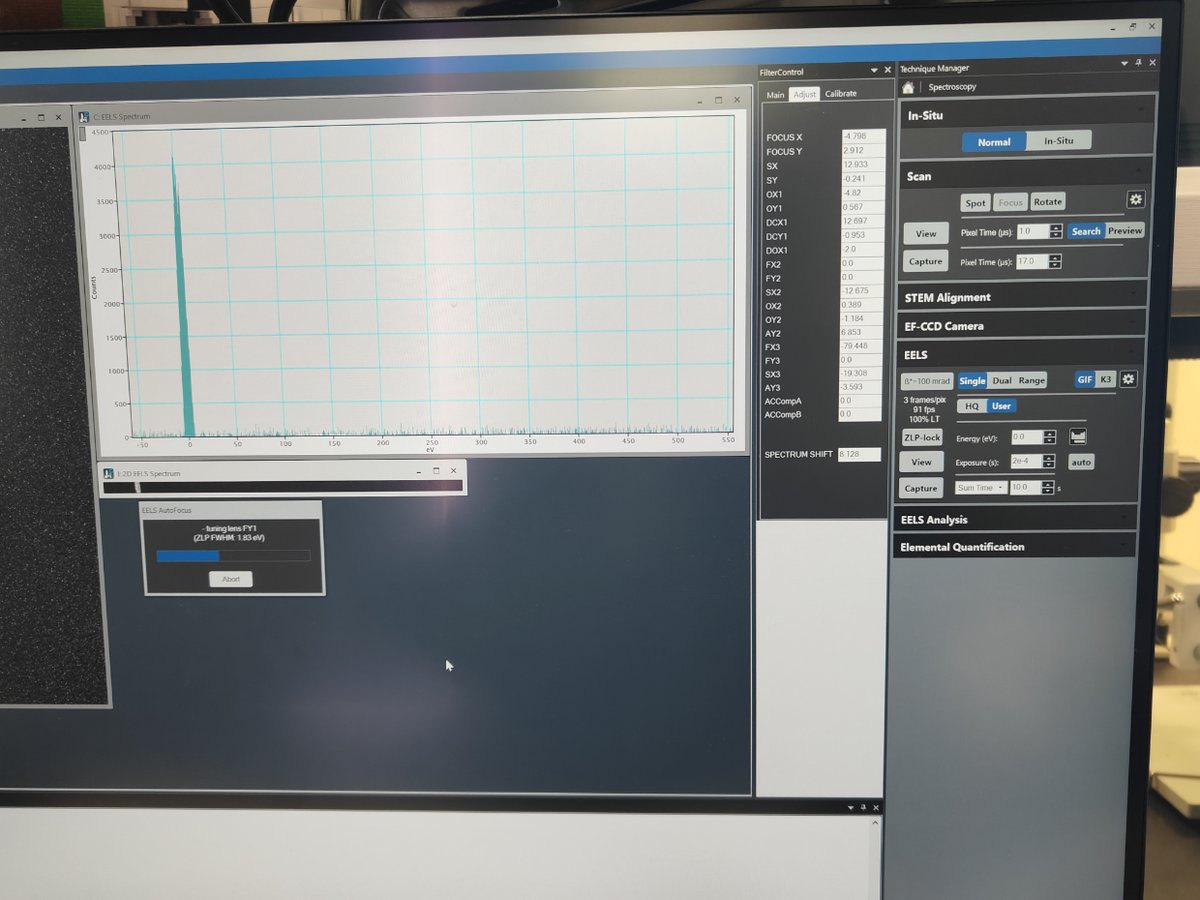
FIXME: what’s plasma peak?
-
-
Align the zero loss peak
-
Set exposure to 2e-4 in View mode
-
Click
Align ZLP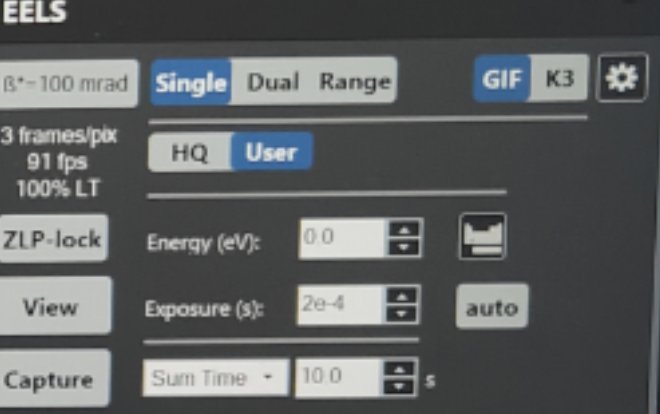
FIXME: where is ZLP click?
-
Part 3: STEM EELS spectrum imaging
-
Set camera length
-
In
Velox, change camera length to 29 mm or 37 mm -
Notice the beam size decreases
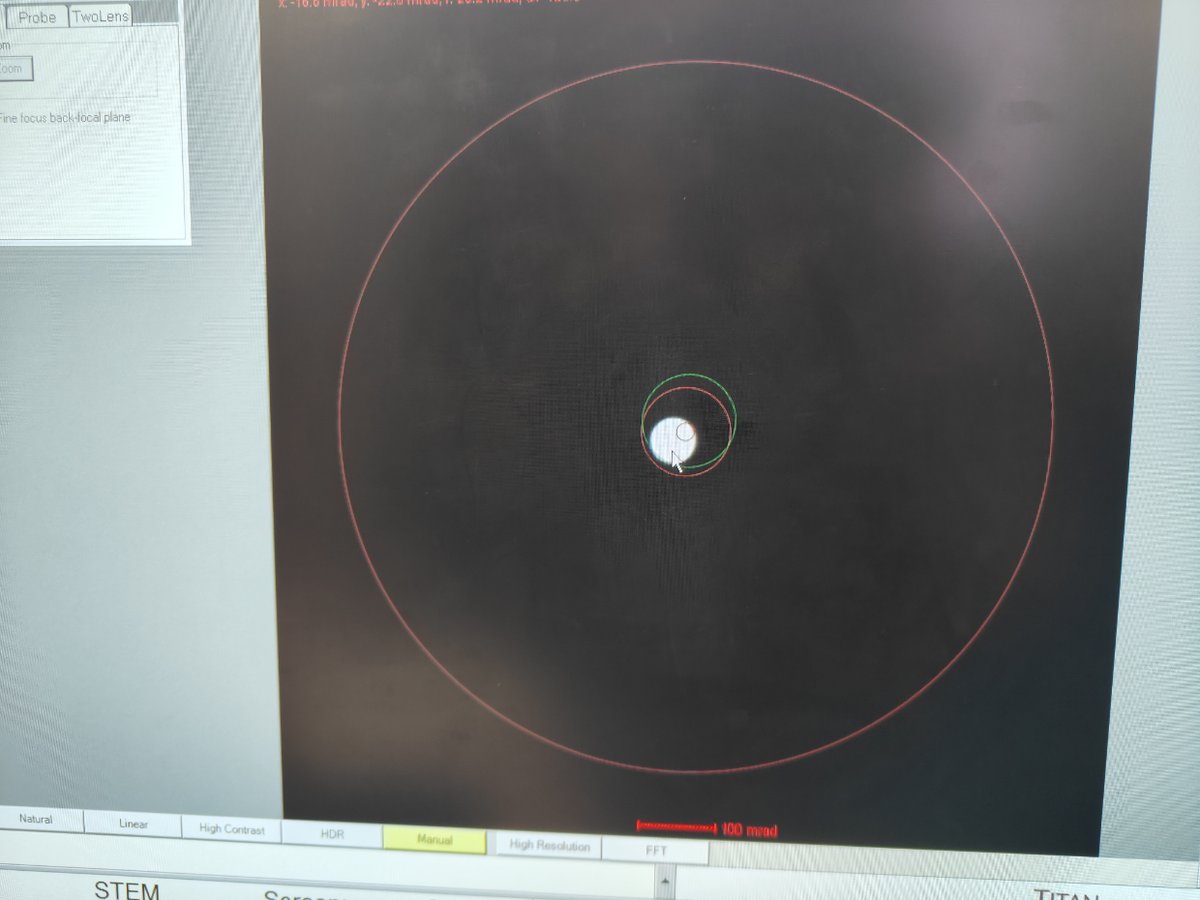
-
-
Enable EELS scanning
- Press
EELS Scanin the software
FIXME: attach image
- Press
-
Find a vacuum area
- Navigate to a vacuum region for initial alignment
-
Center the beam
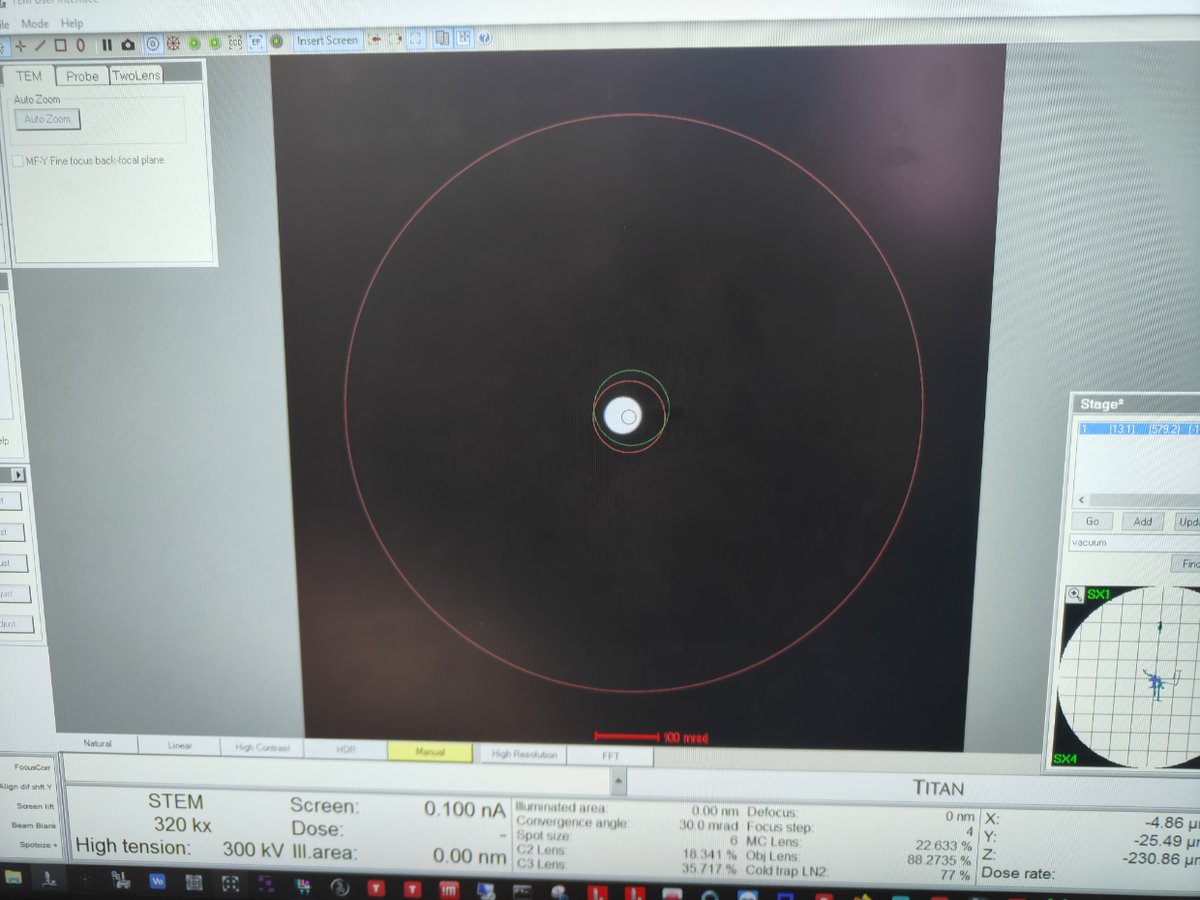
-
In EFTEM mode, use
mulXYknobs to center the beamBefore centering:
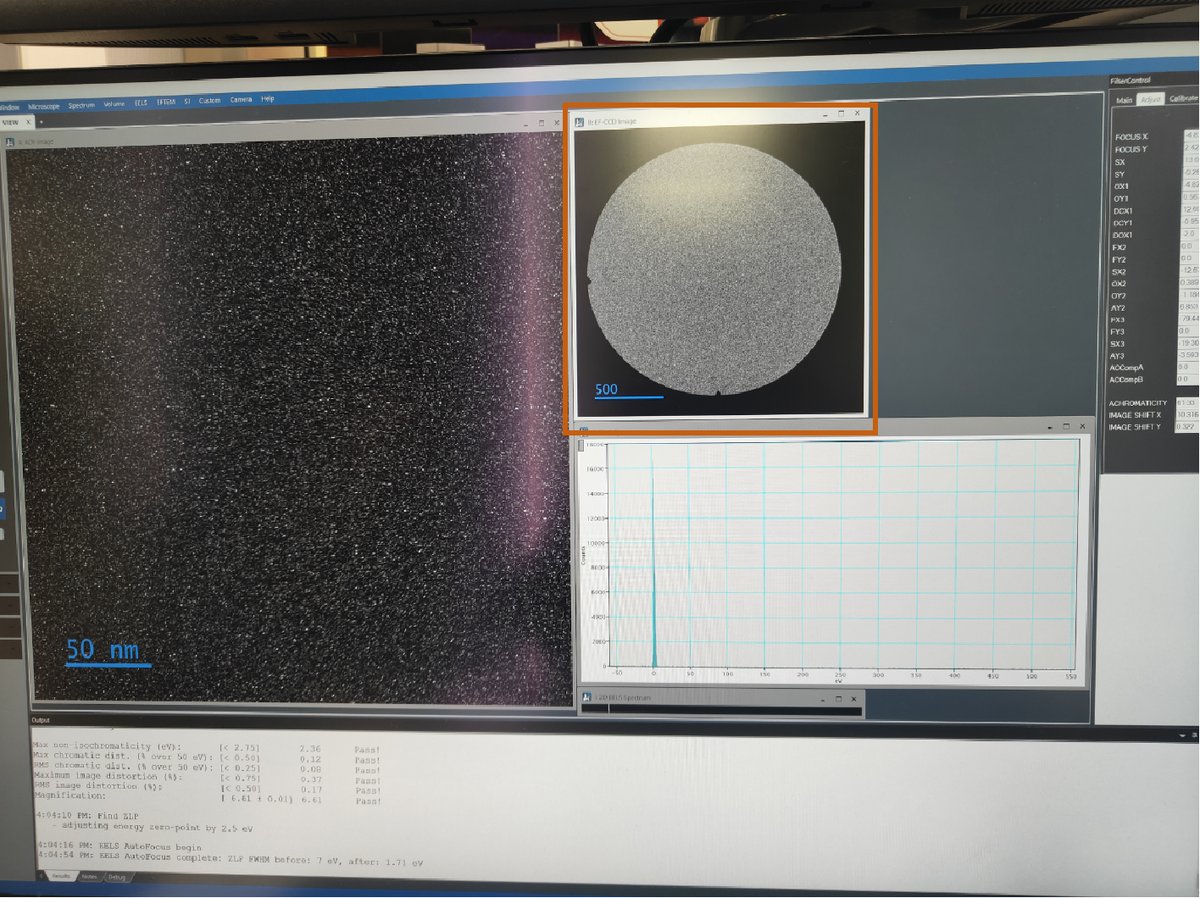
After centering:
-
-
Switch to STEM SI mode
-
Click
STEM SIto switch to Spectrum Imaging mode
-
-
Find sample area
- Navigate to an area of interest on your sample
-
Start scanning
-
Click
Scan→Viewto see the image
-
-
Adjust gain
- Right-click on ADF image → click
Auto Gain
FIXME: add image showing auto gain result
- Right-click on ADF image → click
-
Stop viewing
- Click
Viewagain to stop live scanning
- Click
-
Capture line scan (1D EELS)
-
Click
Captureand draw a line across the region of interest
-
Go to
EELS→User Mode
-
Go to
EELS→Zero Loss→Extract Zero Loss
FIXME: clarify what “extract zero loss” does and expected result
-
-
Capture area scan (2D EELS)
-
Click
Captureand select a rectangular area for 2D spectrum imaging
FIXME: add steps for analyzing 2D EELS data, expected output
-
Part 4: End session
Follow the steps in End session.
Changelog
- Dec 18, 2024 - initial rough draft by Guoliang Hu and @bobleesj
EDS
Caution
VERY ROUGH DRAFT - @bobleesj and Guoliang Hu took notes and pictures during training. This document will be updated with more detailed steps and images.
TODO:
- Add step-by-step images for TEM mode
- Clarify STEM mode beam settings
- Add screenshots for drift correction setup
This guide covers Energy Dispersive X-ray Spectroscopy (EDS) on the Spectra 300.
Prerequisite: Complete the STEM alignment procedure before starting.
Acronyms:
EDS- Energy Dispersive X-ray SpectroscopySI- Spectrum Imaging
Part 1: TEM mode EDS
-
Select area
- Select the area of interest on your sample
-
Open EDS
-
Click on the EDS icon in Velox

-
-
Check experiment log
- Review the experiment log for acquisition parameters
Part 2: STEM mode EDS
-
Set beam parameters
- Go to
Beam Setting→Probe→ click onMF-Y - Change convergence angle to 10 mrad (larger area to focus)
- Increase screen current to 0.4 nA
- Go to
-
Start spectrum imaging
-
Go to
SI(Spectrum Imaging) -
Click on the rectangle selection tool
-
Select an area on the sample
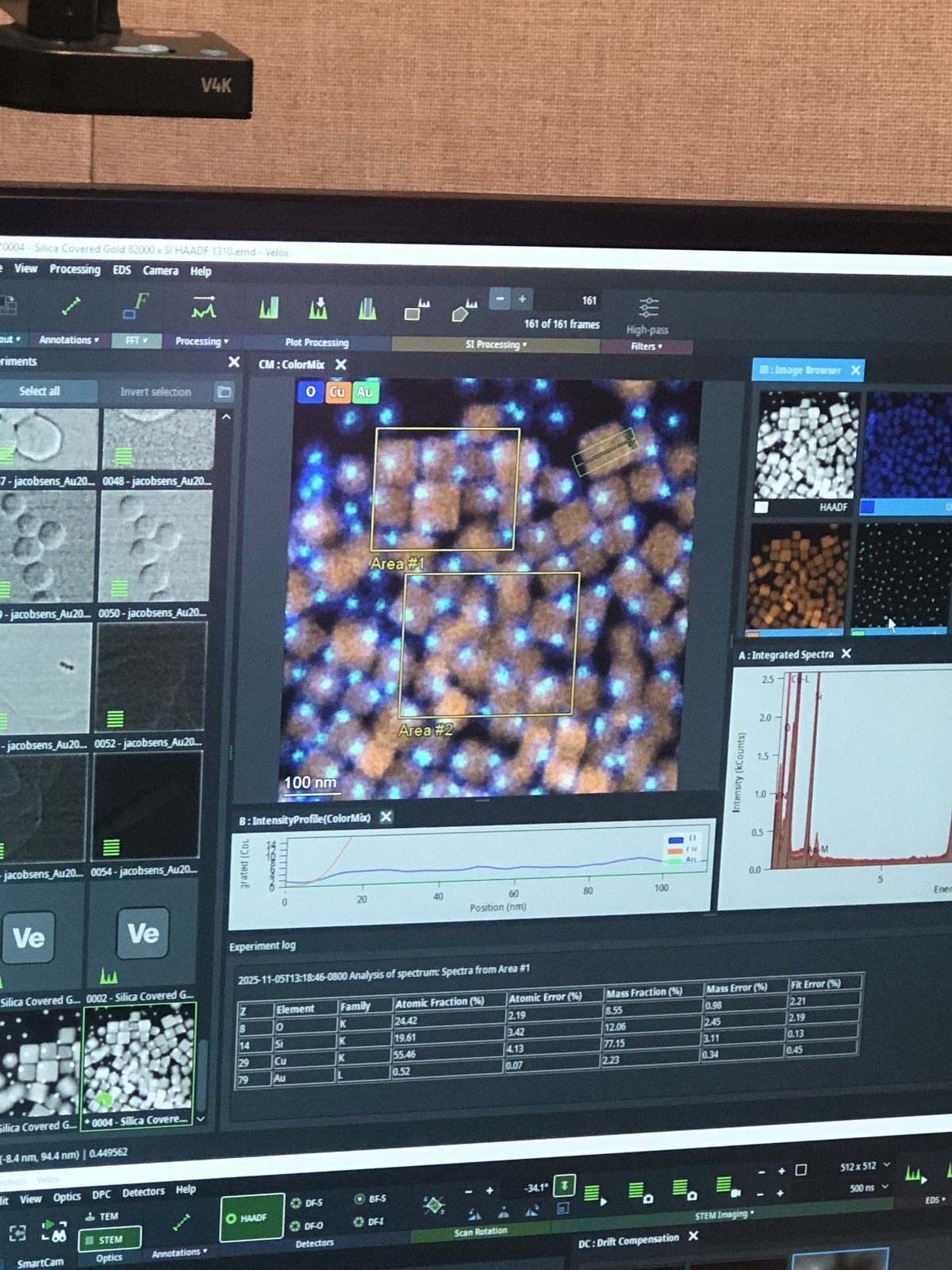
-
For nanoscale resolution, choose 20 microsecond dwell time
-
-
Set drift correction
- Click on
Drift Areato enable drift correction
FIXME: add image for drift area selection
- Click on
-
Adjust parameters
- Use
Object Propertiesto change acquisition parameters
- Use
-
Acquire data
-
Wait until pixel count stabilizes (watch for 8 pixel indicator)
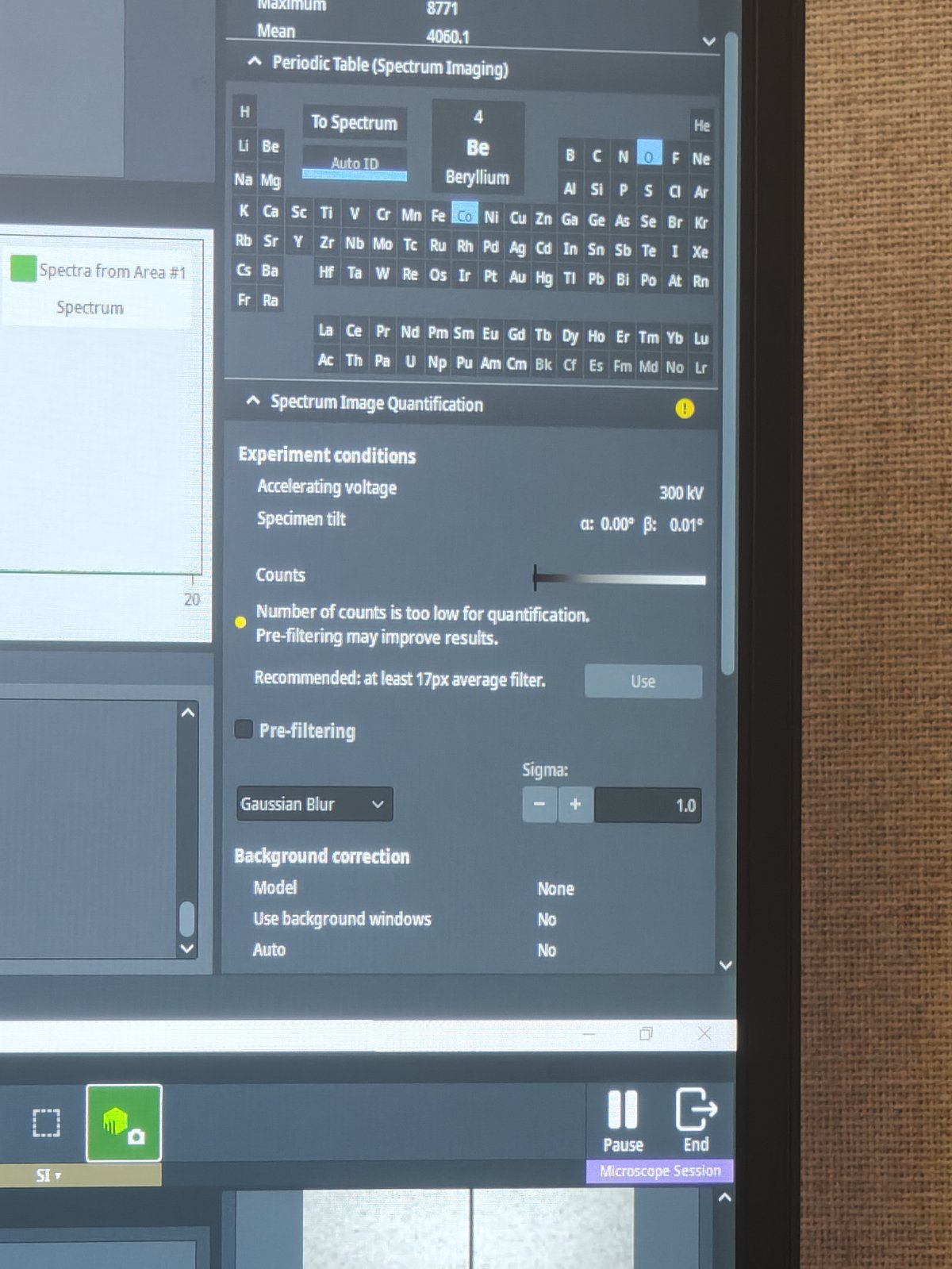
-
Stop recording when complete
-
Part 3: Data processing
-
Draw line profile
-
Go to
Processingto draw a line profile across features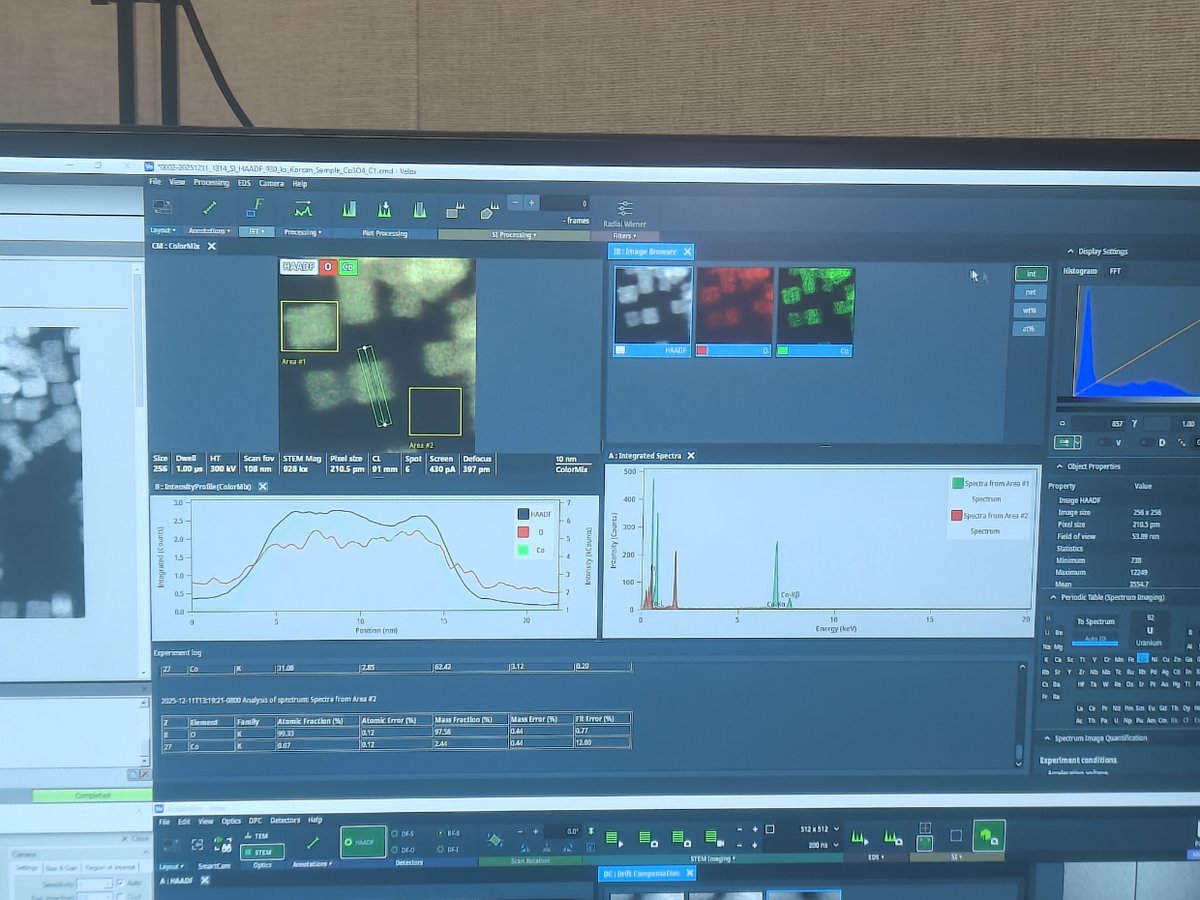
-
-
Select arbitrary area
- You can also select an arbitrary area for analysis
-
Atomic mapping
-
For atomic resolution mapping, use Average or Gaussian filtering
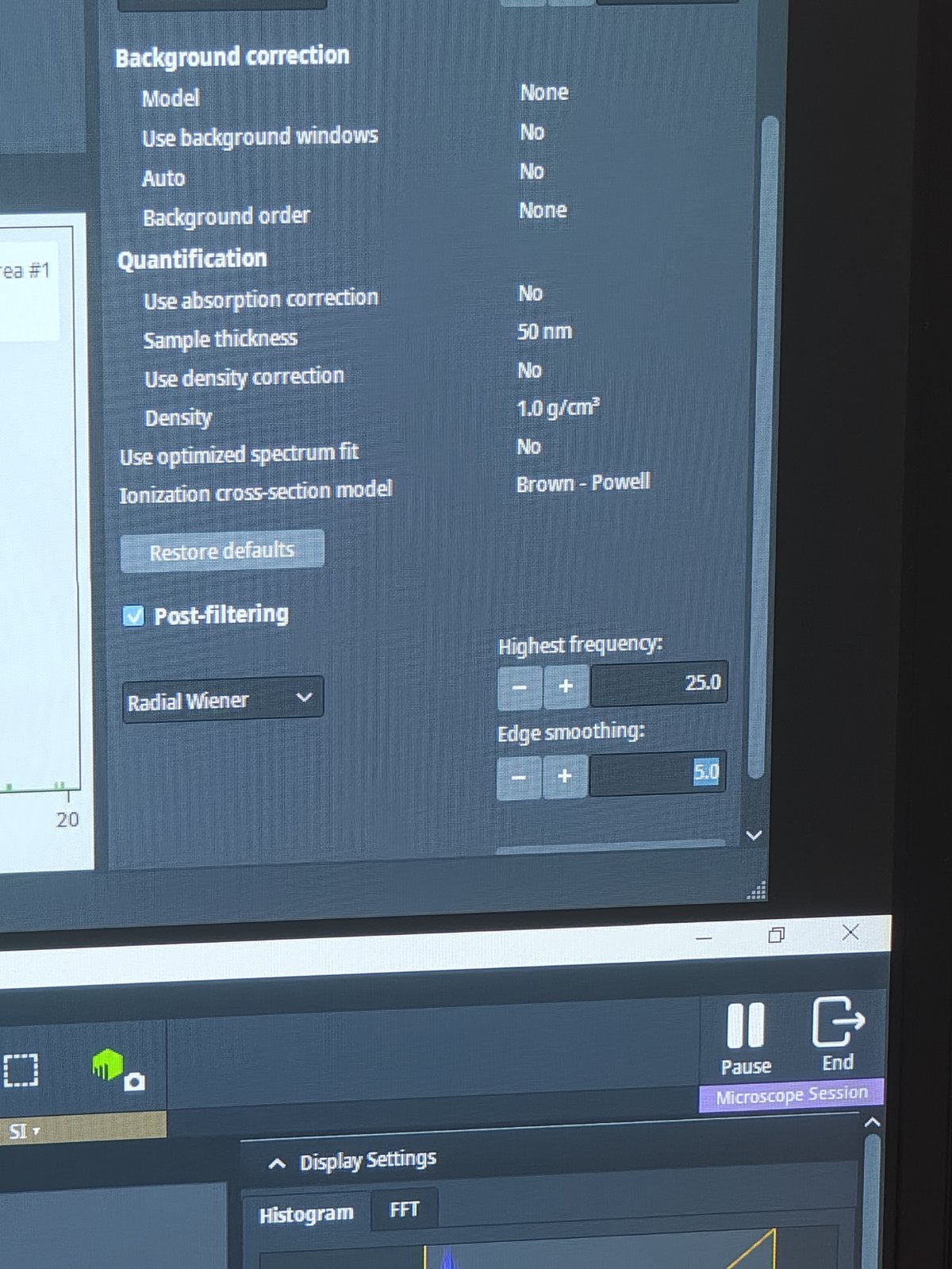
-
Use highest frequency and edge smoothing settings to obtain atomic resolution
-
Part 4: End session
Follow the steps in End session.
Changelog
- Dec 18, 2025 - initial rough draft by Guoliang Hu and @bobleesj
Tomography
TODO: Add tomography tutorial content
This guide covers electron tomography on the Spectra 300.
Changelog
- Dec 17, 2025 - Add placeholder by @bobleesj
Ptychography
For the experiment, record the global meta data:
- keV 300 keV
- convergence angle
- spot size
For region of interest, record the following
- Defocus (nm)
- Mag (Mx)
- Scan steps
- dwell time (μm)
- Camera length
- Data collection notes
- Mono
- Current on flucam: 75 pA
- Note (interesting shape? zone axis?)
Random notes:
- Why is it hard to find zone axis in nanoparticles? Small particles (1 - 10 nm)
PED
TODO: Add PED tutorial content
This guide covers Precession Electron Diffraction (PED) on the Spectra 300.
Changelog
- Dec 17, 2025 - Add placeholder
Share Your Procedure
We want to bring electron microscopy techniques and SOPs together across institutions.
Why contribute?
- Seamless experiments - Researchers traveling between facilities can quickly get up to speed on different microscopes
- Reduced friction - Standardized visual guides reduce training time and errors
- Community knowledge - Your expertise helps the broader EM community
What we’re looking for
- Step-by-step operating procedures for your microscope
- Visual guides with screenshots or photos
- Tips and tricks from experienced operators
- Troubleshooting guides
How to contribute
We can help with writing, formatting, and organizing your content. You don’t need to be a programmer!
- Prepare your notes - Word docs, Google Docs, handwritten notes, photos from your phone - anything works
- Reach out - Contact @bobleesj or open an issue on the GitHub repo
- We’ll handle the rest - We’ll format your content and add it to the site under your institution’s section
Institutions we’d love to hear from
- National labs
- University facilities
- Industry labs
- International facilities
Your procedures will be credited to you and your institution.
Changelog
- Dec 18, 2025 - Page created
Session notes
by Sangjoon Bob Lee
This is a working scratchpad for raw notes taken during microscopy training visits and sessions. Notes here capture practical tips, questions, and observations from hands-on time at the instrument. Over time, useful content gets refined and incorporated into the proper guide sections.
Experiment notes
Gold standard SOP practice: Mar 14, 2026
How to find the ronchigram again?
- Move the joystick around and watch for a small screen current change.
- Then use Diffraction Shift and Focus alignment to find the beam. You will see a faint bit of light.
Remaining questions
- When do you run Stigmator? During real space alignment?
- For C1A1, during the first run, what are the target values? I started with C1 6 nm, A1 27 nm, B2 928 nm. My notes say C1 < 1 nm and A1 < 3 nm. Add target values to the table.
- When do you do manual tuning? With real samples?
- Add a table at the beginning with the target values for aberration corrections.
Open questions
- Ideal Velox play setting? 1024x1024 and 500 ns for gold standard sample.
- When to change condenser stig?
- Camera length and ronchigram size? Proportional: 91 mm to 115 mm, ronchigram size also increases.
- When do you adjust Condenser?
Lessons learned
- Beam condition from FFT: 4-6 rings, each ring with discrete peaks, ~70 pm. Further rings (bigger k) correspond to sharper features resolved in real space.
- Beam condition from Probe corrector: flatness around the “green” aberration surface is the key.
- Overfocus means focal point above the sample; underfocus means below it.
- Defocus change DP? Barely, but real-space probe size is changed (needed for ptycho overlaps).
- Defocus on BF? Expect it to get worse. Ptycho will perform better.
- Focus knob to sharpen? Minimize it. Don’t add degrees of freedom. 20 nm max, use stage piezo and knobs.
- Finding ptycho defocus step size: use ronchigram shadow image to determine feature size variations. Step size of 1 nm is too small.
- Why drift? Inserting the holder itself induces aberrations. After stage movement, find ROI, then wait ~5 min for mechanical stabilization.
Extra notes on aberrations
- Practice getting atomic imaging without Sherpa. Example: Samsung sample, too beam-sensitive for Sherpa.
- Tableau with A5 selected measures up to 5th order aberrations. The following screenshot shows a Tableau measurement result with A5 enabled.

Manual aberration correction without Sherpa (beam-sensitive samples)
- Figure out the full workflow for correcting aberrations without Sherpa.
- Where do you run C1A1 if you can’t expose the sample?
- How do you manually adjust A1 and B2?
- What does “good” look like without Sherpa? Define criteria for the ronchigram, FFT, and probe shape.
- What is the minimum correction quality needed for atomic resolution?
Action steps for future practice sessions
- Try going back to TEM and STEM, confirm aberrations getting worse.
- Try going to LM and then back to regular STEM, confirm resolution getting worse.
- Try correcting the probe without Sherpa, in the case of beam sensitivity.
Cobalt oxide nanoparticles STEM — Mar 3, 2026
It was my first time staying in STEM mode and find samples after STEM probe correction and loading my own sample. The following notes were taken in my attempt to find the feature of interest right after the sample was loaded.
Finding the sample after the sample is loaded
- Go to 5,000× magnification. If there is no beam, it means the beam is blocked on the grid. Move the stage around with the joystick.
- Move the stage until the screen current increases to about 0.150 nA. At this point, the beam has been found. Notice the Kikuchi bands: this is a good clue that you are in a good starting place.
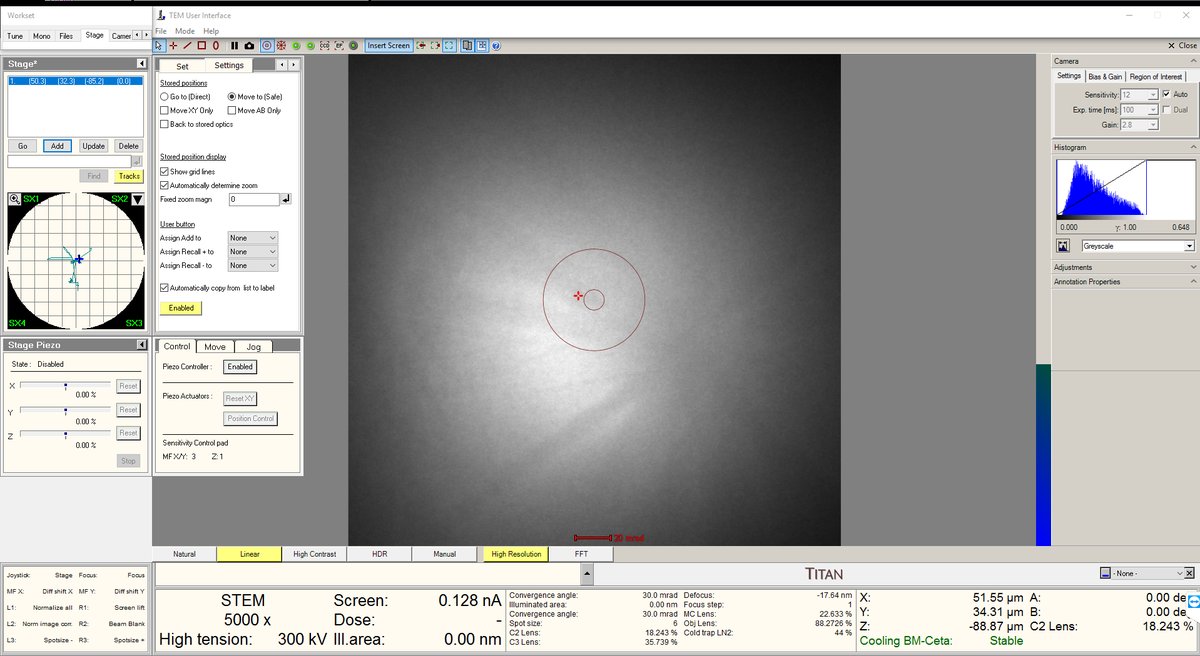
- Increase the magnification to 20,000× or higher. The features will still look blurry since we are not yet at the correct focus.
- Press the Z-axis up and down until the Ronchigram blow-up point appears. Adjust z-axis from 5kx to 20kx to find the blow-up point.
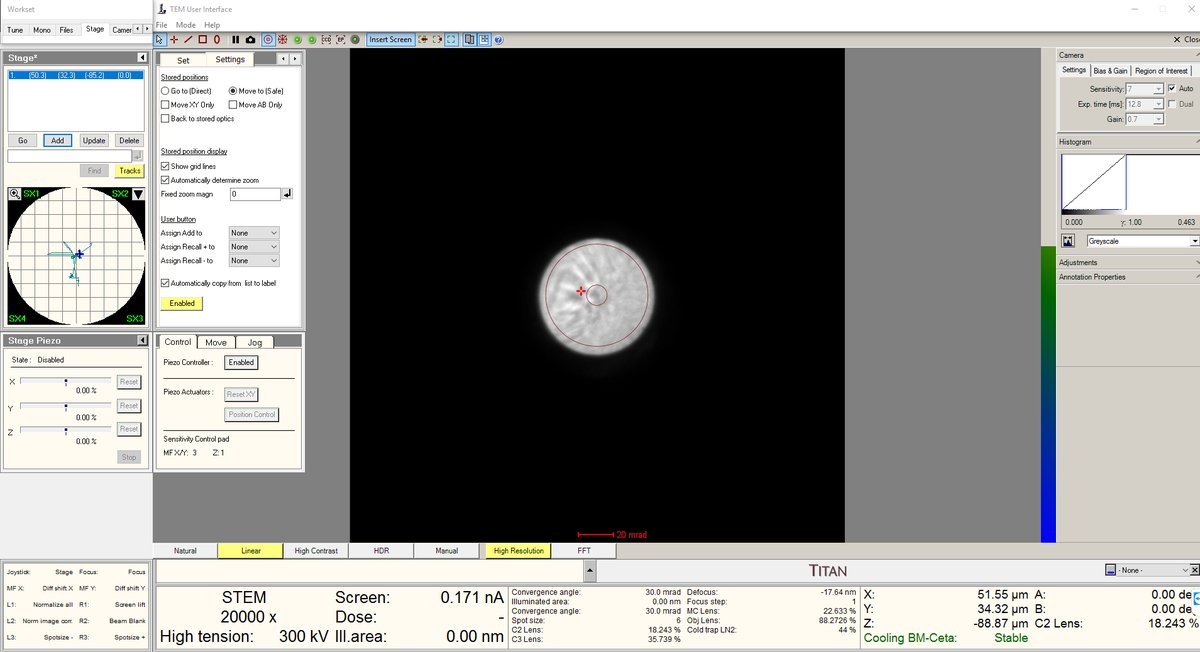
- After the blow-up point, you will see features. In this session, cubes were found after the Ronchigram blow-up point.
- Stay at the blow-up point. We are now at the eucentric height.
- Turn on the HAADF camera in Velox to observe those features.
Getting sharper features after you’ve found them
- Use the stage piezo to move the sample around to ensure you have the sharpest features.
- Use Sharpa to correct C1A1. You do not need to correct B2A2.
- Take the image as usual.
Notes on aberration correctinos
MAPED experience at NCEM, Mar 2, 2026
I had a chance to join TEAM MAPED session at NCEM with Stephanie Ribet and Henry Bell.
General STEM notes for aberration
- LM mode warning: In TEAM, LM mode isn’t used generally. It turns off the aberration corrector, a set of multipole electromagnetic lenses (hexapoles, octupoles) that correct for spherical aberration. When switched back on, the corrector needs hours to restabilize both thermally (coils heat up, causing alignment drift from thermal expansion) and electromagnetically (currents must settle to precise values).
- aberration knobs: in the probe corrector software, manually fix and it can be faster since software can take a while. For example, Correct A1 (twofold astigmatism) and B2 (axial coma) by hand. (@bobleesj, verifiy this)
- Use the stigmator button on the hand panel, it makes the beam round.
- C2 adjust is used to make the beam concentric, by alinigng C2 aperture.
- Rotation center: don’t care about the edges. Use a magnified image to see whether the features are pulsing out of the page.
Sample loading and vacumm
- After loading a sample, watch PPL. It should go down to low 10⁻³ or 10⁻⁴.
- Octagon must be below 10 after sample loading.
Finding sample ROIs
- In Spectra, you can switch between TEM and STEM modes and it’s stable. On TEAM, this is not the case, so it’s better to use STEM at 5k mag to navigate and find samples.
- Use stage double-click to move around.
Zone axis
- Use alpha and beta on the hand panel to get an approximation, then go to Stage, flap out, and use alpha and beta for fine adjustment.
- Feel free to use camera length to make it easier to see. Ensure the ronchigram is symmetric.
Convergence angle
- Change the convergence angle by changing the aperture.
- C2 for 70 µm aperture gives ~9 mrad max. For higher convergence, use another aperture.
- If you change the C2 aperture, the software may still display the old value (e.g., “20”) because it doesn’t know how to get to the new position. Click “Adjust” to move to the new aperture where it has the intended aperature like 10 micrometer bull’s eye aperture.
- Then move C2 to ~30 (instead of 1,000) to block out other apertures.
Arina at NCEM
- HAADF must be out before you insert Arina. Verify on Digital Micrograph.
- Shutdown order: voltage can be turned off from the software. No need to physically turn it off unlike at SNSF.
Colume valve must be closed even during a quick break
- Always set column valve off for lunch. It does not affect aberrations.
Modify current
- To gain more current, go to Focus and Shift under the
Monotab. This controls the monochromator lens excitation, which determines how tightly the beam is focused at the energy-selecting slit. Lowering focus means a less tight crossover at the slit, so more electrons pass through.
Descan pivot point
- TODO: needs definition and procedure.
TODOs
- Investigate the effect of descan when you integrate or sum across k-space
Reconstruction notes
Ptychography reconstruction basics — Mar 9, 2026
Notes from Arthur on reconstructing data collected from ARINA detector at Stanford.
- Sign convention: in
quantem,C10 > 0means underfocus. The beam focal point is below the sample, hence negative defocus. - Aberrations: SSB is somewhat an “eye test.” One may use the aberrations from SSB or not. There are many degrees of freedom: batch size, the “dose” step size (finer can be better), probe size, center of mass/transpose, and initial aberrations.
- Cropping strategy: in real space, it’s fine to crop, encouraged since faster. In k space, we generally don’t want to crop since we lose the max scattering angle, i.e. we lose fine details in real space.
- Probe: aberrations should be identical across all scan regions in theory. However, for ptycho-tomo, defocus will change with tilt.
- CNN reconstruction: reconstruction weight is different between reconstructions since these are weights being trained.
- Memory: It’s hard to manage memory well in Jupyter notebook but it’s something we can work on.
- Virus samples: there is no zone axis, so we can’t do atomic resolution.
- Descan: the beam is tilted from the source and then tilted back after a short travel perpendicular to the sample. During this second tilt, instability can be introduced and the diffraction pattern isn’t perfectly aligned.
- Mixed probe: mixed probe is good and orthogonality is imposed, so probes should look different from each other.